| Sequence ID | dm3.chr3R |
|---|---|
| Location | 10,975,315 – 10,975,410 |
| Length | 95 |
| Max. P | 0.974354 |

| Location | 10,975,315 – 10,975,410 |
|---|---|
| Length | 95 |
| Sequences | 10 |
| Columns | 114 |
| Reading direction | forward |
| Mean pairwise identity | 67.66 |
| Shannon entropy | 0.64590 |
| G+C content | 0.46548 |
| Mean single sequence MFE | -18.51 |
| Consensus MFE | -10.45 |
| Energy contribution | -11.20 |
| Covariance contribution | 0.75 |
| Combinations/Pair | 1.23 |
| Mean z-score | -1.28 |
| Structure conservation index | 0.56 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.91 |
| SVM RNA-class probability | 0.974354 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
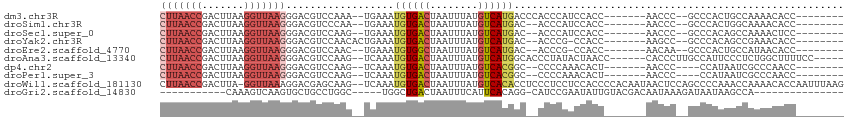
>dm3.chr3R 10975315 95 + 27905053 CUUAACCGACUUAAGGUUAAGGGACGUCCAAA--UGAAAUGUGACUAAUUUAUGUCAUGACCCACCCAUCCACC-------AACCC--GCCCACUGCCAAAACACC-------- (((((((.......)))))))((((((....)--))....(((((........)))))..........)))...-------.....--..................-------- ( -15.30, z-score = -0.53, R) >droSim1.chr3R 17030976 93 - 27517382 CUUAACCGACUUAAGGUUAAGGGACGUCCCAA--UGAAAUGUGACUAAUUUAUGUCAUGAC--ACCCAUCCACC-------AACCC--GCCCACUGGCAAAACACC-------- (((((((.......)))))))((..(((....--......(((((........))))))))--..)).......-------.....--(((....)))........-------- ( -19.80, z-score = -1.48, R) >droSec1.super_0 10149115 93 - 21120651 CUUAACCGACUUAAGGUUAAGGGACGUCCAAG--UGAAAUGUGACUAAUUUAUGUCAUGAC--ACCCAUCCACC-------AACCC--GCCCACAGCCAAAACUCC-------- .((((((.......))))))(((.((.....(--((....(((((........)))))((.--.....))))).-------....)--))))..............-------- ( -17.20, z-score = -0.80, R) >droYak2.chr3R 13377280 94 - 28832112 CUUAACCGACUUAAGGUUAAGGGACGUCCAACACUGAAAUGUGACUAAUUUAUGUCAUGAC--ACCCG-CCACC-------AAGCC--GCCCACAGCCGAAACACC-------- .((((((.......))))))(((.((..............(((((........)))))...--....(-(....-------..)))--))))..............-------- ( -17.10, z-score = -0.53, R) >droEre2.scaffold_4770 10501511 92 + 17746568 CUUAACCGACUUAAGGUUAAGGGACGUCCAAC--UGAAAUGUGGCUAAUUUAUGUCAUGAC--ACCCG-CCACC-------AACAA--GCCCACUGCCAUAACACC-------- (((((((.......)))))))((....))...--.....((((((.......(((....))--)...(-(....-------.....--)).....)))))).....-------- ( -17.50, z-score = -0.33, R) >droAna3.scaffold_13340 6063807 101 + 23697760 CUUAACCGACUUAAGGUUAAGGGACGUCCAAG--UCAAAUGUGACUAAUUUAUGUCAUGGCACCCUAUACUAACC------CACCCUUGCCAUUCCCUCUGGCUUUUCC----- (((((((.......)))))))((....))(((--(((...(((((........)))))((((.............------......))))........))))))....----- ( -23.01, z-score = -1.62, R) >dp4.chr2 20958579 91 + 30794189 CUUAACCGACUUAAGGUUAAGGGACGUCCAAG--UCAAAUGUGACUAAUUUAUGUCACGGC--CCCCAAACACU-------AACCC----CCAUAAUCGCCCAACC-------- (((((((.......))))))).(((......)--))....(((((........)))))(((--...........-------.....----........))).....-------- ( -19.66, z-score = -1.92, R) >droPer1.super_3 3708555 91 + 7375914 CUUAACCGACUUAAGGUUAAGGGACGUCCAAG--UCAAAUGUGACUAAUUUAUGUCACGGC--CCCCAAACACU-------AACCC----CCAUAAUCGCCCAACC-------- (((((((.......))))))).(((......)--))....(((((........)))))(((--...........-------.....----........))).....-------- ( -19.66, z-score = -1.92, R) >droWil1.scaffold_181130 9371389 111 - 16660200 CUUAACCGACUUA-GGUUAAAGGACGAGCAAG--UCAAAUGUGACUAAUUUAUGUCACACCUCCCUCCUCCACCCCACAAUAACUCCAGCCCCAAACCAAAACACCAAUUUAAG .((((((......-)))))).(((.(((..((--.....((((((........)))))).))..))).)))........................................... ( -20.20, z-score = -3.64, R) >droGri2.scaffold_14830 3510460 82 + 6267026 -----------CAAAGUCAAGUGCUGCCUGGC-----UGGCUGACUAAUUUCAUUCACAGG-CAUCCGAAUAUUGUACGACAAUAAAGAUAAUAAGCCA--------------- -----------.........(.((((((((..-----((..(((......)))..))))))-).......(((((.....))))).........)))).--------------- ( -15.70, z-score = -0.03, R) >consensus CUUAACCGACUUAAGGUUAAGGGACGUCCAAG__UGAAAUGUGACUAAUUUAUGUCAUGAC__ACCCAACCACC_______AACCC__GCCCACAGCCAAAACACC________ (((((((.......)))))))..................((((((........))))))....................................................... (-10.45 = -11.20 + 0.75)
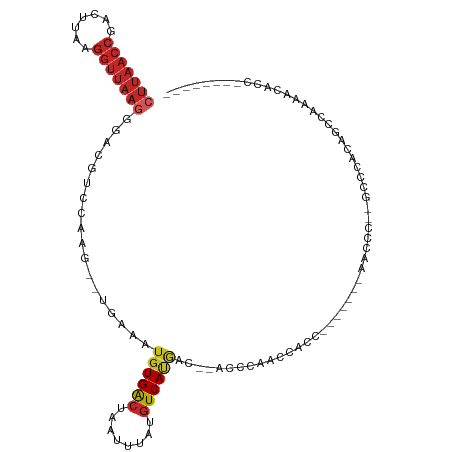
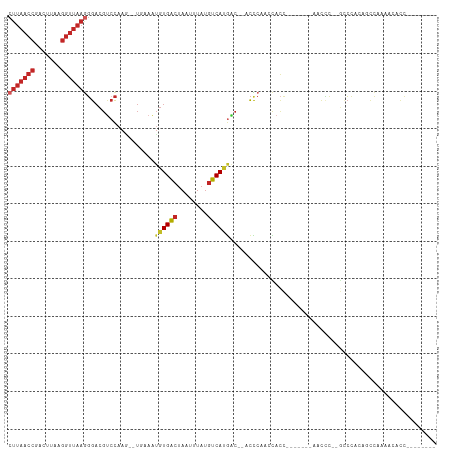
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:16:43 2011