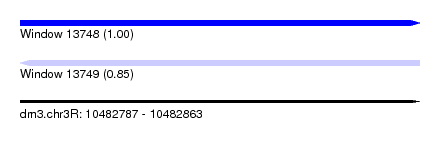

| Sequence ID | dm3.chr3R |
|---|---|
| Location | 10,482,787 – 10,482,863 |
| Length | 76 |
| Max. P | 0.995108 |

| Location | 10,482,787 – 10,482,863 |
|---|---|
| Length | 76 |
| Sequences | 3 |
| Columns | 87 |
| Reading direction | forward |
| Mean pairwise identity | 77.20 |
| Shannon entropy | 0.30321 |
| G+C content | 0.43618 |
| Mean single sequence MFE | -21.93 |
| Consensus MFE | -19.90 |
| Energy contribution | -20.68 |
| Covariance contribution | 0.78 |
| Combinations/Pair | 1.13 |
| Mean z-score | -2.56 |
| Structure conservation index | 0.91 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 2.77 |
| SVM RNA-class probability | 0.995108 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3R 10482787 76 + 27905053 CGGUCAUGGUUAAUUCACACCACAA-----------CUAAACGCCGAGAUGCCGAACAUAUGAGAGGCUCGACUUAAGAUGUUCGGC ((((..(((((............))-----------)))...))))....(((((((((....(((......)))...))))))))) ( -20.60, z-score = -1.93, R) >droSec1.super_0 9566333 76 - 21120651 CAGUCUUGGUUCAUAGAUACCAUAU-----------AUAAACGCUGAGAUGCUGAAUAUCUAAGAGGCUCGACUUUAGAUGUUCGGC ..((((..((......(((...)))-----------......))..))))((((((((((((.(((......))))))))))))))) ( -22.30, z-score = -2.59, R) >droSim1.chr3R 16537479 87 - 27517382 CUGUCUUUGUUCAUAGAUACCAUAUUCAUACCACAAAUAAACGCUGAGAUGCCGAACAUCUAACAGGCUCGACUUUAGAUGUUCGGC .(((((........)))))...............................(((((((((((((.((......))))))))))))))) ( -22.90, z-score = -3.15, R) >consensus CAGUCUUGGUUCAUAGAUACCAUAU___________AUAAACGCUGAGAUGCCGAACAUCUAAGAGGCUCGACUUUAGAUGUUCGGC ..((((((((................................))))))))(((((((((((((.((......))))))))))))))) (-19.90 = -20.68 + 0.78)


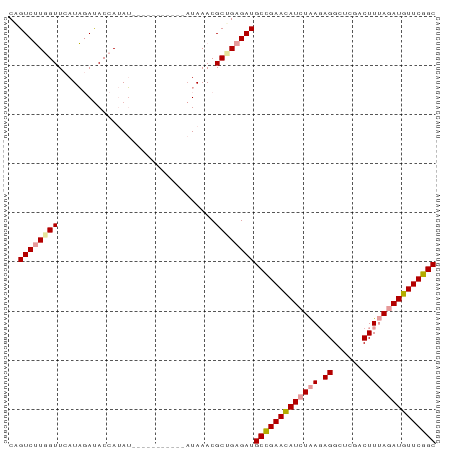
| Location | 10,482,787 – 10,482,863 |
|---|---|
| Length | 76 |
| Sequences | 3 |
| Columns | 87 |
| Reading direction | reverse |
| Mean pairwise identity | 77.20 |
| Shannon entropy | 0.30321 |
| G+C content | 0.43618 |
| Mean single sequence MFE | -20.63 |
| Consensus MFE | -15.27 |
| Energy contribution | -16.93 |
| Covariance contribution | 1.67 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.81 |
| Structure conservation index | 0.74 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.91 |
| SVM RNA-class probability | 0.851048 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3R 10482787 76 - 27905053 GCCGAACAUCUUAAGUCGAGCCUCUCAUAUGUUCGGCAUCUCGGCGUUUAG-----------UUGUGGUGUGAAUUAACCAUGACCG (((((((((....((........))...))))))))).............(-----------(..((((........))))..)).. ( -22.50, z-score = -2.15, R) >droSec1.super_0 9566333 76 + 21120651 GCCGAACAUCUAAAGUCGAGCCUCUUAGAUAUUCAGCAUCUCAGCGUUUAU-----------AUAUGGUAUCUAUGAACCAAGACUG ((.(((.((((((((......)).)))))).))).))......(.((((((-----------(.........))))))))....... ( -12.00, z-score = -0.02, R) >droSim1.chr3R 16537479 87 + 27517382 GCCGAACAUCUAAAGUCGAGCCUGUUAGAUGUUCGGCAUCUCAGCGUUUAUUUGUGGUAUGAAUAUGGUAUCUAUGAACAAAGACAG (((((((((((((((......)).)))))))))))))........((((((..(((.(((....))).)))..))))))........ ( -27.40, z-score = -3.28, R) >consensus GCCGAACAUCUAAAGUCGAGCCUCUUAGAUGUUCGGCAUCUCAGCGUUUAU___________AUAUGGUAUCUAUGAACCAAGACAG (((((((((((((((......)).)))))))))))))............................((((........))))...... (-15.27 = -16.93 + 1.67)


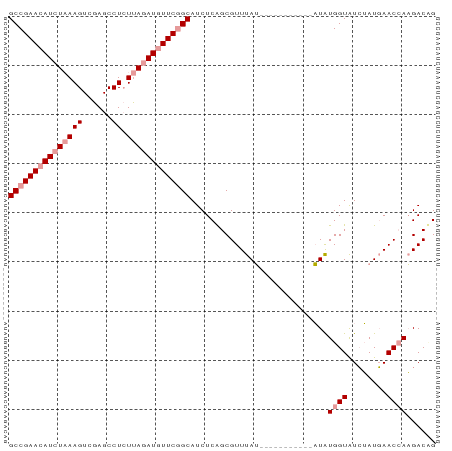
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:15:33 2011