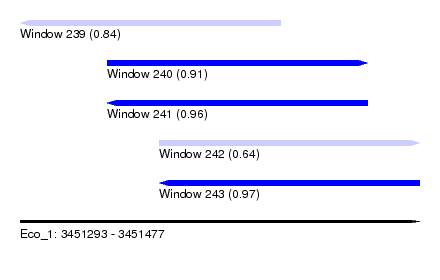
| Sequence ID | Eco_1 |
|---|---|
| Location | 3,451,293 – 3,451,477 |
| Length | 184 |
| Max. P | 0.966998 |

| Location | 3,451,293 – 3,451,413 |
|---|---|
| Length | 120 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 98.33 |
| Mean single sequence MFE | -34.07 |
| Consensus MFE | -34.00 |
| Energy contribution | -34.00 |
| Covariance contribution | 0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.80 |
| Structure conservation index | 1.00 |
| SVM decision value | 0.75 |
| SVM RNA-class probability | 0.840367 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>Eco_1 3451293 120 - 1 GGCUACCUAACAAUGCUCCCAAUCGGGGAGCUACGUAAGAACGGUUACACUCUCCCAUCAAUCGUAAUGGGUCUGAGGAGUAAUCAUUUUCGUUUAUAAAAUAAUUGGAGCUCUGGUCUC ((((.((.......((((((.....)))))).(((.((((..((((((.(((.(((((........)))))...)))..)))))).))))))).............))))))........ ( -34.10) >Sfl_1 1 120 - 1 GGCUACCUAACAAUGCUCCCAAUCGGGGAGCUACGUAAGAACGGUUACACUCUCCCAUCAAUCGUAAUGGGUCUGAGGAGUAAUCAUUUUCGUUUAUAAAAUAAUUGGAGCUCUGGUCUC ((((.((.......((((((.....)))))).(((.((((..((((((.(((.(((((........)))))...)))..)))))).))))))).............))))))........ ( -34.10) >Sty_1 1 120 - 1 GGCUAUUUAACAAUGCUCCCAAUCGGGGAGCUACGUAAGAACGGUUACACUCUCCCAUCAAUCGUAAUGGGUGUGAGGAGUAAUCAUUUUCGUUUAUAAAAUAAUUGGAGCUCUGGUCUC (((((.....((((((((((.....)))))).(((.((((..((((((.(((.(((((........)))))...)))..)))))).)))))))..........))))......))))).. ( -34.00) >consensus GGCUACCUAACAAUGCUCCCAAUCGGGGAGCUACGUAAGAACGGUUACACUCUCCCAUCAAUCGUAAUGGGUCUGAGGAGUAAUCAUUUUCGUUUAUAAAAUAAUUGGAGCUCUGGUCUC (((((.....((((((((((.....)))))).(((.((((..((((((.(((.(((((........)))))...)))..)))))).)))))))..........))))......))))).. (-34.00 = -34.00 + 0.00)

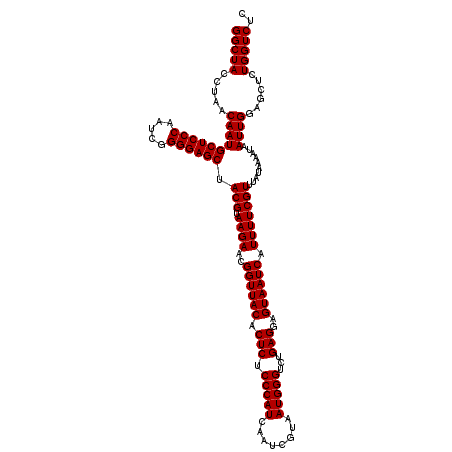
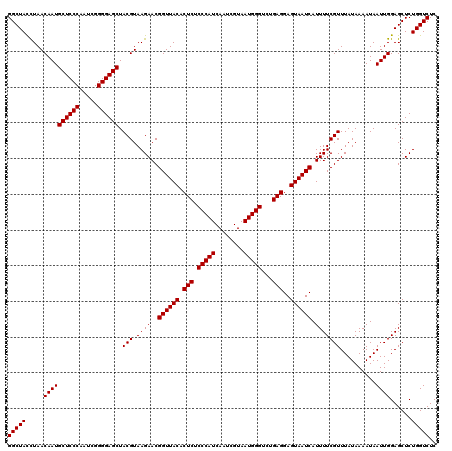
| Location | 3,451,333 – 3,451,453 |
|---|---|
| Length | 120 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 94.44 |
| Mean single sequence MFE | -38.50 |
| Consensus MFE | -37.34 |
| Energy contribution | -37.23 |
| Covariance contribution | -0.11 |
| Combinations/Pair | 1.03 |
| Mean z-score | -2.02 |
| Structure conservation index | 0.97 |
| SVM decision value | 1.09 |
| SVM RNA-class probability | 0.913444 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
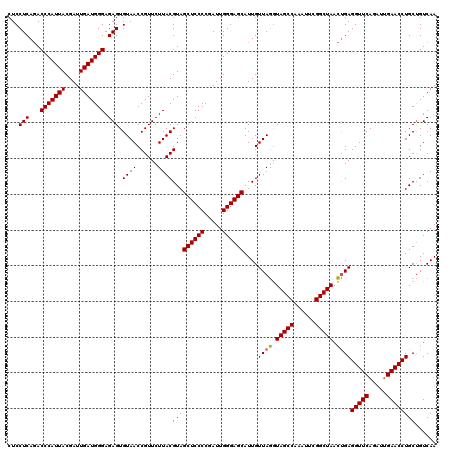
>Eco_1 3451333 120 + 1 CUCCUCAGACCCAUUACGAUUGAUGGGAGAGUGUAACCGUUCUUACGUAGCUCCCCGAUUGGGAGCAUUGUUAGGUAGCCAAAUUCGGCUAACUGAGGUUCAGAUUGAACCUGCUGUCAA ...(((...(((((((....))))))).))).((((......))))(((((((((.....))))))....((((.(((((......))))).))))(((((.....))))))))...... ( -38.70) >Sfl_1 41 120 + 1 CUCCUCAGACCCAUUACGAUUGAUGGGAGAGUGUAACCGUUCUUACGUAGCUCCCCGAUUGGGAGCAUUGUUAGGUAGCCAAAUUCGGCUAACUGAGGUUCAGAUUGAACCUGCUGUCAA ...(((...(((((((....))))))).))).((((......))))(((((((((.....))))))....((((.(((((......))))).))))(((((.....))))))))...... ( -38.70) >Sty_1 41 119 + 1 CUCCUCACACCCAUUACGAUUGAUGGGAGAGUGUAACCGUUCUUACGUAGCUCCCCGAUUGGGAGCAUUGUUAAAUAGCCAAAU-CGGCUAUUCGAGGUUCAAAUCGAACCUGCCGUCAA ...(((...(((((((....))))))).)))..(((((((....)))..((((((.....))))))...))))(((((((....-.)))))))..((((((.....))))))........ ( -38.10) >consensus CUCCUCAGACCCAUUACGAUUGAUGGGAGAGUGUAACCGUUCUUACGUAGCUCCCCGAUUGGGAGCAUUGUUAGGUAGCCAAAUUCGGCUAACUGAGGUUCAGAUUGAACCUGCUGUCAA ...(((...(((((((....))))))).))).((((......))))(((((((((.....))))))....((((.(((((......))))).))))(((((.....))))))))...... (-37.34 = -37.23 + -0.11)

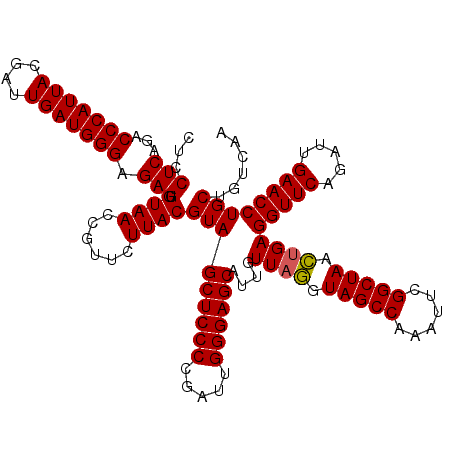
| Location | 3,451,333 – 3,451,453 |
|---|---|
| Length | 120 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 94.44 |
| Mean single sequence MFE | -38.77 |
| Consensus MFE | -38.30 |
| Energy contribution | -37.63 |
| Covariance contribution | -0.66 |
| Combinations/Pair | 1.06 |
| Mean z-score | -2.16 |
| Structure conservation index | 0.99 |
| SVM decision value | 1.48 |
| SVM RNA-class probability | 0.958020 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
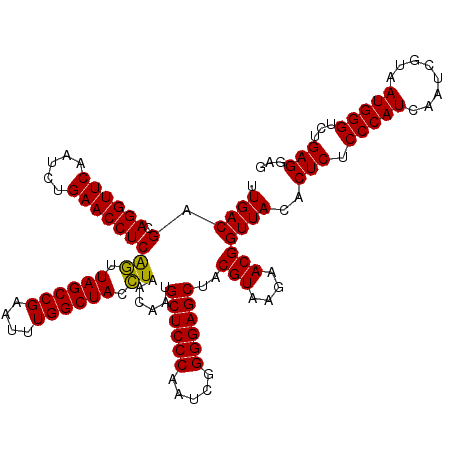
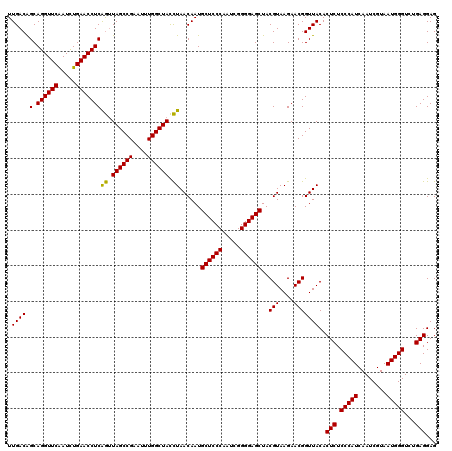
>Eco_1 3451333 120 - 1 UUGACAGCAGGUUCAAUCUGAACCUCAGUUAGCCGAAUUUGGCUACCUAACAAUGCUCCCAAUCGGGGAGCUACGUAAGAACGGUUACACUCUCCCAUCAAUCGUAAUGGGUCUGAGGAG .((((.(.((((((.....)))))))((.((((((....)))))).))......((((((.....))))))..(((....)))))))..(((.(((((........)))))...)))... ( -38.50) >Sfl_1 41 120 - 1 UUGACAGCAGGUUCAAUCUGAACCUCAGUUAGCCGAAUUUGGCUACCUAACAAUGCUCCCAAUCGGGGAGCUACGUAAGAACGGUUACACUCUCCCAUCAAUCGUAAUGGGUCUGAGGAG .((((.(.((((((.....)))))))((.((((((....)))))).))......((((((.....))))))..(((....)))))))..(((.(((((........)))))...)))... ( -38.50) >Sty_1 41 119 - 1 UUGACGGCAGGUUCGAUUUGAACCUCGAAUAGCCG-AUUUGGCUAUUUAACAAUGCUCCCAAUCGGGGAGCUACGUAAGAACGGUUACACUCUCCCAUCAAUCGUAAUGGGUGUGAGGAG .((((.(.((((((.....)))))))((((((((.-....))))))))......((((((.....))))))..(((....)))))))..(((.(((((........)))))...)))... ( -39.30) >consensus UUGACAGCAGGUUCAAUCUGAACCUCAGUUAGCCGAAUUUGGCUACCUAACAAUGCUCCCAAUCGGGGAGCUACGUAAGAACGGUUACACUCUCCCAUCAAUCGUAAUGGGUCUGAGGAG .((((.(.((((((.....)))))))((.((((((....)))))).))......((((((.....))))))..(((....)))))))..(((.(((((........)))))...)))... (-38.30 = -37.63 + -0.66)

| Location | 3,451,357 – 3,451,477 |
|---|---|
| Length | 120 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 94.44 |
| Mean single sequence MFE | -38.30 |
| Consensus MFE | -36.68 |
| Energy contribution | -36.90 |
| Covariance contribution | 0.22 |
| Combinations/Pair | 1.03 |
| Mean z-score | -1.57 |
| Structure conservation index | 0.96 |
| SVM decision value | 0.23 |
| SVM RNA-class probability | 0.642766 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
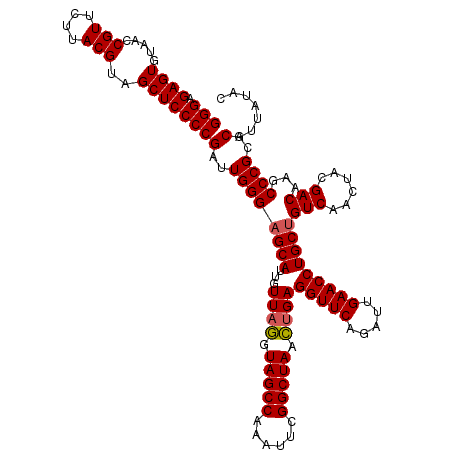
>Eco_1 3451357 120 + 1 GGGAGAGUGUAACCGUUCUUACGUAGCUCCCCGAUUGGGAGCAUUGUUAGGUAGCCAAAUUCGGCUAACUGAGGUUCAGAUUGAACCUGCUGUCAACUACGACAAGCCCGCGCAUUAUAC (((.((((.....(((....)))..)))))))(..((((((((...((((.(((((......))))).))))(((((.....)))))))))(((......)))...))))..)....... ( -39.20) >Sfl_1 65 120 + 1 GGGAGAGUGUAACCGUUCUUACGUAGCUCCCCGAUUGGGAGCAUUGUUAGGUAGCCAAAUUCGGCUAACUGAGGUUCAGAUUGAACCUGCUGUCAACUACGACAAGCCCGCGCAUUAUAC (((.((((.....(((....)))..)))))))(..((((((((...((((.(((((......))))).))))(((((.....)))))))))(((......)))...))))..)....... ( -39.20) >Sty_1 65 119 + 1 GGGAGAGUGUAACCGUUCUUACGUAGCUCCCCGAUUGGGAGCAUUGUUAAAUAGCCAAAU-CGGCUAUUCGAGGUUCAAAUCGAACCUGCCGUCAAUUACGACAAGCCCGCGCAUUAUAC (((.((((.....(((....)))..)))))))(..((((.(((...(..(((((((....-.)))))))..)(((((.....)))))))).(((......)))...))))..)....... ( -36.50) >consensus GGGAGAGUGUAACCGUUCUUACGUAGCUCCCCGAUUGGGAGCAUUGUUAGGUAGCCAAAUUCGGCUAACUGAGGUUCAGAUUGAACCUGCUGUCAACUACGACAAGCCCGCGCAUUAUAC (((.((((.....(((....)))..)))))))(..((((((((...((((.(((((......))))).))))(((((.....)))))))))(((......)))...))))..)....... (-36.68 = -36.90 + 0.22)

| Location | 3,451,357 – 3,451,477 |
|---|---|
| Length | 120 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 94.44 |
| Mean single sequence MFE | -40.63 |
| Consensus MFE | -39.92 |
| Energy contribution | -39.03 |
| Covariance contribution | -0.88 |
| Combinations/Pair | 1.07 |
| Mean z-score | -2.28 |
| Structure conservation index | 0.98 |
| SVM decision value | 1.60 |
| SVM RNA-class probability | 0.966998 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
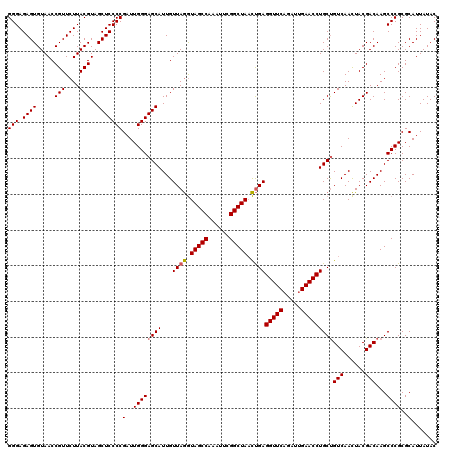
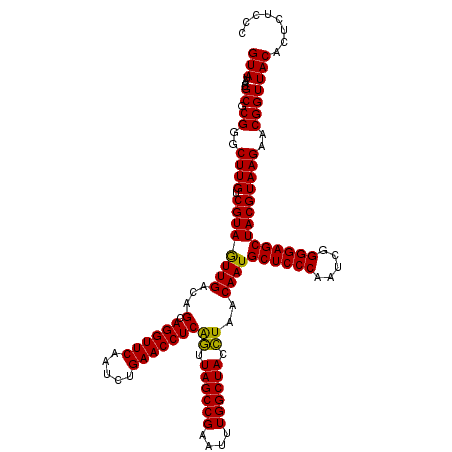
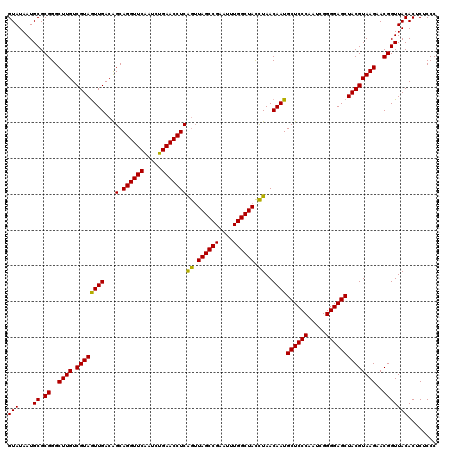
>Eco_1 3451357 120 - 1 GUAUAAUGCGCGGGCUUGUCGUAGUUGACAGCAGGUUCAAUCUGAACCUCAGUUAGCCGAAUUUGGCUACCUAACAAUGCUCCCAAUCGGGGAGCUACGUAAGAACGGUUACACUCUCCC (((....((.((..((((.((((((((...(.((((((.....)))))))((.((((((....)))))).))..))))((((((.....))))))))))))))..)))))))........ ( -40.00) >Sfl_1 65 120 - 1 GUAUAAUGCGCGGGCUUGUCGUAGUUGACAGCAGGUUCAAUCUGAACCUCAGUUAGCCGAAUUUGGCUACCUAACAAUGCUCCCAAUCGGGGAGCUACGUAAGAACGGUUACACUCUCCC (((....((.((..((((.((((((((...(.((((((.....)))))))((.((((((....)))))).))..))))((((((.....))))))))))))))..)))))))........ ( -40.00) >Sty_1 65 119 - 1 GUAUAAUGCGCGGGCUUGUCGUAAUUGACGGCAGGUUCGAUUUGAACCUCGAAUAGCCG-AUUUGGCUAUUUAACAAUGCUCCCAAUCGGGGAGCUACGUAAGAACGGUUACACUCUCCC ..........((((((((((((.....))))))))))))...((((((..((((((((.-....)))))))).((...((((((.....))))))...))......)))).))....... ( -41.90) >consensus GUAUAAUGCGCGGGCUUGUCGUAGUUGACAGCAGGUUCAAUCUGAACCUCAGUUAGCCGAAUUUGGCUACCUAACAAUGCUCCCAAUCGGGGAGCUACGUAAGAACGGUUACACUCUCCC (((....((.((..((((.((((((((...(.((((((.....)))))))((.((((((....)))))).))..))))((((((.....))))))))))))))..)))))))........ (-39.92 = -39.03 + -0.88)

Generated by rnazCluster.pl (part of RNAz 1.0) on Fri Dec 1 11:05:31 2006