
| Sequence ID | dm3.chr3R |
|---|---|
| Location | 10,615,629 – 10,615,705 |
| Length | 76 |
| Max. P | 0.593042 |

| Location | 10,615,629 – 10,615,705 |
|---|---|
| Length | 76 |
| Sequences | 12 |
| Columns | 76 |
| Reading direction | forward |
| Mean pairwise identity | 86.00 |
| Shannon entropy | 0.31478 |
| G+C content | 0.59063 |
| Mean single sequence MFE | -29.57 |
| Consensus MFE | -17.67 |
| Energy contribution | -18.15 |
| Covariance contribution | 0.48 |
| Combinations/Pair | 1.33 |
| Mean z-score | -1.90 |
| Structure conservation index | 0.60 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.21 |
| SVM RNA-class probability | 0.593042 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
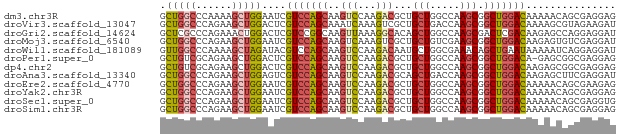
>dm3.chr3R 10615629 76 + 27905053 GCUGGCCCAAAAGCUGGAAUCGUCCAGCAAGUCCAAGACGCUGCUGGCCAAGCGGCUGGACAAAAACAGCGAGGAG ((((........((((((....))))))..(((((....((((((.....))))))))))).....))))...... ( -32.40, z-score = -2.85, R) >droVir3.scaffold_13047 11698739 76 + 19223366 GCUGGCCCAGAAGCUGGACUCGUCCAGCAAAUCAAAGUCGCUGCUGACCAAGCGGCUGGACAAAAGCGUAGAAGAU (((.(.(((...((((((....))))))...........((((((.....))))))))).)...)))......... ( -27.60, z-score = -2.11, R) >droGri2.scaffold_14624 1777477 76 + 4233967 GCUCGCCCAGAAACUGGACUCGUCCGGCAAGUUAAAGGCACAGCUGGCCAAGCGACUCGACAAGAGCCAGGAGGAU .((((((...((.(((((....)))))....))...)))....((((((...((...))....).))))))))... ( -20.30, z-score = 0.57, R) >droMoj3.scaffold_6540 4077586 76 - 34148556 GCUGGCCCAGAAGCUGGAAUCGUCCAGCAAGUCAAAGUCGCUGCUGUCGAAGCGGCUGGACAAGAGUGUCGAGGAU .(((((.(....((((((....))))))..(((..(((((((........))))))).)))....).))).))... ( -26.40, z-score = -0.89, R) >droWil1.scaffold_181089 9750867 76 - 12369635 GUUGGCCCAAAAGCUAGAUACGUCCAGCAAGUCCAAGACAAUGCUGGCGAAACAGCUGAAUAAAAAUCAGGAGGAU .(((((......)))))...((.((((((.(((...)))..))))))))......((((.......))))...... ( -18.00, z-score = -1.30, R) >droPer1.super_0 1512083 75 + 11822988 GCUGUCGCAGAAGCUGGACUCGUCCAGCAAGUCCAAGACGCUGCUGGCCAAGCGGCUGGACA-GAGCGGCGAGGAG .(((((((....((((((....))))))..(((((....((((((.....))))))))))).-..))))).))... ( -36.10, z-score = -2.21, R) >dp4.chr2 7689288 76 + 30794189 GCUGUCGCAGAAGCUGGACUCGUCCAGCAAGUCCAAGACGCUGCUGGCCAAGCGGCUGGACAAGAGCGGCGAGGAG .(((((((....((((((....))))))..(((((....((((((.....)))))))))))....))))).))... ( -36.10, z-score = -2.30, R) >droAna3.scaffold_13340 13616192 76 - 23697760 GCUGGCCCAGAAGCUGGAGUCGUCCAGCAAGUCCAAGACGCAGCUGACCAAGCGGCUGGACAAGAGCUUCGAGGAU ((((((((((...)))).)))(((((((..(((...)))...(((.....))).)))))))...)))......... ( -28.30, z-score = -1.18, R) >droEre2.scaffold_4770 10149917 76 + 17746568 GCUGGCCCAGAAGCUGGAAUCGUCCAGCAAGUCCAAGACGCUGCUGGCCAAGCGGCUGGACAAAAACAGCGAAGAG ((((........((((((....))))))..(((((....((((((.....))))))))))).....))))...... ( -32.40, z-score = -2.96, R) >droYak2.chr3R 13018398 76 - 28832112 GCUGGCCCAGAAGCUGGAAUCGUCCAGCAAGUCCAAGACGCUGCUGGCCAAGCGGCUGGACAAAAACAGCGAGGAG ((((........((((((....))))))..(((((....((((((.....))))))))))).....))))...... ( -32.40, z-score = -2.58, R) >droSec1.super_0 9801582 76 - 21120651 GCUGGCCCAGAAGCUGGAAUCGUCCAGCAAGUCCAAGACGCUGCUGGCCAAGCGGCUGGACAAAAACAGCGAGGUG ((((........((((((....))))))..(((((....((((((.....))))))))))).....))))...... ( -32.40, z-score = -2.41, R) >droSim1.chr3R 16672336 76 - 27517382 GCUGGCCCAGAAGCUGGAAUCGUCCAGCAAGUCCAAGACGCUGCUGGCCAAGCGGCUGGACAAAAACAGCGAGGAG ((((........((((((....))))))..(((((....((((((.....))))))))))).....))))...... ( -32.40, z-score = -2.58, R) >consensus GCUGGCCCAGAAGCUGGAAUCGUCCAGCAAGUCCAAGACGCUGCUGGCCAAGCGGCUGGACAAAAACAGCGAGGAG .(((((......)))))....(((((((..(((...)))...(((.....))).)))))))............... (-17.67 = -18.15 + 0.48)


Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:15:51 2011