
| Sequence ID | Eco_1 |
|---|---|
| Location | 4,035,569 – 4,035,769 |
| Length | 200 |
| Max. P | 0.999541 |

| Location | 4,035,569 – 4,035,689 |
|---|---|
| Length | 120 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 92.50 |
| Mean single sequence MFE | -42.00 |
| Consensus MFE | -43.54 |
| Energy contribution | -41.23 |
| Covariance contribution | -2.31 |
| Combinations/Pair | 1.24 |
| Mean z-score | -1.96 |
| Structure conservation index | 1.04 |
| SVM decision value | 1.98 |
| SVM RNA-class probability | 0.984654 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>Eco_1 4035569 120 + 1 AUGCCCUGGCAGUCAGAGGCGAUGAAGGACGUGCUAAUCUGCGAUAAGCGUCGGUAAGGUGAUAUGAACCGUUAUAACCGGCGAUUUCCGAAUGGGGAAACCCAGUGUGUUUCGACACAC .(((((((.....))).)))).....(((..(((......))).....(((((((..(((.......)))......)))))))...)))...((((....))))((((((....)))))) ( -40.80) >Sfl_1 41 120 + 1 AUGCCCUGGCAGUCAGAGGCGAUGAAGGACGUGCUAAUCUGCGAUAAGCGUCGGUAAGGUGAUAUGAACCGUUAUAACCGGCGAUUUCCGAAUGGGGAAACCCAGUGUGAUUCGUCACAC ..((((((.....))).)))(((((((((..(((......))).....(((((((..(((.......)))......)))))))...)))...((((....))))......)))))).... ( -41.10) >Sty_1 41 120 + 1 AUGCCCUGGCAGUCAGAGGCGAUGAAGGGCGUGCUAAUCUGCGAUAAGCGCCGGUAAGGUGAUAUGAACCGUUAUAACCGGCGAUACCCGAAUGGGGAAACCCAGUGUGAUUCGUCACAC ..((((((.....))).)))(((((((((..(((......))......(((((((..(((.......)))......))))))))..)))...((((....))))......)))))).... ( -45.40) >Ype_1 41 120 + 1 AUGCCUAGGCAGUCAGAGGCGAUGAAGGGCGUGCUAAUCUGCGAAAAGCGUCGGUAAGCUGAUAUGAAGCGUUAUAACCGACGAUACCCGAAUGGGGAAACCCAGUGCAAUACGUUGCAC .(((....))).......((((((..(((..(((......))......(((((((..(((.......)))......))))))))..)))(..((((....))))...)....)))))).. ( -40.70) >consensus AUGCCCUGGCAGUCAGAGGCGAUGAAGGACGUGCUAAUCUGCGAUAAGCGUCGGUAAGGUGAUAUGAACCGUUAUAACCGGCGAUACCCGAAUGGGGAAACCCAGUGUGAUUCGUCACAC .(((((((.....))).)))).....(((..(((......))).....(((((((..(((.......)))......)))))))...)))...((((....))))((((((....)))))) (-43.54 = -41.23 + -2.31)

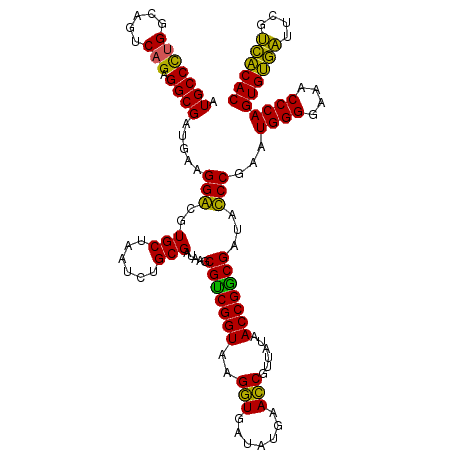

| Location | 4,035,569 – 4,035,689 |
|---|---|
| Length | 120 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 92.50 |
| Mean single sequence MFE | -35.25 |
| Consensus MFE | -33.44 |
| Energy contribution | -32.12 |
| Covariance contribution | -1.31 |
| Combinations/Pair | 1.22 |
| Mean z-score | -1.35 |
| Structure conservation index | 0.95 |
| SVM decision value | 0.20 |
| SVM RNA-class probability | 0.631277 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>Eco_1 4035569 120 - 1 GUGUGUCGAAACACACUGGGUUUCCCCAUUCGGAAAUCGCCGGUUAUAACGGUUCAUAUCACCUUACCGACGCUUAUCGCAGAUUAGCACGUCCUUCAUCGCCUCUGACUGCCAGGGCAU ((((((....))))))..(((((((......)))))))((..(((....((((............))))..((.....)).)))..))............(((.(((.....)))))).. ( -32.20) >Sfl_1 41 120 - 1 GUGUGACGAAUCACACUGGGUUUCCCCAUUCGGAAAUCGCCGGUUAUAACGGUUCAUAUCACCUUACCGACGCUUAUCGCAGAUUAGCACGUCCUUCAUCGCCUCUGACUGCCAGGGCAU ((((((....))))))..(((((((......)))))))((..(((....((((............))))..((.....)).)))..))............(((.(((.....)))))).. ( -32.30) >Sty_1 41 120 - 1 GUGUGACGAAUCACACUGGGUUUCCCCAUUCGGGUAUCGCCGGUUAUAACGGUUCAUAUCACCUUACCGGCGCUUAUCGCAGAUUAGCACGCCCUUCAUCGCCUCUGACUGCCAGGGCAU ((((((....))))))((((....))))...((((...((((((......(((.......)))..))))))(((.(((...))).)))..))))......(((.(((.....)))))).. ( -41.30) >Ype_1 41 120 - 1 GUGCAACGUAUUGCACUGGGUUUCCCCAUUCGGGUAUCGUCGGUUAUAACGCUUCAUAUCAGCUUACCGACGCUUUUCGCAGAUUAGCACGCCCUUCAUCGCCUCUGACUGCCUAGGCAU ((((((....))))))((((....))))...((((...((((((......(((.......)))..))))))(((..((...))..)))..))))......((((..(....)..)))).. ( -35.20) >consensus GUGUGACGAAUCACACUGGGUUUCCCCAUUCGGAAAUCGCCGGUUAUAACGGUUCAUAUCACCUUACCGACGCUUAUCGCAGAUUAGCACGCCCUUCAUCGCCUCUGACUGCCAGGGCAU ((((((....))))))..(((((((......)))))))((((((......(((.......)))..))))))((((...((((.((((...((........))..))))))))..)))).. (-33.44 = -32.12 + -1.31)

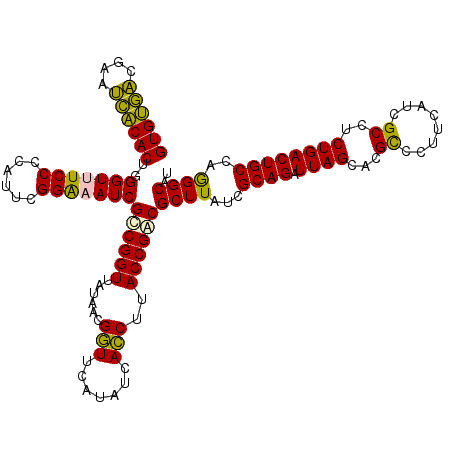

| Location | 4,035,609 – 4,035,729 |
|---|---|
| Length | 120 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 90.97 |
| Mean single sequence MFE | -43.30 |
| Consensus MFE | -46.45 |
| Energy contribution | -43.20 |
| Covariance contribution | -3.25 |
| Combinations/Pair | 1.29 |
| Mean z-score | -3.45 |
| Structure conservation index | 1.07 |
| SVM decision value | 3.70 |
| SVM RNA-class probability | 0.999541 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>Eco_1 4035609 120 + 1 GCGAUAAGCGUCGGUAAGGUGAUAUGAACCGUUAUAACCGGCGAUUUCCGAAUGGGGAAACCCAGUGUGUUUCGACACACUAUCAUUAACUGAAUCCAUAGGUUAAUGAGGCGAACCGGG ((.....))((((((..(((.......)))......)))))).....(((...(((....)))(((((((....))))))).(((((((((.........))))))))).......))). ( -40.60) >Sfl_1 81 120 + 1 GCGAUAAGCGUCGGUAAGGUGAUAUGAACCGUUAUAACCGGCGAUUUCCGAAUGGGGAAACCCAGUGUGAUUCGUCACACUAUCAUUAACUGAAUCCAUAGGUUAAUGAGGCGAACCGGG ((.....))((((((..(((.......)))......)))))).....(((...(((....)))(((((((....))))))).(((((((((.........))))))))).......))). ( -40.90) >Sty_1 81 120 + 1 GCGAUAAGCGCCGGUAAGGUGAUAUGAACCGUUAUAACCGGCGAUACCCGAAUGGGGAAACCCAGUGUGAUUCGUCACACUAUCAUUAACUGAAUCCAUAGGUUAAUGAGGCGAACCGGG ((.....))((((((..(((.......)))......))))))....((((...(((....)))(((((((....))))))).(((((((((.........))))))))).......)))) ( -46.00) >Ype_1 81 120 + 1 GCGAAAAGCGUCGGUAAGCUGAUAUGAAGCGUUAUAACCGACGAUACCCGAAUGGGGAAACCCAGUGCAAUACGUUGCACUAUCGUUAGAUGAAUACAUAGUCUAACGAGGCGAACCGGG ((.....))((((((..(((.......)))......))))))....((((...(((....)))(((((((....))))))).(((((((((.........))))))))).......)))) ( -45.70) >consensus GCGAUAAGCGUCGGUAAGGUGAUAUGAACCGUUAUAACCGGCGAUACCCGAAUGGGGAAACCCAGUGUGAUUCGUCACACUAUCAUUAACUGAAUCCAUAGGUUAAUGAGGCGAACCGGG ((.....))((((((..(((.......)))......))))))....((((...(((....)))(((((((....))))))).(((((((((.........))))))))).......)))) (-46.45 = -43.20 + -3.25)


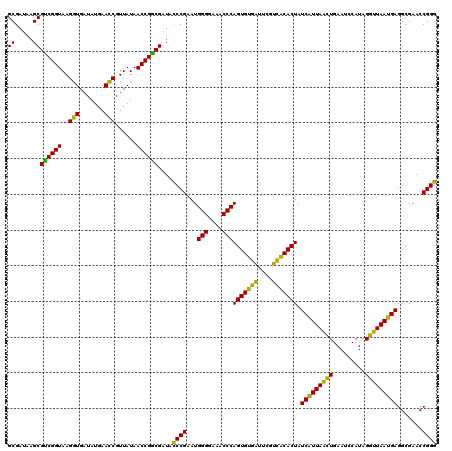
| Location | 4,035,609 – 4,035,729 |
|---|---|
| Length | 120 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 90.97 |
| Mean single sequence MFE | -34.78 |
| Consensus MFE | -34.20 |
| Energy contribution | -32.45 |
| Covariance contribution | -1.75 |
| Combinations/Pair | 1.24 |
| Mean z-score | -2.06 |
| Structure conservation index | 0.98 |
| SVM decision value | 2.02 |
| SVM RNA-class probability | 0.985750 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>Eco_1 4035609 120 - 1 CCCGGUUCGCCUCAUUAACCUAUGGAUUCAGUUAAUGAUAGUGUGUCGAAACACACUGGGUUUCCCCAUUCGGAAAUCGCCGGUUAUAACGGUUCAUAUCACCUUACCGACGCUUAUCGC .(((((.....((((((((...........))))))))((((((((....))))))))(((((((......))))))))))))......((((............))))........... ( -33.10) >Sfl_1 81 120 - 1 CCCGGUUCGCCUCAUUAACCUAUGGAUUCAGUUAAUGAUAGUGUGACGAAUCACACUGGGUUUCCCCAUUCGGAAAUCGCCGGUUAUAACGGUUCAUAUCACCUUACCGACGCUUAUCGC .(((((.....((((((((...........))))))))((((((((....))))))))(((((((......))))))))))))......((((............))))........... ( -33.20) >Sty_1 81 120 - 1 CCCGGUUCGCCUCAUUAACCUAUGGAUUCAGUUAAUGAUAGUGUGACGAAUCACACUGGGUUUCCCCAUUCGGGUAUCGCCGGUUAUAACGGUUCAUAUCACCUUACCGGCGCUUAUCGC (((((......((((((((...........)))))))).(((((((....)))))))(((....)))..)))))...(((((((......(((.......)))..)))))))........ ( -36.50) >Ype_1 81 120 - 1 CCCGGUUCGCCUCGUUAGACUAUGUAUUCAUCUAACGAUAGUGCAACGUAUUGCACUGGGUUUCCCCAUUCGGGUAUCGUCGGUUAUAACGCUUCAUAUCAGCUUACCGACGCUUUUCGC (((((......((((((((...((....)))))))))).(((((((....)))))))(((....)))..)))))...(((((((......(((.......)))..)))))))........ ( -36.30) >consensus CCCGGUUCGCCUCAUUAACCUAUGGAUUCAGUUAAUGAUAGUGUGACGAAUCACACUGGGUUUCCCCAUUCGGAAAUCGCCGGUUAUAACGGUUCAUAUCACCUUACCGACGCUUAUCGC ..((((..((.((((((((...........))))))))((((((((....))))))))(((((((......)))))))((((((......(((.......)))..))))))))..)))). (-34.20 = -32.45 + -1.75)

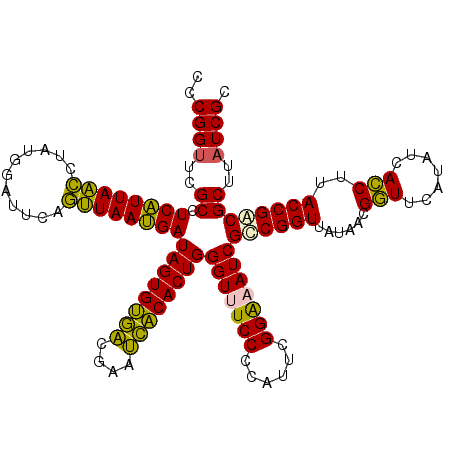
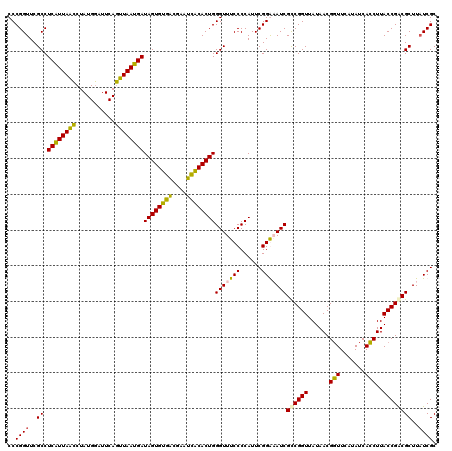
| Location | 4,035,649 – 4,035,769 |
|---|---|
| Length | 120 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 92.64 |
| Mean single sequence MFE | -41.62 |
| Consensus MFE | -42.53 |
| Energy contribution | -41.27 |
| Covariance contribution | -1.25 |
| Combinations/Pair | 1.17 |
| Mean z-score | -4.52 |
| Structure conservation index | 1.02 |
| SVM decision value | 2.72 |
| SVM RNA-class probability | 0.996602 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>Eco_1 4035649 120 + 1 GCGAUUUCCGAAUGGGGAAACCCAGUGUGUUUCGACACACUAUCAUUAACUGAAUCCAUAGGUUAAUGAGGCGAACCGGGGGAACUGAAACAUCUAAGUACCCCGAGGAAAAGAAAUCAA ..((((((((...(((....)))(((((((....))))))).(((((((((.........)))))))))..))..((.((((.(((..........))).))))..))....)))))).. ( -42.70) >Sfl_1 121 120 + 1 GCGAUUUCCGAAUGGGGAAACCCAGUGUGAUUCGUCACACUAUCAUUAACUGAAUCCAUAGGUUAAUGAGGCGAACCGGGGGAACUGAAACAUCUAAGUACCCCGAGGAAAAGAAAUCAA ..((((((((...(((....)))(((((((....))))))).(((((((((.........)))))))))..))..((.((((.(((..........))).))))..))....)))))).. ( -43.00) >Sty_1 121 120 + 1 GCGAUACCCGAAUGGGGAAACCCAGUGUGAUUCGUCACACUAUCAUUAACUGAAUCCAUAGGUUAAUGAGGCGAACCGGGGGAACUGAAACAUCUAAGUACCCCGAGGAAAAGAAAUCAA ..(((.(((....(((....)))(((((((....))))))).(((((((((.........))))))))))).)..((.((((.(((..........))).))))..)).......))).. ( -39.50) >Ype_1 121 120 + 1 ACGAUACCCGAAUGGGGAAACCCAGUGCAAUACGUUGCACUAUCGUUAGAUGAAUACAUAGUCUAACGAGGCGAACCGGGGGAACUGAAACAUCUAAGUACCCCGAGGAAAAGAAAUCAA ..(((.(((....(((....)))(((((((....))))))).(((((((((.........))))))))))).)..((.((((.(((..........))).))))..)).......))).. ( -41.30) >consensus GCGAUACCCGAAUGGGGAAACCCAGUGUGAUUCGUCACACUAUCAUUAACUGAAUCCAUAGGUUAAUGAGGCGAACCGGGGGAACUGAAACAUCUAAGUACCCCGAGGAAAAGAAAUCAA ..((((((((...(((....)))(((((((....))))))).(((((((((.........)))))))))..))..((.((((.(((..........))).))))..))....)))))).. (-42.53 = -41.27 + -1.25)


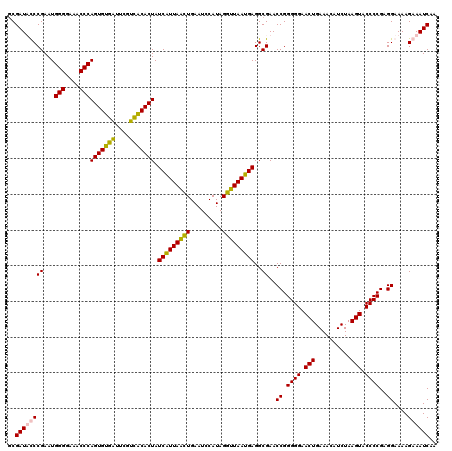
| Location | 4,035,649 – 4,035,769 |
|---|---|
| Length | 120 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 92.64 |
| Mean single sequence MFE | -33.25 |
| Consensus MFE | -35.83 |
| Energy contribution | -33.82 |
| Covariance contribution | -2.00 |
| Combinations/Pair | 1.19 |
| Mean z-score | -1.81 |
| Structure conservation index | 1.08 |
| SVM decision value | 1.79 |
| SVM RNA-class probability | 0.977469 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>Eco_1 4035649 120 - 1 UUGAUUUCUUUUCCUCGGGGUACUUAGAUGUUUCAGUUCCCCCGGUUCGCCUCAUUAACCUAUGGAUUCAGUUAAUGAUAGUGUGUCGAAACACACUGGGUUUCCCCAUUCGGAAAUCGC ..(((((((.......((((.(((..((....))))).)))).((...(((((((((((...........))))))))((((((((....)))))))))))..))......))))))).. ( -34.70) >Sfl_1 121 120 - 1 UUGAUUUCUUUUCCUCGGGGUACUUAGAUGUUUCAGUUCCCCCGGUUCGCCUCAUUAACCUAUGGAUUCAGUUAAUGAUAGUGUGACGAAUCACACUGGGUUUCCCCAUUCGGAAAUCGC ..(((((((.......((((.(((..((....))))).)))).((...(((((((((((...........))))))))((((((((....)))))))))))..))......))))))).. ( -34.80) >Sty_1 121 120 - 1 UUGAUUUCUUUUCCUCGGGGUACUUAGAUGUUUCAGUUCCCCCGGUUCGCCUCAUUAACCUAUGGAUUCAGUUAAUGAUAGUGUGACGAAUCACACUGGGUUUCCCCAUUCGGGUAUCGC ..............((((((.(((..((....)))))..))))))...(((((((((((...........))))))))((((((((....)))))))))))...(((....)))...... ( -30.80) >Ype_1 121 120 - 1 UUGAUUUCUUUUCCUCGGGGUACUUAGAUGUUUCAGUUCCCCCGGUUCGCCUCGUUAGACUAUGUAUUCAUCUAACGAUAGUGCAACGUAUUGCACUGGGUUUCCCCAUUCGGGUAUCGU ..............((((((.(((..((....)))))..))))))...(((((((((((...((....))))))))))((((((((....)))))))))))...(((....)))...... ( -32.70) >consensus UUGAUUUCUUUUCCUCGGGGUACUUAGAUGUUUCAGUUCCCCCGGUUCGCCUCAUUAACCUAUGGAUUCAGUUAAUGAUAGUGUGACGAAUCACACUGGGUUUCCCCAUUCGGAAAUCGC ..(((((((.......((((.(((..((....))))).)))).((...(((((((((((...........))))))))((((((((....)))))))))))..))......))))))).. (-35.83 = -33.82 + -2.00)

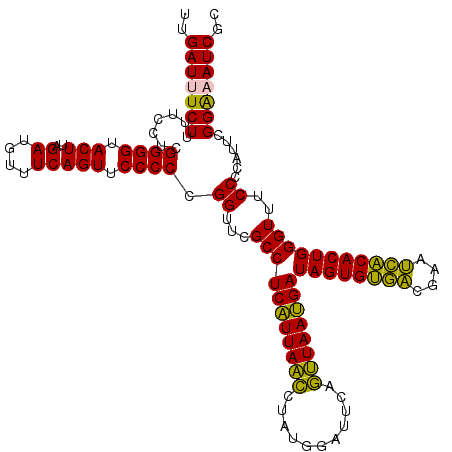
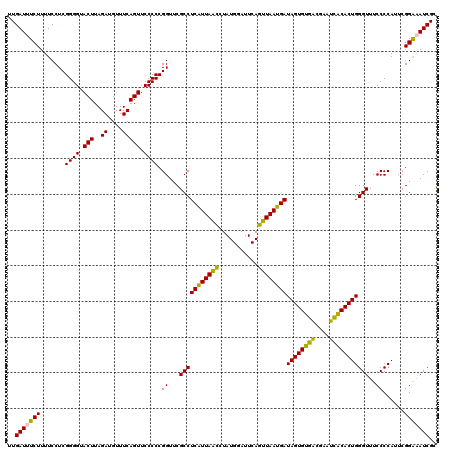
Generated by rnazCluster.pl (part of RNAz 1.0) on Fri Dec 1 11:06:31 2006