
| Sequence ID | dm3.chr2LHet |
|---|---|
| Location | 36,615 – 36,721 |
| Length | 106 |
| Max. P | 0.515802 |

| Location | 36,615 – 36,721 |
|---|---|
| Length | 106 |
| Sequences | 3 |
| Columns | 106 |
| Reading direction | reverse |
| Mean pairwise identity | 60.69 |
| Shannon entropy | 0.56690 |
| G+C content | 0.44166 |
| Mean single sequence MFE | -27.07 |
| Consensus MFE | -13.61 |
| Energy contribution | -11.97 |
| Covariance contribution | -1.65 |
| Combinations/Pair | 1.62 |
| Mean z-score | -0.95 |
| Structure conservation index | 0.50 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.04 |
| SVM RNA-class probability | 0.515802 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2LHet 36615 106 - 368872 AGCACGGGGGAUCAAACAUCGUGGUAUGGGCGUGCUCUUCAAACUAUGGAGUAGGUCCGAUCCAUUGAAUUCAAGGCAUCAUGGAUCAGCACAUUUACACAGAUAU ..((((..(.......)..))))(((.((((.((((((.........)))))).))))(((((((.((..........)))))))))........)))........ ( -29.60, z-score = -1.70, R) >anoGam1.chr2L 616851 106 - 48795086 AACACAGAGGAGCAAAACUCAUGGUGUGGGGAUGUUUUUCGUACAAUGGUGUUGGUCCAAUUUAUCGCAUACCGGGUAUAAUGGAUCAACAUGGAUACAUAGCAAU ......((((((((...(((((...)))))..))))))))(((...(.(((((((((((...((((........))))...))))))))))).).)))........ ( -29.60, z-score = -1.95, R) >triCas2.ChLG4 653835 105 + 13894384 AACAUGGUCAUGGUAGCGUUUUUCUGUGGGGUUGUUUUUCGUGCAAUGGUGUGGGUCCGUUGAGCUUUAUUGAGGGAAU-AUGGAUCGGUUCAUCUAUAGGGACAU ..(((....)))........(..((((((((((((.......))))).(..(.(((((((....((((....))))...-))))))).)..).)))))))..)... ( -22.00, z-score = 0.79, R) >consensus AACACGGAGGAGCAAACAUCAUGGUGUGGGGAUGUUUUUCGUACAAUGGUGUAGGUCCGAUCAAUUGAAUUCAGGGAAU_AUGGAUCAGCACAUAUACACAGAAAU .......................(((((((.(((((......(((....))).((((((.....(((....))).......))))))))))).)))))))...... (-13.61 = -11.97 + -1.65)


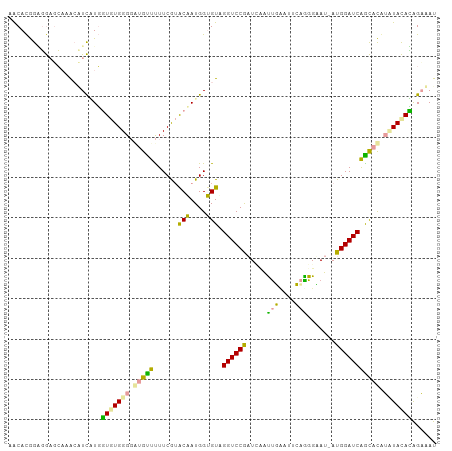
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:04:41 2011