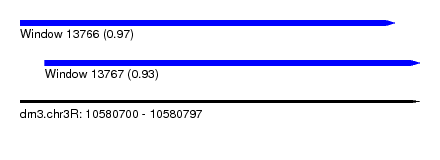

| Sequence ID | dm3.chr3R |
|---|---|
| Location | 10,580,700 – 10,580,797 |
| Length | 97 |
| Max. P | 0.968057 |

| Location | 10,580,700 – 10,580,791 |
|---|---|
| Length | 91 |
| Sequences | 5 |
| Columns | 106 |
| Reading direction | forward |
| Mean pairwise identity | 77.73 |
| Shannon entropy | 0.36965 |
| G+C content | 0.45178 |
| Mean single sequence MFE | -18.82 |
| Consensus MFE | -13.56 |
| Energy contribution | -12.64 |
| Covariance contribution | -0.92 |
| Combinations/Pair | 1.32 |
| Mean z-score | -2.08 |
| Structure conservation index | 0.72 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.79 |
| SVM RNA-class probability | 0.968057 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
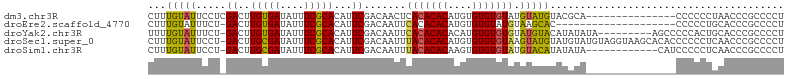
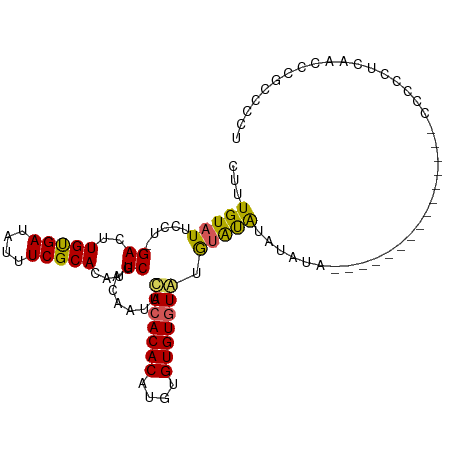
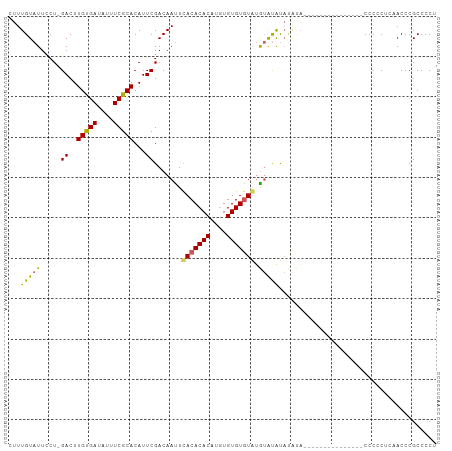
>dm3.chr3R 10580700 91 + 27905053 CUUUGUAUUCCUCGACUUGUGAUAUUUCGCACAUUCGACAACUCACACACAUGUGUGUGUAUGUAUGUACGCA---------------CCCCCCUAACCCGCCCCU ....((((...((((..(((((....)))))...))))..((..((((((....))))))..))..))))...---------------.................. ( -17.80, z-score = -2.03, R) >droEre2.scaffold_4770 10114426 85 + 17746568 CUUUGUAUUUCU-GACUUGUGAUAUUUCGCACAUUCGACAAUUCACACACAUGUGUGUAUGUAAGCAC--------------------CCCCCUGCACCCGCCCCU ...........(-((..(((((....)))))...)))(((...(((((....)))))..)))..(((.--------------------.....))).......... ( -14.00, z-score = -1.14, R) >droYak2.chr3R 12982671 96 - 28832112 UUUUGUAUUUCU-GACUUGUGAUAUUUCGCACAUUCGACAAUUCACACACACAUGUGUGUGUAUGUACAUAUAUA---------AGCCCCCACUGCACCCGCCCCU .(((((((...(-(...(((((....)))))......(((...(((((((....)))))))..))).)).)))))---------))........((....)).... ( -18.10, z-score = -1.44, R) >droSec1.super_0 9767603 105 - 21120651 CUUUGUAUUCCU-GACUUGCGAUAUUUCGCACAUUCGACAAUUUACACACAUGUGUGUGUAAGUAUGUAUGUAUGUAGGUAAGCACACCCCCCUCAACCCGCCCCU ...(((...(((-(...(((((....)))))(((.(((((((((((((((....)))))))))).))).)).)))))))...)))..................... ( -25.80, z-score = -2.55, R) >droSim1.chr3R 16637009 93 - 27517382 CUUUGUAUUCCU-GACUUGCGAUAUUUCGCACAUUCGACAAUUUACACACAAGUGUGUGUAUGUACAUAUAUA------------CAUCCCCCUCAACCCGCCCCU ...(((((...(-(((.(((((....)))))............(((((((....))))))).)).))...)))------------))................... ( -18.40, z-score = -3.22, R) >consensus CUUUGUAUUCCU_GACUUGUGAUAUUUCGCACAUUCGACAAUUCACACACAUGUGUGUGUAUGUAUAUAUAUA_______________CCCCCUCAACCCGCCCCU ...(((((.....((..(((((....)))))...)).......(((((((....))))))).)))))....................................... (-13.56 = -12.64 + -0.92)
| Location | 10,580,706 – 10,580,797 |
|---|---|
| Length | 91 |
| Sequences | 5 |
| Columns | 106 |
| Reading direction | forward |
| Mean pairwise identity | 78.14 |
| Shannon entropy | 0.36284 |
| G+C content | 0.46456 |
| Mean single sequence MFE | -21.96 |
| Consensus MFE | -17.36 |
| Energy contribution | -17.00 |
| Covariance contribution | -0.36 |
| Combinations/Pair | 1.20 |
| Mean z-score | -1.63 |
| Structure conservation index | 0.79 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.33 |
| SVM RNA-class probability | 0.927387 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
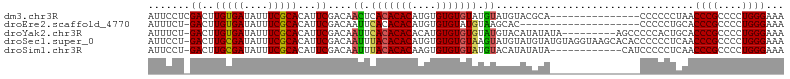
>dm3.chr3R 10580706 91 + 27905053 AUUCCUCGACUUGUGAUAUUUCGCACAUUCGACAACUCACACACAUGUGUGUGUAUGUAUGUACGCA---------------CCCCCCUAACCCGCCCCUGGGAAA .....((((..(((((....)))))...))))................((((((((....)))))))---------------)........((((....))))... ( -20.70, z-score = -1.45, R) >droEre2.scaffold_4770 10114432 85 + 17746568 AUUUCU-GACUUGUGAUAUUUCGCACAUUCGACAAUUCACACACAUGUGUGUAUGUAAGCAC--------------------CCCCCUGCACCCGCCCCUGGGAAA .....(-((..(((((....)))))...)))(((...(((((....)))))..)))..(((.--------------------.....))).((((....))))... ( -18.20, z-score = -1.02, R) >droYak2.chr3R 12982677 96 - 28832112 AUUUCU-GACUUGUGAUAUUUCGCACAUUCGACAAUUCACACACACAUGUGUGUGUAUGUACAUAUAUA---------AGCCCCCACUGCACCCGCCCCUGGGAAA .....(-((..(((((....)))))...)))(((...(((((((....)))))))..))).........---------....((((..((....))...))))... ( -22.50, z-score = -1.52, R) >droSec1.super_0 9767609 105 - 21120651 AUUCCU-GACUUGCGAUAUUUCGCACAUUCGACAAUUUACACACAUGUGUGUGUAAGUAUGUAUGUAUGUAGGUAAGCACACCCCCCUCAACCCGCCCCUGGGAAA ...(((-(...(((((....)))))(((.(((((((((((((((....)))))))))).))).)).)))))))..................((((....))))... ( -27.90, z-score = -1.89, R) >droSim1.chr3R 16637015 93 - 27517382 AUUCCU-GACUUGCGAUAUUUCGCACAUUCGACAAUUUACACACAAGUGUGUGUAUGUACAUAUAUA------------CAUCCCCCUCAACCCGCCCCUGGGAAA .(((((-((..(((((....)))))...))((.....(((((((....)))))))((((......))------------))......))...........))))). ( -20.50, z-score = -2.26, R) >consensus AUUCCU_GACUUGUGAUAUUUCGCACAUUCGACAAUUCACACACAUGUGUGUGUAUGUAUAUAUAUA_______________CCCCCUCAACCCGCCCCUGGGAAA .......((..(((((....)))))...))....((.(((((((....))))))).)).................................((((....))))... (-17.36 = -17.00 + -0.36)
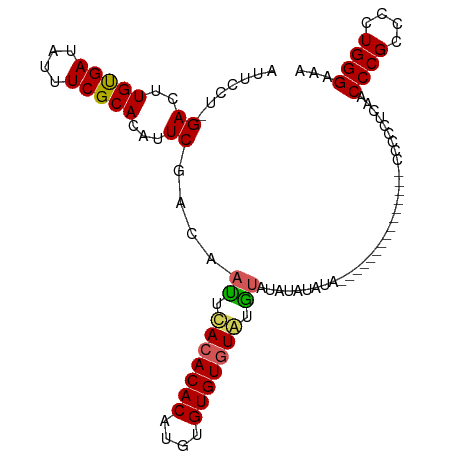
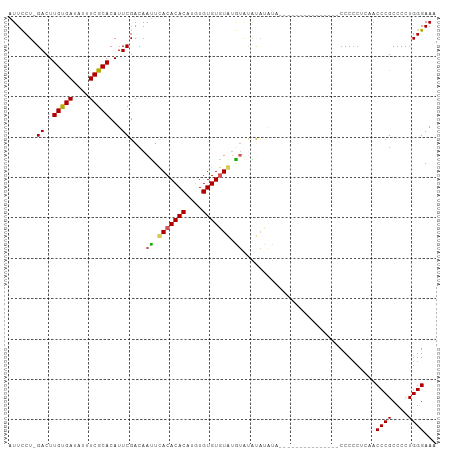
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:15:47 2011