
| Sequence ID | dm3.chr3R |
|---|---|
| Location | 11,221,280 – 11,221,372 |
| Length | 92 |
| Max. P | 0.881892 |

| Location | 11,221,280 – 11,221,372 |
|---|---|
| Length | 92 |
| Sequences | 6 |
| Columns | 98 |
| Reading direction | forward |
| Mean pairwise identity | 82.47 |
| Shannon entropy | 0.32353 |
| G+C content | 0.42406 |
| Mean single sequence MFE | -23.38 |
| Consensus MFE | -14.31 |
| Energy contribution | -14.20 |
| Covariance contribution | -0.11 |
| Combinations/Pair | 1.25 |
| Mean z-score | -2.25 |
| Structure conservation index | 0.61 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.05 |
| SVM RNA-class probability | 0.881892 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
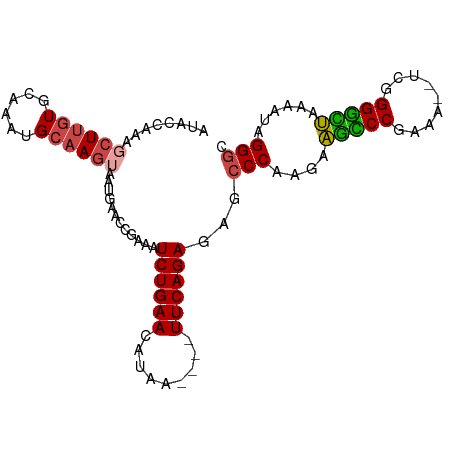
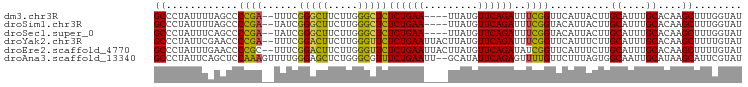
>dm3.chr3R 11221280 92 + 27905053 AUACCAAAGCUUGUGCAAAUGCAAGUAAUGAACCGAAAUCUGAACAUAA----UUCAGAGAGCCCAAGAAGCCCGAAA--UCGGGGCUAAAAUAGGGC ....((..((((((......))))))..))........((((((.....----))))))..((((....(((((....--...)))))......)))) ( -24.90, z-score = -2.72, R) >droSim1.chr3R 17266466 92 - 27517382 AUACCAAAGCUUGUGCAAAUGCAAGUAAUGUACCGAAAUCUGAACAUAA----UUCAGAGAGCCCAAGAAGCCCGAUA--UCGGGGCUAAAAUAGGGC ........((((((......))))))............((((((.....----))))))..((((....(((((....--...)))))......)))) ( -24.70, z-score = -2.39, R) >droSec1.super_0 10393897 92 - 21120651 AUACCAAAGCUUGUGCAAAUGCAAGUAAUGUACCGAAAUCUGAACAUAA----UUCAGAGAGCCCAAGAAGCCCGAUA--UCGGGGCUGAAAUAGGGC ........((((((......))))))............((((((.....----))))))..((((....(((((....--...)))))......)))) ( -25.80, z-score = -2.52, R) >droYak2.chr3R 13621569 96 - 28832112 AUACAAAAGCUUGUGCAAAUGCAAGAAAUGAACCGAAAUCUGAACAUAAGUAAUUCAGAGAACCCAAGAAGUCCGAAA--UCGGGGUUCGAAUAGGGC ...((....(((((......)))))...))........((((((.........))))))...(((..(((.((((...--.)))).))).....))). ( -20.80, z-score = -1.67, R) >droEre2.scaffold_4770 10747383 96 + 17746568 AUACAAAAGCUUGUGCAAAUGCAAGAAAUGAACCGAUAUCUGAACAUAAGUAAUUCAGAGAACCCAAGAAGUCCGAAA--GCGGGGUUCAAAUAGGGC ...((....(((((......)))))...))..((....((((((.........))))))((((((.....((......--)).)))))).....)).. ( -20.40, z-score = -1.60, R) >droAna3.scaffold_13340 12805764 96 + 23697760 AUACGAAUGCUUAUGCAAUUGCCACUAAAGAACAAAACUCUGAAUAUGC--AAUUCAGAAACGCCCAGAGCUCCCAAAACUUUGGAGCUGAAUAGGGC ...(((.(((....))).))).................(((((((....--.)))))))...((((..((((((.........)))))).....)))) ( -23.70, z-score = -2.61, R) >consensus AUACCAAAGCUUGUGCAAAUGCAAGUAAUGAACCGAAAUCUGAACAUAA____UUCAGAGAGCCCAAGAAGCCCGAAA__UCGGGGCUAAAAUAGGGC ........((((((......))))))............((((((.........))))))...(((....(((((.........)))))......))). (-14.31 = -14.20 + -0.11)


| Location | 11,221,280 – 11,221,372 |
|---|---|
| Length | 92 |
| Sequences | 6 |
| Columns | 98 |
| Reading direction | reverse |
| Mean pairwise identity | 82.47 |
| Shannon entropy | 0.32353 |
| G+C content | 0.42406 |
| Mean single sequence MFE | -23.80 |
| Consensus MFE | -14.61 |
| Energy contribution | -13.50 |
| Covariance contribution | -1.11 |
| Combinations/Pair | 1.32 |
| Mean z-score | -1.78 |
| Structure conservation index | 0.61 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.13 |
| SVM RNA-class probability | 0.555505 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
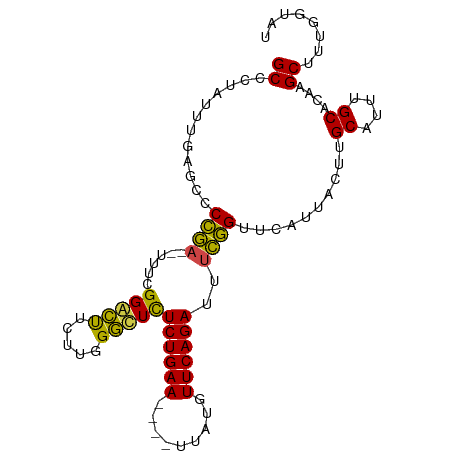
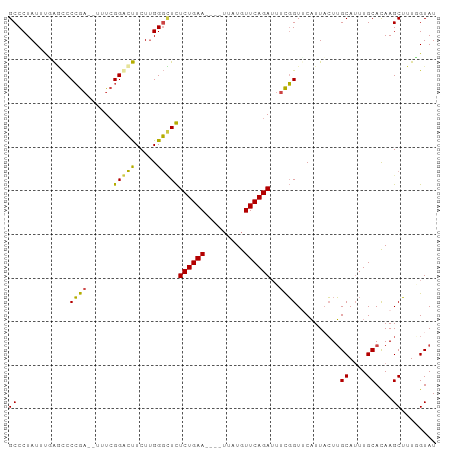
>dm3.chr3R 11221280 92 - 27905053 GCCCUAUUUUAGCCCCGA--UUUCGGGCUUCUUGGGCUCUCUGAA----UUAUGUUCAGAUUUCGGUUCAUUACUUGCAUUUGCACAAGCUUUGGUAU ((((......(((((.(.--...))))))....))))..((((((----(...)))))))...............(((....((....))....))). ( -23.50, z-score = -1.45, R) >droSim1.chr3R 17266466 92 + 27517382 GCCCUAUUUUAGCCCCGA--UAUCGGGCUUCUUGGGCUCUCUGAA----UUAUGUUCAGAUUUCGGUACAUUACUUGCAUUUGCACAAGCUUUGGUAU ((((......(((((.(.--...))))))....))))..((((((----(...))))))).....((((....((((........)))).....)))) ( -24.50, z-score = -1.72, R) >droSec1.super_0 10393897 92 + 21120651 GCCCUAUUUCAGCCCCGA--UAUCGGGCUUCUUGGGCUCUCUGAA----UUAUGUUCAGAUUUCGGUACAUUACUUGCAUUUGCACAAGCUUUGGUAU ((((......(((((.(.--...))))))....))))..((((((----(...))))))).....((((....((((........)))).....)))) ( -23.80, z-score = -1.40, R) >droYak2.chr3R 13621569 96 + 28832112 GCCCUAUUCGAACCCCGA--UUUCGGACUUCUUGGGUUCUCUGAAUUACUUAUGUUCAGAUUUCGGUUCAUUUCUUGCAUUUGCACAAGCUUUUGUAU .........((((((((.--...))).......(((...(((((((.......))))))).))))))))......((((...((....))...)))). ( -21.30, z-score = -1.93, R) >droEre2.scaffold_4770 10747383 96 - 17746568 GCCCUAUUUGAACCCCGC--UUUCGGACUUCUUGGGUUCUCUGAAUUACUUAUGUUCAGAUAUCGGUUCAUUUCUUGCAUUUGCACAAGCUUUUGUAU ........((((((....--....(((((.....)))))(((((((.......)))))))....)))))).....((((...((....))...)))). ( -21.60, z-score = -2.12, R) >droAna3.scaffold_13340 12805764 96 - 23697760 GCCCUAUUCAGCUCCAAAGUUUUGGGAGCUCUGGGCGUUUCUGAAUU--GCAUAUUCAGAGUUUUGUUCUUUAGUGGCAAUUGCAUAAGCAUUCGUAU ((((.....((((((.........))))))..))))((((.((((((--((.((((.((((.......)))))))))))))).)).))))........ ( -28.10, z-score = -2.07, R) >consensus GCCCUAUUUGAGCCCCGA__UUUCGGACUUCUUGGGCUCUCUGAA____UUAUGUUCAGAUUUCGGUUCAUUACUUGCAUUUGCACAAGCUUUGGUAU ((............((((......(((((.....)))))((((((.........))))))..))))..........((....))....))........ (-14.61 = -13.50 + -1.11)
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:17:14 2011