
| Sequence ID | Eco_1 |
|---|---|
| Location | 3,939,685 – 3,940,285 |
| Length | 600 |
| Max. P | 0.994689 |

| Location | 3,939,685 – 3,939,805 |
|---|---|
| Length | 120 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 98.33 |
| Mean single sequence MFE | -30.13 |
| Consensus MFE | -30.13 |
| Energy contribution | -30.13 |
| Covariance contribution | 0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.39 |
| Structure conservation index | 1.00 |
| SVM decision value | 1.65 |
| SVM RNA-class probability | 0.969714 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
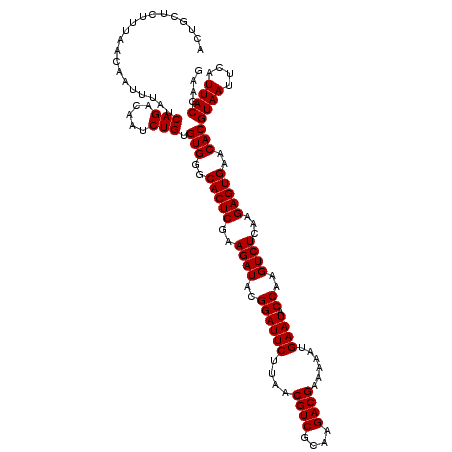
>Eco_1 3939685 120 + 1 ACUGCUCUUUAACAAUUUAUCAGACAAUCUGUGUGGGCACUCGAAGAUACGGAUUCUUAACGUCGCAAGACGAAAAAUGAAUACCAAGUCUCAAGAGUGAACACGUAAUUCAUUACGAAG ....................(((.....))).(((..(((((..((((..((((((....((((....))))......)))).))..))))...)))))..)))((((....)))).... ( -30.90) >Sfl_1 1 120 + 1 ACUGCUCUUUAACAAUUUAUCAGACAAUCUGUGUGGGCACUCGAAGAUACGGAUUCUUAACGUCGCAAGACGAAAAAUGAAUACCAAGUCUCAAGAGUGAACACGUAAUUCAUUACGAAG ....................(((.....))).(((..(((((..((((..((((((....((((....))))......)))).))..))))...)))))..)))((((....)))).... ( -30.90) >Sty_1 1 120 + 1 ACUGCUCUUUAACAAUUUAUCAGACAAUCUGUGUGGGCACUCGAAGAUACGGAUUCUUAACGUCCUAGGACGAAAAAUGAAUACCAAGUCUCAAGAGUGAACACGUAAUUCAUUACGAAG ....................(((.....))).(((..(((((..((((..((((((....((((....))))......)))).))..))))...)))))..)))((((....)))).... ( -28.60) >consensus ACUGCUCUUUAACAAUUUAUCAGACAAUCUGUGUGGGCACUCGAAGAUACGGAUUCUUAACGUCGCAAGACGAAAAAUGAAUACCAAGUCUCAAGAGUGAACACGUAAUUCAUUACGAAG ....................(((.....))).(((..(((((..((((..((((((....((((....))))......)))).))..))))...)))))..)))((((....)))).... (-30.13 = -30.13 + 0.00)



| Location | 3,939,685 – 3,939,805 |
|---|---|
| Length | 120 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 98.33 |
| Mean single sequence MFE | -31.00 |
| Consensus MFE | -31.00 |
| Energy contribution | -31.00 |
| Covariance contribution | 0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.80 |
| Structure conservation index | 1.00 |
| SVM decision value | 0.77 |
| SVM RNA-class probability | 0.845854 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>Eco_1 3939685 120 - 1 CUUCGUAAUGAAUUACGUGUUCACUCUUGAGACUUGGUAUUCAUUUUUCGUCUUGCGACGUUAAGAAUCCGUAUCUUCGAGUGCCCACACAGAUUGUCUGAUAAAUUGUUAAAGAGCAGU (((..(((..(.(((.(((..(((((..((((..(((.((((.((...((((....))))..)))))))))..)))).)))))..))).(((.....))).))).)..)))..))).... ( -30.50) >Sfl_1 1 120 - 1 CUUCGUAAUGAAUUACGUGUUCACUCUUGAGACUUGGUAUUCAUUUUUCGUCUUGCGACGUUAAGAAUCCGUAUCUUCGAGUGCCCACACAGAUUGUCUGAUAAAUUGUUAAAGAGCAGU (((..(((..(.(((.(((..(((((..((((..(((.((((.((...((((....))))..)))))))))..)))).)))))..))).(((.....))).))).)..)))..))).... ( -30.50) >Sty_1 1 120 - 1 CUUCGUAAUGAAUUACGUGUUCACUCUUGAGACUUGGUAUUCAUUUUUCGUCCUAGGACGUUAAGAAUCCGUAUCUUCGAGUGCCCACACAGAUUGUCUGAUAAAUUGUUAAAGAGCAGU (((..(((..(.(((.(((..(((((..((((..(((.((((.((...((((....))))..)))))))))..)))).)))))..))).(((.....))).))).)..)))..))).... ( -32.00) >consensus CUUCGUAAUGAAUUACGUGUUCACUCUUGAGACUUGGUAUUCAUUUUUCGUCUUGCGACGUUAAGAAUCCGUAUCUUCGAGUGCCCACACAGAUUGUCUGAUAAAUUGUUAAAGAGCAGU (((..(((..(.(((.(((..(((((..((((..(((.((((.((...((((....))))..)))))))))..)))).)))))..))).(((.....))).))).)..)))..))).... (-31.00 = -31.00 + 0.00)


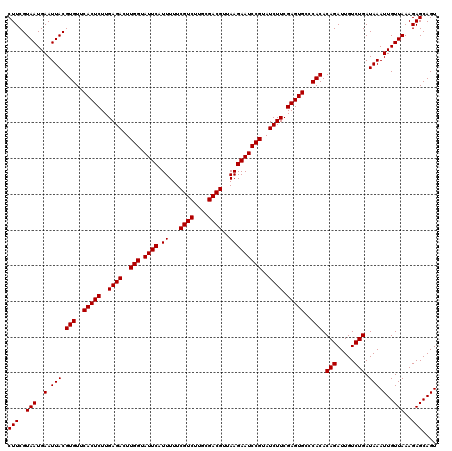
| Location | 3,939,725 – 3,939,845 |
|---|---|
| Length | 120 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 97.78 |
| Mean single sequence MFE | -29.03 |
| Consensus MFE | -29.03 |
| Energy contribution | -29.03 |
| Covariance contribution | 0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.86 |
| Structure conservation index | 1.00 |
| SVM decision value | 2.19 |
| SVM RNA-class probability | 0.989904 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>Eco_1 3939725 120 + 1 UCGAAGAUACGGAUUCUUAACGUCGCAAGACGAAAAAUGAAUACCAAGUCUCAAGAGUGAACACGUAAUUCAUUACGAAGUUUAAUUCUUUGAGCAUCAAACUUUUAAAUUGAAGAGUUU .....(((..((((((....((((....))))......)))).))..(.(((((((((((((.(((((....)))))..)))).)))).)))))))))((((((((......)))))))) ( -29.80) >Sfl_1 41 120 + 1 UCGAAGAUACGGAUUCUUAACGUCGCAAGACGAAAAAUGAAUACCAAGUCUCAAGAGUGAACACGUAAUUCAUUACGAAGUUUAAUUCUUUGAGCAUCAAACUUUUAAAUUGAAGAGUUU .....(((..((((((....((((....))))......)))).))..(.(((((((((((((.(((((....)))))..)))).)))).)))))))))((((((((......)))))))) ( -29.80) >Sty_1 41 120 + 1 UCGAAGAUACGGAUUCUUAACGUCCUAGGACGAAAAAUGAAUACCAAGUCUCAAGAGUGAACACGUAAUUCAUUACGAAGUUUAAUUCAUUGAGCAUCAAACUUUUAAAUUGAAGAGUUU .....(((..((((((....((((....))))......)))).))..(.(((((((((((((.(((((....)))))..)))).)))).)))))))))((((((((......)))))))) ( -27.50) >consensus UCGAAGAUACGGAUUCUUAACGUCGCAAGACGAAAAAUGAAUACCAAGUCUCAAGAGUGAACACGUAAUUCAUUACGAAGUUUAAUUCUUUGAGCAUCAAACUUUUAAAUUGAAGAGUUU .....(((..((((((....((((....))))......)))).))..(.(((((((((((((.(((((....)))))..)))).)))).)))))))))((((((((......)))))))) (-29.03 = -29.03 + 0.00)

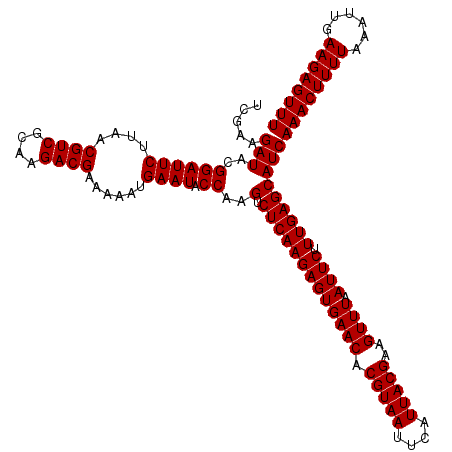

| Location | 3,939,765 – 3,939,885 |
|---|---|
| Length | 120 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 99.44 |
| Mean single sequence MFE | -31.37 |
| Consensus MFE | -30.70 |
| Energy contribution | -30.70 |
| Covariance contribution | -0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.20 |
| Structure conservation index | 0.98 |
| SVM decision value | 1.18 |
| SVM RNA-class probability | 0.926488 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>Eco_1 3939765 120 + 1 AUACCAAGUCUCAAGAGUGAACACGUAAUUCAUUACGAAGUUUAAUUCUUUGAGCAUCAAACUUUUAAAUUGAAGAGUUUGAUCAUGGCUCAGAUUGAACGCUGGCGGCAGGCCUAACAC ...(((.((.((((((((((((.(((((....)))))..)))).))))((((((((((((((((((......)))))))))))....)))))))))))))..))).((....))...... ( -31.50) >Sfl_1 81 120 + 1 AUACCAAGUCUCAAGAGUGAACACGUAAUUCAUUACGAAGUUUAAUUCUUUGAGCAUCAAACUUUUAAAUUGAAGAGUUUGAUCAUGGCUCAGAUUGAACGCUGGCGGCAGGCCUAACAC ...(((.((.((((((((((((.(((((....)))))..)))).))))((((((((((((((((((......)))))))))))....)))))))))))))..))).((....))...... ( -31.50) >Sty_1 81 120 + 1 AUACCAAGUCUCAAGAGUGAACACGUAAUUCAUUACGAAGUUUAAUUCAUUGAGCAUCAAACUUUUAAAUUGAAGAGUUUGAUCAUGGCUCAGAUUGAACGCUGGCGGCAGGCCUAACAC ...(((.((.((((((((((((.(((((....)))))..)))).)))).(((((((((((((((((......)))))))))))....)))))).))))))..))).((....))...... ( -31.10) >consensus AUACCAAGUCUCAAGAGUGAACACGUAAUUCAUUACGAAGUUUAAUUCUUUGAGCAUCAAACUUUUAAAUUGAAGAGUUUGAUCAUGGCUCAGAUUGAACGCUGGCGGCAGGCCUAACAC ...(((.((.((((((((((((.(((((....)))))..)))).)))).(((((((((((((((((......)))))))))))....)))))).))))))..))).((....))...... (-30.70 = -30.70 + -0.00)

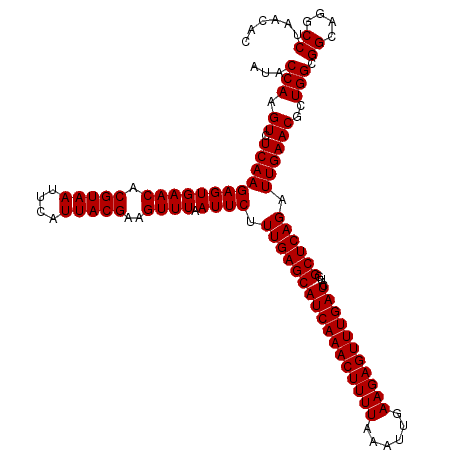
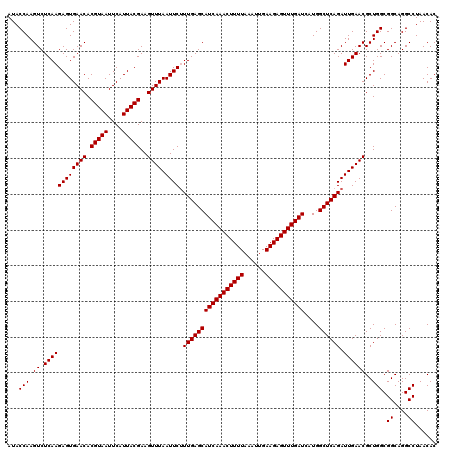
| Location | 3,939,805 – 3,939,925 |
|---|---|
| Length | 120 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 98.33 |
| Mean single sequence MFE | -36.57 |
| Consensus MFE | -36.94 |
| Energy contribution | -36.50 |
| Covariance contribution | -0.44 |
| Combinations/Pair | 1.05 |
| Mean z-score | -2.25 |
| Structure conservation index | 1.01 |
| SVM decision value | 1.55 |
| SVM RNA-class probability | 0.963107 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>Eco_1 3939805 120 + 1 UUUAAUUCUUUGAGCAUCAAACUUUUAAAUUGAAGAGUUUGAUCAUGGCUCAGAUUGAACGCUGGCGGCAGGCCUAACACAUGCAAGUCGAACGGUAACAGGAAACAGCUUGCUGUUUCG (((((((...((((((((((((((((......)))))))))))....)))))))))))).((((.((((..((.........))..))))..)))).....(((((((....))))))). ( -35.70) >Sfl_1 121 120 + 1 UUUAAUUCUUUGAGCAUCAAACUUUUAAAUUGAAGAGUUUGAUCAUGGCUCAGAUUGAACGCUGGCGGCAGGCCUAACACAUGCAAGUCGAACGGUAACAGGAAGCAGCUUGCUGUUUCG (((((((...((((((((((((((((......)))))))))))....)))))))))))).((((.((((..((.........))..))))..)))).....(((((((....))))))). ( -35.70) >Sty_1 121 120 + 1 UUUAAUUCAUUGAGCAUCAAACUUUUAAAUUGAAGAGUUUGAUCAUGGCUCAGAUUGAACGCUGGCGGCAGGCCUAACACAUGCAAGUCGAACGGUAACAGGAAGCAGCUUGCUGCUUCG .....(((((((((((((((((((((......)))))))))))....))))))..)))).((((.((((..((.........))..))))..)))).....(((((((....))))))). ( -38.30) >consensus UUUAAUUCUUUGAGCAUCAAACUUUUAAAUUGAAGAGUUUGAUCAUGGCUCAGAUUGAACGCUGGCGGCAGGCCUAACACAUGCAAGUCGAACGGUAACAGGAAGCAGCUUGCUGUUUCG (((((((...((((((((((((((((......)))))))))))....)))))))))))).((((.((((..((.........))..))))..)))).....(((((((....))))))). (-36.94 = -36.50 + -0.44)

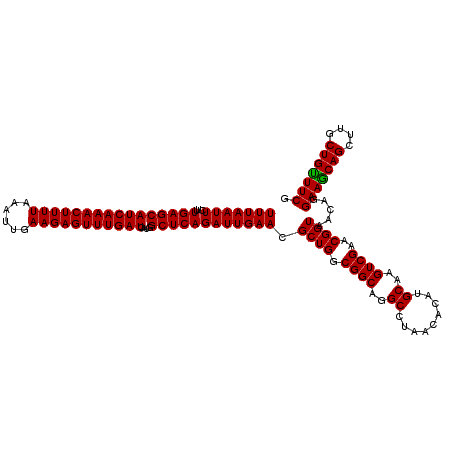

| Location | 3,939,885 – 3,940,005 |
|---|---|
| Length | 120 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 98.33 |
| Mean single sequence MFE | -37.60 |
| Consensus MFE | -38.26 |
| Energy contribution | -37.60 |
| Covariance contribution | -0.66 |
| Combinations/Pair | 1.08 |
| Mean z-score | -1.60 |
| Structure conservation index | 1.02 |
| SVM decision value | 0.55 |
| SVM RNA-class probability | 0.777989 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>Eco_1 3939885 120 + 1 AUGCAAGUCGAACGGUAACAGGAAACAGCUUGCUGUUUCGCUGACGAGUGGCGGACGGGUGAGUAAUGUCUGGGAAACUGCCUGAUGGAGGGGGAUAACUACUGGAAACGGUAGCUAAUA .(((.(.(((..((.((.((((((((((....))))))).))).....)).))..))).)..))).(((((((....)).(((......)))))))).((((((....))))))...... ( -36.90) >Sfl_1 201 120 + 1 AUGCAAGUCGAACGGUAACAGGAAGCAGCUUGCUGUUUCGCUGACGAGUGGCGGACGGGUGAGUAAUGUCUGGGAAACUGCCUGAUGGAGGGGGAUAACUACUGGAAACGGUAGCUAAUA .(((.(.(((..((.((.((((((((((....))))))).))).....)).))..))).)..))).(((((((....)).(((......)))))))).((((((....))))))...... ( -36.90) >Sty_1 201 120 + 1 AUGCAAGUCGAACGGUAACAGGAAGCAGCUUGCUGCUUCGCUGACGAGUGGCGGACGGGUGAGUAAUGUCUGGGAAACUGCCUGAUGGAGGGGGAUAACUACUGGAAACGGUGGCUAAUA .(((.(.(((..((.((.((((((((((....))))))).))).....)).))..))).)..))).(((((((....)).(((......)))))))).((((((....))))))...... ( -39.00) >consensus AUGCAAGUCGAACGGUAACAGGAAGCAGCUUGCUGUUUCGCUGACGAGUGGCGGACGGGUGAGUAAUGUCUGGGAAACUGCCUGAUGGAGGGGGAUAACUACUGGAAACGGUAGCUAAUA .(((.(.(((..((.((.((((((((((....))))))).))).....)).))..))).)..))).(((((((....)).(((......)))))))).((((((....))))))...... (-38.26 = -37.60 + -0.66)


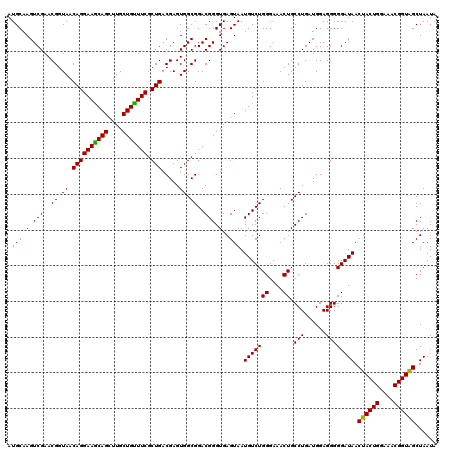
| Location | 3,939,925 – 3,940,045 |
|---|---|
| Length | 120 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 99.44 |
| Mean single sequence MFE | -41.00 |
| Consensus MFE | -41.22 |
| Energy contribution | -41.00 |
| Covariance contribution | -0.22 |
| Combinations/Pair | 1.03 |
| Mean z-score | -1.57 |
| Structure conservation index | 1.01 |
| SVM decision value | 0.40 |
| SVM RNA-class probability | 0.723224 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>Eco_1 3939925 120 + 1 CUGACGAGUGGCGGACGGGUGAGUAAUGUCUGGGAAACUGCCUGAUGGAGGGGGAUAACUACUGGAAACGGUAGCUAAUACCGCAUAACGUCGCAAGACCAAAGAGGGGGACCUUCGGGC .......((((((.....(((.(((.(((((((....)).(((......)))))))).((((((....))))))....))))))....))))))....((...((((....))))..)). ( -41.10) >Sfl_1 241 120 + 1 CUGACGAGUGGCGGACGGGUGAGUAAUGUCUGGGAAACUGCCUGAUGGAGGGGGAUAACUACUGGAAACGGUAGCUAAUACCGCAUAACGUCGCAAGACCAAAGAGGGGGACCUUCGGGC .......((((((.....(((.(((.(((((((....)).(((......)))))))).((((((....))))))....))))))....))))))....((...((((....))))..)). ( -41.10) >Sty_1 241 120 + 1 CUGACGAGUGGCGGACGGGUGAGUAAUGUCUGGGAAACUGCCUGAUGGAGGGGGAUAACUACUGGAAACGGUGGCUAAUACCGCAUAACGUCGCAAGACCAAAGAGGGGGACCUUCGGGC .......((((((.....(((.(((.(((((((....)).(((......)))))))).((((((....))))))....))))))....))))))....((...((((....))))..)). ( -40.80) >consensus CUGACGAGUGGCGGACGGGUGAGUAAUGUCUGGGAAACUGCCUGAUGGAGGGGGAUAACUACUGGAAACGGUAGCUAAUACCGCAUAACGUCGCAAGACCAAAGAGGGGGACCUUCGGGC .......((((((.....(((.(((.(((((((....)).(((......)))))))).((((((....))))))....))))))....))))))....((...((((....))))..)). (-41.22 = -41.00 + -0.22)


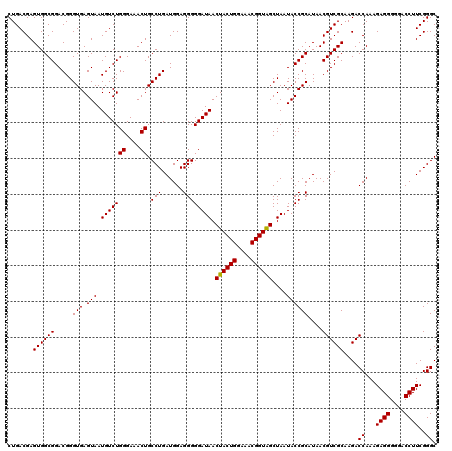
| Location | 3,939,965 – 3,940,085 |
|---|---|
| Length | 120 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 97.78 |
| Mean single sequence MFE | -48.67 |
| Consensus MFE | -49.11 |
| Energy contribution | -48.67 |
| Covariance contribution | -0.44 |
| Combinations/Pair | 1.06 |
| Mean z-score | -3.32 |
| Structure conservation index | 1.01 |
| SVM decision value | 2.50 |
| SVM RNA-class probability | 0.994689 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>Eco_1 3939965 120 + 1 CCUGAUGGAGGGGGAUAACUACUGGAAACGGUAGCUAAUACCGCAUAACGUCGCAAGACCAAAGAGGGGGACCUUCGGGCCUCUUGCCAUCAGAUGUGCCCAGAUGGGAUUAGCUAGUAG .(((((((((((((....((((((....))))))......(((......(((....)))....((((....))))))).)))))).)))))))..((.(((....)))....))...... ( -49.00) >Sfl_1 281 120 + 1 CCUGAUGGAGGGGGAUAACUACUGGAAACGGUAGCUAAUACCGCAUAACGUCGCAAGACCAAAGAGGGGGACCUUCGGGCCUCUUGCCAUCGGAUGUGCCCAGAUGGGAUUAGCUAGUAG .(((((((((((((....((((((....))))))......(((......(((....)))....((((....))))))).)))))).)))))))..((.(((....)))....))...... ( -48.30) >Sty_1 281 120 + 1 CCUGAUGGAGGGGGAUAACUACUGGAAACGGUGGCUAAUACCGCAUAACGUCGCAAGACCAAAGAGGGGGACCUUCGGGCCUCUUGCCAUCAGAUGUGCCCAGAUGGGAUUAGCUUGUUG .(((((((((((((....((((((....))))))......(((......(((....)))....((((....))))))).)))))).)))))))..((.(((....)))....))...... ( -48.70) >consensus CCUGAUGGAGGGGGAUAACUACUGGAAACGGUAGCUAAUACCGCAUAACGUCGCAAGACCAAAGAGGGGGACCUUCGGGCCUCUUGCCAUCAGAUGUGCCCAGAUGGGAUUAGCUAGUAG .(((((((((((((....((((((....))))))......(((......(((....)))....((((....))))))).)))))).)))))))..((.(((....)))....))...... (-49.11 = -48.67 + -0.44)

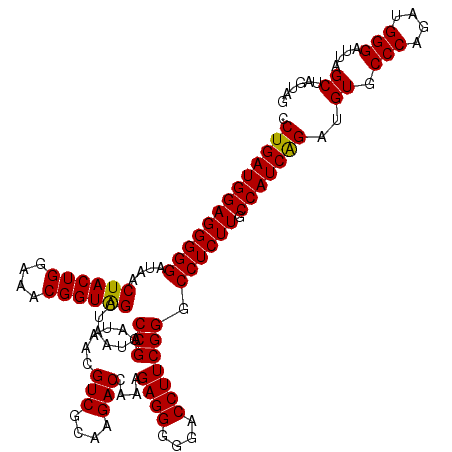

| Location | 3,940,005 – 3,940,125 |
|---|---|
| Length | 120 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 97.22 |
| Mean single sequence MFE | -52.00 |
| Consensus MFE | -50.93 |
| Energy contribution | -50.27 |
| Covariance contribution | -0.66 |
| Combinations/Pair | 1.05 |
| Mean z-score | -2.73 |
| Structure conservation index | 0.98 |
| SVM decision value | 1.93 |
| SVM RNA-class probability | 0.982911 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>Eco_1 3940005 120 + 1 CCGCAUAACGUCGCAAGACCAAAGAGGGGGACCUUCGGGCCUCUUGCCAUCAGAUGUGCCCAGAUGGGAUUAGCUAGUAGGUGGGGUAACGGCUCACCUAGGCGACGAUCCCUAGCUGGU ..(((((.((((....))).....((((((.((....)))))))).......).)))))((((..((((((.(((..((((((((.......)))))))))))...))))))...)))). ( -51.60) >Sfl_1 321 120 + 1 CCGCAUAACGUCGCAAGACCAAAGAGGGGGACCUUCGGGCCUCUUGCCAUCGGAUGUGCCCAGAUGGGAUUAGCUAGUAGGUGGGGUAACGGCUCACCUAGGCGACGAUCCCUAGCUGGU ..(((((.((..((((((((...((((....))))..))..))))))...))..)))))((((..((((((.(((..((((((((.......)))))))))))...))))))...)))). ( -52.30) >Sty_1 321 120 + 1 CCGCAUAACGUCGCAAGACCAAAGAGGGGGACCUUCGGGCCUCUUGCCAUCAGAUGUGCCCAGAUGGGAUUAGCUUGUUGGUGAGGUAACGGCUCACCAAGGCGACGAUCCCUAGCUGGU ..(((((.((((....))).....((((((.((....)))))))).......).)))))((((..((((((.(((..((((((((.......)))))))))))...))))))...)))). ( -52.10) >consensus CCGCAUAACGUCGCAAGACCAAAGAGGGGGACCUUCGGGCCUCUUGCCAUCAGAUGUGCCCAGAUGGGAUUAGCUAGUAGGUGGGGUAACGGCUCACCUAGGCGACGAUCCCUAGCUGGU .(((((...((.((((((((...((((....))))..))..)))))).))...))))).((((..((((((.(((..((((((((.......)))))))))))...))))))...)))). (-50.93 = -50.27 + -0.66)


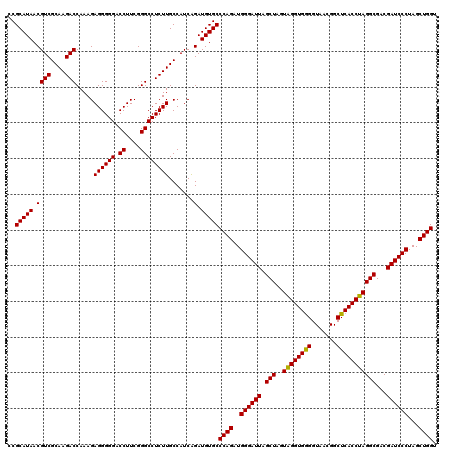
| Location | 3,940,085 – 3,940,205 |
|---|---|
| Length | 120 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 98.89 |
| Mean single sequence MFE | -47.53 |
| Consensus MFE | -44.72 |
| Energy contribution | -47.00 |
| Covariance contribution | 2.28 |
| Combinations/Pair | 1.03 |
| Mean z-score | -2.25 |
| Structure conservation index | 0.94 |
| SVM decision value | 0.94 |
| SVM RNA-class probability | 0.886151 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>Eco_1 3940085 120 + 1 GUGGGGUAACGGCUCACCUAGGCGACGAUCCCUAGCUGGUCUGAGAGGAUGACCAGCCACACUGGAACUGAGACACGGUCCAGACUCCUACGGGAGGCAGCAGUGGGGAAUAUUGCACAA (((.((((.((.((.....)).))....((((..(((((((.........)))))))(((.(((((.(((.....)))))))).((((....))))......))))))).)))).))).. ( -47.50) >Sfl_1 401 120 + 1 GUGGGGUAACGGCUCACCUAGGCGACGAUCCCUAGCUGGUCUGAGAGGAUGACCAGCCACACUGGAACUGAGACACGGUCCAGACUCCUACGGGAGGCAGCAGUGGGGAAUAUUGCACAA (((.((((.((.((.....)).))....((((..(((((((.........)))))))(((.(((((.(((.....)))))))).((((....))))......))))))).)))).))).. ( -47.50) >Sty_1 401 120 + 1 GUGAGGUAACGGCUCACCAAGGCGACGAUCCCUAGCUGGUCUGAGAGGAUGACCAGCCACACUGGAACUGAGACACGGUCCAGACUCCUACGGGAGGCAGCAGUGGGGAAUAUUGCACAA (((((.......)))))....((((.(.((((..(((((((.........)))))))(((.(((((.(((.....)))))))).((((....))))......))))))).).)))).... ( -47.60) >consensus GUGGGGUAACGGCUCACCUAGGCGACGAUCCCUAGCUGGUCUGAGAGGAUGACCAGCCACACUGGAACUGAGACACGGUCCAGACUCCUACGGGAGGCAGCAGUGGGGAAUAUUGCACAA (((((.......)))))....((((.(.((((..(((((((.........)))))))(((.(((((.(((.....)))))))).((((....))))......))))))).).)))).... (-44.72 = -47.00 + 2.28)

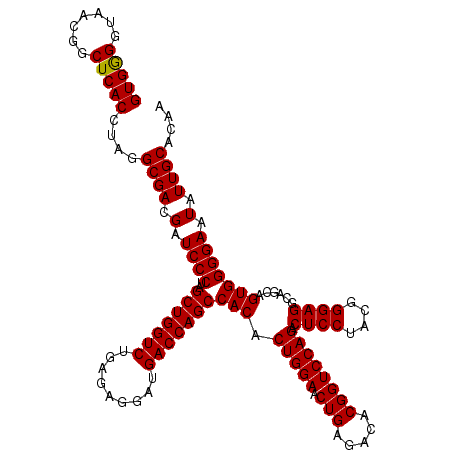

| Location | 3,940,165 – 3,940,285 |
|---|---|
| Length | 120 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 100.00 |
| Mean single sequence MFE | -45.80 |
| Consensus MFE | -41.70 |
| Energy contribution | -44.70 |
| Covariance contribution | 3.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.19 |
| Structure conservation index | 0.91 |
| SVM decision value | 0.58 |
| SVM RNA-class probability | 0.789479 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>Eco_1 3940165 120 + 1 CAGACUCCUACGGGAGGCAGCAGUGGGGAAUAUUGCACAAUGGGCGCAAGCCUGAUGCAGCCAUGCCGCGUGUAUGAAGAAGGCCUUCGGGUUGUAAAGUACUUUCAGCGGGGAGGAAGG ....((((....))))(((((((((.....)))))).....((((....))))..)))..((.(.((((..((((......((((....)))).....)))).....)))).).)).... ( -45.80) >Sfl_1 481 120 + 1 CAGACUCCUACGGGAGGCAGCAGUGGGGAAUAUUGCACAAUGGGCGCAAGCCUGAUGCAGCCAUGCCGCGUGUAUGAAGAAGGCCUUCGGGUUGUAAAGUACUUUCAGCGGGGAGGAAGG ....((((....))))(((((((((.....)))))).....((((....))))..)))..((.(.((((..((((......((((....)))).....)))).....)))).).)).... ( -45.80) >Sty_1 481 120 + 1 CAGACUCCUACGGGAGGCAGCAGUGGGGAAUAUUGCACAAUGGGCGCAAGCCUGAUGCAGCCAUGCCGCGUGUAUGAAGAAGGCCUUCGGGUUGUAAAGUACUUUCAGCGGGGAGGAAGG ....((((....))))(((((((((.....)))))).....((((....))))..)))..((.(.((((..((((......((((....)))).....)))).....)))).).)).... ( -45.80) >consensus CAGACUCCUACGGGAGGCAGCAGUGGGGAAUAUUGCACAAUGGGCGCAAGCCUGAUGCAGCCAUGCCGCGUGUAUGAAGAAGGCCUUCGGGUUGUAAAGUACUUUCAGCGGGGAGGAAGG ....((((....((..(((((((((.....)))))).....((((....))))..)))..))...((((..((((......((((....)))).....)))).....))))))))..... (-41.70 = -44.70 + 3.00)


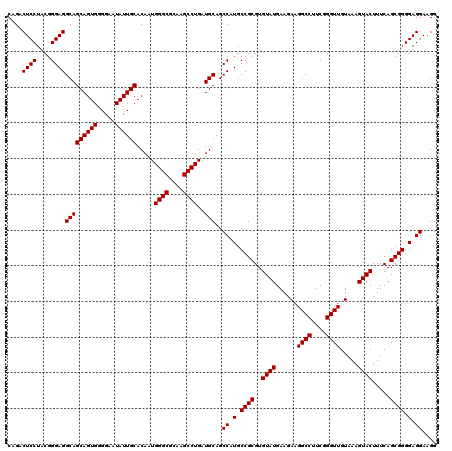
| Location | 3,939,909 – 3,940,029 |
|---|---|
| Length | 120 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 82.78 |
| Mean single sequence MFE | -39.60 |
| Consensus MFE | -36.56 |
| Energy contribution | -34.47 |
| Covariance contribution | -2.09 |
| Combinations/Pair | 1.31 |
| Mean z-score | -1.48 |
| Structure conservation index | 0.92 |
| SVM decision value | 1.19 |
| SVM RNA-class probability | 0.927722 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>Eco_1 3939909 120 + 1 ACAGCUUGCUGUUUCGCUGACGAGUGGCGGACGGGUGAGUAAUGUCUGGGAAACUGCCUGAUGGAGGGGGAUAACUACUGGAAACGGUAGCUAAUACCGCAUAACGUCGCAAGACCAAAG ....(((((.....((((.......))))((((.(((.(((.(((((((....)).(((......)))))))).((((((....))))))....))))))....)))))))))....... ( -39.20) >Hin_1 78 120 + 1 AAAGCUUGCUUUCUUGCUGACGAGUGGCGGACGGGUGAGUAAUGCUUGGGAAUCUGGCUUAUGGAGGGGGAUAACGACGGGAAACUGUCGCUAAUACCGCGUAUUAUCGGAAGAUGAAAG .((((((((((.(((((((........))).)))).)))))).))))((..((((..(((...)))..))))..((((((....))))))......)).....(((((....)))))... ( -36.30) >Ype_1 81 120 + 1 GUAGUUUACUACUUUGCCGGCGAGCGGCGGACGGGUGAGUAAUGUCUGGGGAUCUGCCUGAUGGAGGGGGAUAACUACUGGAAACGGUAGCUAAUACCGCAUGACCUCGCAAGAGCAAAG ....((((((..(((((((.....)))))))..))))))....(((((.((((((.(((.....))).))))..((((((....))))))......)).)).)))(((....)))..... ( -43.30) >consensus AAAGCUUGCUAUUUUGCUGACGAGUGGCGGACGGGUGAGUAAUGUCUGGGAAUCUGCCUGAUGGAGGGGGAUAACUACUGGAAACGGUAGCUAAUACCGCAUAACAUCGCAAGACCAAAG ...(((((((..(((((((.....)))))))..))))))).((((..((..((((.(((.....))).))))..((((((....))))))......))))))...(((....)))..... (-36.56 = -34.47 + -2.09)



| Location | 3,939,949 – 3,940,069 |
|---|---|
| Length | 120 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 80.28 |
| Mean single sequence MFE | -51.57 |
| Consensus MFE | -52.46 |
| Energy contribution | -47.37 |
| Covariance contribution | -5.10 |
| Combinations/Pair | 1.35 |
| Mean z-score | -4.68 |
| Structure conservation index | 1.02 |
| SVM decision value | 2.32 |
| SVM RNA-class probability | 0.992343 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>Eco_1 3939949 120 + 1 AAUGUCUGGGAAACUGCCUGAUGGAGGGGGAUAACUACUGGAAACGGUAGCUAAUACCGCAUAACGUCGCAAGACCAAAGAGGGGGACCUUCGGGCCUCUUGCCAUCAGAUGUGCCCAGA ....((((((..((...(((((((((((((....((((((....))))))......(((......(((....)))....((((....))))))).)))))).)))))))..)).)))))) ( -53.90) >Hin_1 118 120 + 1 AAUGCUUGGGAAUCUGGCUUAUGGAGGGGGAUAACGACGGGAAACUGUCGCUAAUACCGCGUAUUAUCGGAAGAUGAAAGUGCGGGACUGAGAGGCCGCAUGCCAUAGGAUGAGCCCAAG ....((((((..((...(((((((.(.((..((.((((((....)))))).))...)).)...(((((....)))))..((((((..((....)))))))).)))))))..)).)))))) ( -49.70) >Ype_1 121 120 + 1 AAUGUCUGGGGAUCUGCCUGAUGGAGGGGGAUAACUACUGGAAACGGUAGCUAAUACCGCAUGACCUCGCAAGAGCAAAGUGGGGGACCUUAGGGCCUCACGCCAUCGGAUGAACCCAGA ....((((((..((...(((((((..((((....((((((....))))))((((..((.(((...(((....)))....))).))....))))..))))...)))))))..)).)))))) ( -51.10) >consensus AAUGUCUGGGAAUCUGCCUGAUGGAGGGGGAUAACUACUGGAAACGGUAGCUAAUACCGCAUAACAUCGCAAGACCAAAGUGGGGGACCUUAGGGCCUCAUGCCAUCGGAUGAGCCCAGA ....((((((..((...(((((((.(.((..((.((((((....)))))).))...)).).....(((....)))....((((((..((...)).)))))).)))))))..)).)))))) (-52.46 = -47.37 + -5.10)

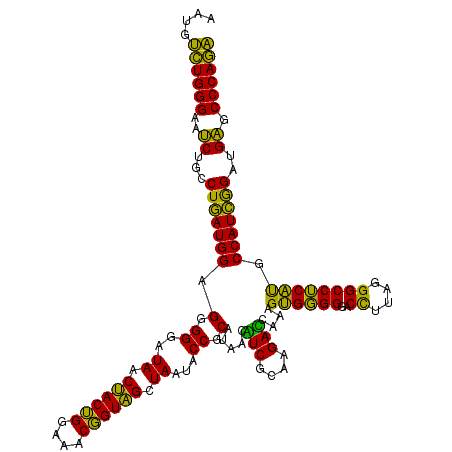

| Location | 3,939,949 – 3,940,069 |
|---|---|
| Length | 120 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 80.28 |
| Mean single sequence MFE | -37.33 |
| Consensus MFE | -30.50 |
| Energy contribution | -28.73 |
| Covariance contribution | -1.77 |
| Combinations/Pair | 1.28 |
| Mean z-score | -2.19 |
| Structure conservation index | 0.82 |
| SVM decision value | 1.71 |
| SVM RNA-class probability | 0.973597 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>Eco_1 3939949 120 - 1 UCUGGGCACAUCUGAUGGCAAGAGGCCCGAAGGUCCCCCUCUUUGGUCUUGCGACGUUAUGCGGUAUUAGCUACCGUUUCCAGUAGUUAUCCCCCUCCAUCAGGCAGUUUCCCAGACAUU ((((((.((.(((((((((((((...(((((((......))))))))))))).((.....((((((.....)))))).....)).............)))))))..))..)))))).... ( -39.90) >Hin_1 118 120 - 1 CUUGGGCUCAUCCUAUGGCAUGCGGCCUCUCAGUCCCGCACUUUCAUCUUCCGAUAAUACGCGGUAUUAGCGACAGUUUCCCGUCGUUAUCCCCCUCCAUAAGCCAGAUUCCCAAGCAUU ((((((...(((...((((.(((((..........)))))............(((((((((.((....(((....))).))))).))))))...........))))))).)))))).... ( -30.20) >Ype_1 121 120 - 1 UCUGGGUUCAUCCGAUGGCGUGAGGCCCUAAGGUCCCCCACUUUGCUCUUGCGAGGUCAUGCGGUAUUAGCUACCGUUUCCAGUAGUUAUCCCCCUCCAUCAGGCAGAUCCCCAGACAUU ((((((.((..((((((((((((((((....)))).....((((((....))))))))))))((...(((((((........))))))).)).....)))).))..))..)))))).... ( -41.90) >consensus UCUGGGCUCAUCCGAUGGCAUGAGGCCCCAAGGUCCCCCACUUUGAUCUUGCGACGUUAUGCGGUAUUAGCUACCGUUUCCAGUAGUUAUCCCCCUCCAUCAGGCAGAUUCCCAGACAUU ((((((.((..((((((((((((((((....))))..........(((....))).))))))((...(((((((........))))))).)).....)))).))..))..)))))).... (-30.50 = -28.73 + -1.77)

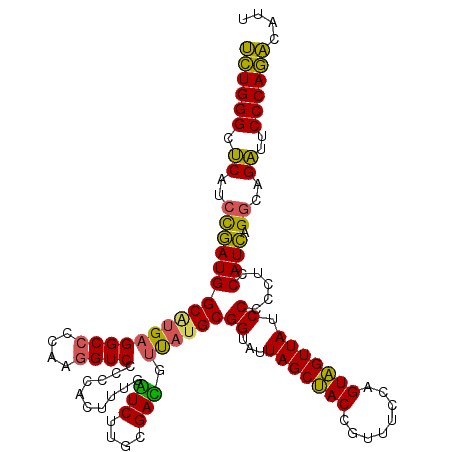

| Location | 3,939,989 – 3,940,109 |
|---|---|
| Length | 120 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 79.44 |
| Mean single sequence MFE | -43.13 |
| Consensus MFE | -38.95 |
| Energy contribution | -35.30 |
| Covariance contribution | -3.65 |
| Combinations/Pair | 1.35 |
| Mean z-score | -1.83 |
| Structure conservation index | 0.90 |
| SVM decision value | 2.08 |
| SVM RNA-class probability | 0.987436 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>Eco_1 3939989 120 + 1 GAAACGGUAGCUAAUACCGCAUAACGUCGCAAGACCAAAGAGGGGGACCUUCGGGCCUCUUGCCAUCAGAUGUGCCCAGAUGGGAUUAGCUAGUAGGUGGGGUAACGGCUCACCUAGGCG .......((((((((..(((((...(((....))).....((((((.((....))))))))........)))))(((....))))))))))).((((((((.......)))))))).... ( -44.80) >Hin_1 158 120 + 1 GAAACUGUCGCUAAUACCGCGUAUUAUCGGAAGAUGAAAGUGCGGGACUGAGAGGCCGCAUGCCAUAGGAUGAGCCCAAGUGGGAUUAGGUAGUUGGUGGGGUAAAUGCCUACCAAGCCU ..(((((.(.(((((.((((...(((((....)))))..((((((..((....))))))))......((......))..)))).)))))))))))((((((.......))))))...... ( -39.80) >Ype_1 161 120 + 1 GAAACGGUAGCUAAUACCGCAUGACCUCGCAAGAGCAAAGUGGGGGACCUUAGGGCCUCACGCCAUCGGAUGAACCCAGAUGGGAUUAGCUAGUAGGUGGGGUAAUGGCUCACCUAGGCG .......((((((((.(((.(((..(((....)))....(((((((.((....)))))).))))))))).....(((....))))))))))).((((((((.......)))))))).... ( -44.80) >consensus GAAACGGUAGCUAAUACCGCAUAACAUCGCAAGACCAAAGUGGGGGACCUUAGGGCCUCAUGCCAUCGGAUGAGCCCAGAUGGGAUUAGCUAGUAGGUGGGGUAAAGGCUCACCUAGGCG ....((((.((..............(((....)))....((((((..((...)).)))))))).))))......(((....)))....(((..((((((((.......))))))))))). (-38.95 = -35.30 + -3.65)

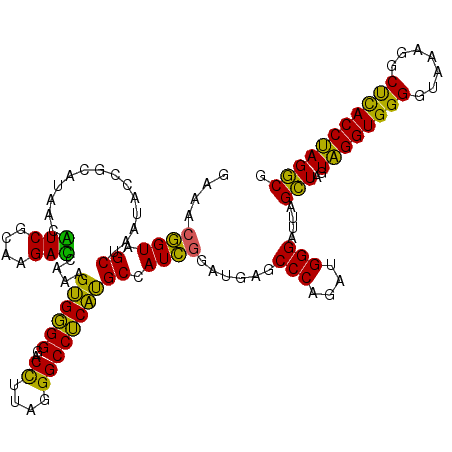

| Location | 3,940,029 – 3,940,149 |
|---|---|
| Length | 120 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 84.44 |
| Mean single sequence MFE | -47.60 |
| Consensus MFE | -42.16 |
| Energy contribution | -39.40 |
| Covariance contribution | -2.76 |
| Combinations/Pair | 1.25 |
| Mean z-score | -1.60 |
| Structure conservation index | 0.89 |
| SVM decision value | 0.60 |
| SVM RNA-class probability | 0.796809 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>Eco_1 3940029 120 + 1 AGGGGGACCUUCGGGCCUCUUGCCAUCAGAUGUGCCCAGAUGGGAUUAGCUAGUAGGUGGGGUAACGGCUCACCUAGGCGACGAUCCCUAGCUGGUCUGAGAGGAUGACCAGCCACACUG ((((((.((....)))))))).....(((.((((.......((((((.(((..((((((((.......)))))))))))...))))))..(((((((.........)))))))))))))) ( -51.30) >Hin_1 198 120 + 1 UGCGGGACUGAGAGGCCGCAUGCCAUAGGAUGAGCCCAAGUGGGAUUAGGUAGUUGGUGGGGUAAAUGCCUACCAAGCCUGCGAUCUCUAGCUGGUCUGAGAGGAUGACCAGCCACACUG ..(((.....((((..(((((.((((.((......))..)))).)).((((..((((((((.......)))))))))))))))..)))).(((((((.........)))))))....))) ( -44.60) >Ype_1 201 120 + 1 UGGGGGACCUUAGGGCCUCACGCCAUCGGAUGAACCCAGAUGGGAUUAGCUAGUAGGUGGGGUAAUGGCUCACCUAGGCGACGAUCCCUAGCUGGUCUGAGAGGAUGACCAGCCACACUG .((....)).(((((.....((((...((.(((.(((....))).))).))..((((((((.......)))))))))))).....)))))(((((((.........)))))))....... ( -46.90) >consensus UGGGGGACCUUAGGGCCUCAUGCCAUCGGAUGAGCCCAGAUGGGAUUAGCUAGUAGGUGGGGUAAAGGCUCACCUAGGCGACGAUCCCUAGCUGGUCUGAGAGGAUGACCAGCCACACUG (((((..((...)).)))))......(((.((.........((((((.(((..((((((((.......)))))))))))...))))))..(((((((.........)))))))..))))) (-42.16 = -39.40 + -2.76)

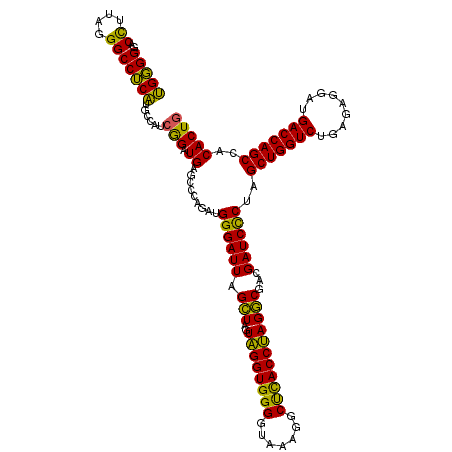

| Location | 3,940,029 – 3,940,149 |
|---|---|
| Length | 120 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 84.44 |
| Mean single sequence MFE | -42.10 |
| Consensus MFE | -33.86 |
| Energy contribution | -33.87 |
| Covariance contribution | 0.00 |
| Combinations/Pair | 1.09 |
| Mean z-score | -1.89 |
| Structure conservation index | 0.80 |
| SVM decision value | 0.41 |
| SVM RNA-class probability | 0.726383 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>Eco_1 3940029 120 - 1 CAGUGUGGCUGGUCAUCCUCUCAGACCAGCUAGGGAUCGUCGCCUAGGUGAGCCGUUACCCCACCUACUAGCUAAUCCCAUCUGGGCACAUCUGAUGGCAAGAGGCCCGAAGGUCCCCCU .....(((((((((.........)))))))))((((((.((((.((((((...........))))))...))...........((((...(((.......))).)))))).))))))... ( -43.70) >Hin_1 198 120 - 1 CAGUGUGGCUGGUCAUCCUCUCAGACCAGCUAGAGAUCGCAGGCUUGGUAGGCAUUUACCCCACCAACUACCUAAUCCCACUUGGGCUCAUCCUAUGGCAUGCGGCCUCUCAGUCCCGCA ..((((((((((((.........)))))))))(((..((((.(((.(((((................)))))....(((....)))..........))).))))..))).......))). ( -40.09) >Ype_1 201 120 - 1 CAGUGUGGCUGGUCAUCCUCUCAGACCAGCUAGGGAUCGUCGCCUAGGUGAGCCAUUACCCCACCUACUAGCUAAUCCCAUCUGGGUUCAUCCGAUGGCGUGAGGCCCUAAGGUCCCCCA .....(((((((((.........)))))))))((((((...(((((((((...........)))))........((.(((((.(((....)))))))).)).)))).....))))))... ( -42.50) >consensus CAGUGUGGCUGGUCAUCCUCUCAGACCAGCUAGGGAUCGUCGCCUAGGUGAGCCAUUACCCCACCUACUAGCUAAUCCCAUCUGGGCUCAUCCGAUGGCAUGAGGCCCCAAGGUCCCCCA .....(((((((((.........)))))))))(((((((((((.((((((...........))))))...))....(((....))).......)))(((.....)))....))))))... (-33.86 = -33.87 + 0.00)

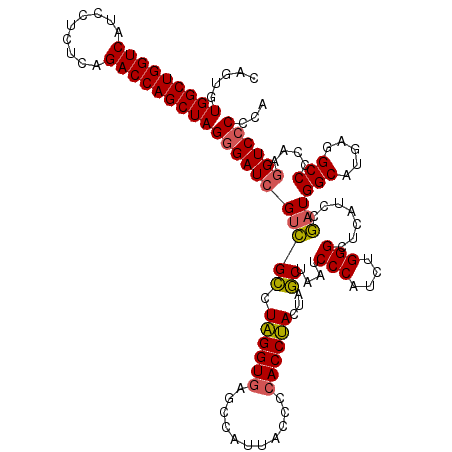
| Location | 3,940,069 – 3,940,189 |
|---|---|
| Length | 120 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 93.61 |
| Mean single sequence MFE | -50.80 |
| Consensus MFE | -49.28 |
| Energy contribution | -50.23 |
| Covariance contribution | 0.95 |
| Combinations/Pair | 1.14 |
| Mean z-score | -2.90 |
| Structure conservation index | 0.97 |
| SVM decision value | 2.23 |
| SVM RNA-class probability | 0.990734 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>Eco_1 3940069 120 + 1 UGGGAUUAGCUAGUAGGUGGGGUAACGGCUCACCUAGGCGACGAUCCCUAGCUGGUCUGAGAGGAUGACCAGCCACACUGGAACUGAGACACGGUCCAGACUCCUACGGGAGGCAGCAGU .((((((.(((..((((((((.......)))))))))))...))))))..(((((((.........)))))))....(((((.(((.....)))))))).((((....))))........ ( -51.40) >Hin_1 238 120 + 1 UGGGAUUAGGUAGUUGGUGGGGUAAAUGCCUACCAAGCCUGCGAUCUCUAGCUGGUCUGAGAGGAUGACCAGCCACACUGGAACUGAGACACGGUCCAGACUCCUACGGGAGGCAGCAGU .((((((((((..((((((((.......))))))))))))..))))))..(((((((.........)))))))....(((((.(((.....)))))))).((((....))))........ ( -49.60) >Ype_1 241 120 + 1 UGGGAUUAGCUAGUAGGUGGGGUAAUGGCUCACCUAGGCGACGAUCCCUAGCUGGUCUGAGAGGAUGACCAGCCACACUGGAACUGAGACACGGUCCAGACUCCUACGGGAGGCAGCAGU .((((((.(((..((((((((.......)))))))))))...))))))..(((((((.........)))))))....(((((.(((.....)))))))).((((....))))........ ( -51.40) >consensus UGGGAUUAGCUAGUAGGUGGGGUAAAGGCUCACCUAGGCGACGAUCCCUAGCUGGUCUGAGAGGAUGACCAGCCACACUGGAACUGAGACACGGUCCAGACUCCUACGGGAGGCAGCAGU .((((((.(((..((((((((.......)))))))))))...))))))..(((((((.........)))))))....(((((.(((.....)))))))).((((....))))........ (-49.28 = -50.23 + 0.95)


| Location | 3,940,109 – 3,940,229 |
|---|---|
| Length | 120 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 96.11 |
| Mean single sequence MFE | -50.43 |
| Consensus MFE | -48.67 |
| Energy contribution | -50.07 |
| Covariance contribution | 1.39 |
| Combinations/Pair | 1.08 |
| Mean z-score | -2.97 |
| Structure conservation index | 0.97 |
| SVM decision value | 2.07 |
| SVM RNA-class probability | 0.987273 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>Eco_1 3940109 120 + 1 ACGAUCCCUAGCUGGUCUGAGAGGAUGACCAGCCACACUGGAACUGAGACACGGUCCAGACUCCUACGGGAGGCAGCAGUGGGGAAUAUUGCACAAUGGGCGCAAGCCUGAUGCAGCCAU ....((((..(((((((.........)))))))(((.(((((.(((.....)))))))).((((....))))......)))))))...(((((...(((((....))))).))))).... ( -51.20) >Hin_1 278 120 + 1 GCGAUCUCUAGCUGGUCUGAGAGGAUGACCAGCCACACUGGAACUGAGACACGGUCCAGACUCCUACGGGAGGCAGCAGUGGGGAAUAUUGCGCAAUGGGGGGAACCCUGACGCAGCCAU ((..((((..(((((((.........)))))))(((.(((((.(((.....)))))))).((((....))))......)))))))....((((...(((((....))))).))))))... ( -48.90) >Ype_1 281 120 + 1 ACGAUCCCUAGCUGGUCUGAGAGGAUGACCAGCCACACUGGAACUGAGACACGGUCCAGACUCCUACGGGAGGCAGCAGUGGGGAAUAUUGCACAAUGGGCGCAAGCCUGAUGCAGCCAU ....((((..(((((((.........)))))))(((.(((((.(((.....)))))))).((((....))))......)))))))...(((((...(((((....))))).))))).... ( -51.20) >consensus ACGAUCCCUAGCUGGUCUGAGAGGAUGACCAGCCACACUGGAACUGAGACACGGUCCAGACUCCUACGGGAGGCAGCAGUGGGGAAUAUUGCACAAUGGGCGCAAGCCUGAUGCAGCCAU ....((((..(((((((.........)))))))(((.(((((.(((.....)))))))).((((....))))......)))))))...(((((...(((((....))))).))))).... (-48.67 = -50.07 + 1.39)


Generated by rnazCluster.pl (part of RNAz 1.0) on Fri Dec 1 11:05:49 2006