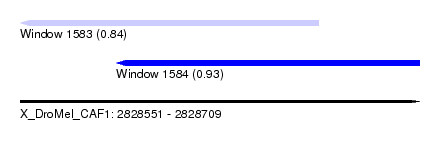

| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 2,828,551 – 2,828,709 |
| Length | 158 |
| Max. P | 0.928196 |

| Location | 2,828,551 – 2,828,669 |
|---|---|
| Length | 118 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 80.80 |
| Mean single sequence MFE | -23.50 |
| Consensus MFE | -15.31 |
| Energy contribution | -15.25 |
| Covariance contribution | -0.06 |
| Combinations/Pair | 1.13 |
| Mean z-score | -2.01 |
| Structure conservation index | 0.65 |
| SVM decision value | 0.74 |
| SVM RNA-class probability | 0.838525 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
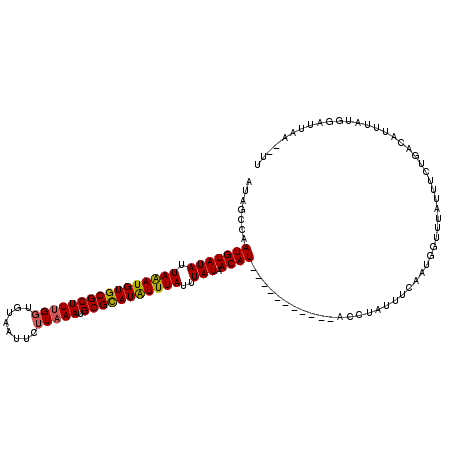
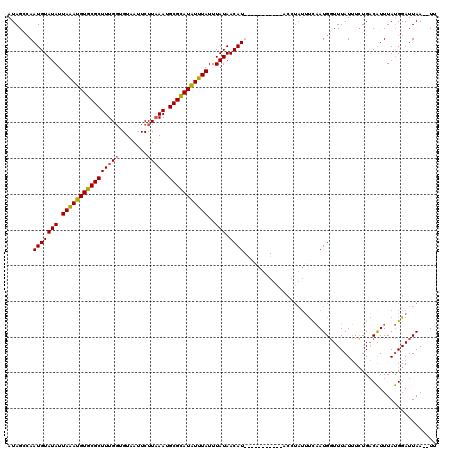
>X_DroMel_CAF1 2828551 118 - 22224390 AUAGGCAAUGUAUAUUAAAUGUGCGCUUUGGUGUAAUUCUUUAAUGCGUAUAUUUAUUUAUAACAUUCGAAUGGUAUACCUAUUUCAAUGGUUUAUUUCUGACAUUUAUGGAUUAA--UU ((((((..((((...(((((((((((.((((.(.....).)))).)))))))))))..)))).((((.((((((.....)))))).))))))))))....................--.. ( -22.70) >DroSec_CAF1 216783 118 - 1 AUAGCGAAUGUAUAUUAGAUGUGCGCUUUGGUGUAAUUCUUAAAUGCGCAUAUUUAUUUAUAACAUUUGAAUUGUAUGCCUAUUUCAAUGGUUUAUUUCUGACAUUUAUGGAUUAA--UU ....((((((((((.((((((((((((((((........))))).)))))))))))..))).)))))))....(((.(((.........))).)))....................--.. ( -24.00) >DroSim_CAF1 189362 101 - 1 AUAGCCAAUGUAUAUUAAAUGUGCGCUUUGGUGUAAUUCUUAAAUGCGCAUAUUUAUUUAUAACAU-----------------UUCAAUGGUUUAUUUCUGACAUUUAUGGAUUAA--UU ..((((((((((((.((((((((((((((((........))))).)))))))))))..))).))))-----------------.....))))).......................--.. ( -20.10) >DroEre_CAF1 236525 109 - 1 AUAGGCAAUGUAUAUUAAAUGUGCGCUUUGCCGUUAUUCUUAAAUGCGCAUGUUUAUUUAUAACAU-----------ACUUAUACUUAUGUUUUAUGUGUGUCGUUUACGGAUUAACAUA ...((((((((((((...)))))))..)))))((((.((((((((((((((((........(((((-----------(........))))))..))))))).)))))).))).))))... ( -27.20) >consensus AUAGCCAAUGUAUAUUAAAUGUGCGCUUUGGUGUAAUUCUUAAAUGCGCAUAUUUAUUUAUAACAU___________ACCUAUUUCAAUGGUUUAUUUCUGACAUUUAUGGAUUAA__UU .......(((((((.((((((((((((((((........))))).)))))))))))..))).))))...................................................... (-15.31 = -15.25 + -0.06)

| Location | 2,828,589 – 2,828,709 |
|---|---|
| Length | 120 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 86.46 |
| Mean single sequence MFE | -23.03 |
| Consensus MFE | -17.61 |
| Energy contribution | -17.18 |
| Covariance contribution | -0.44 |
| Combinations/Pair | 1.17 |
| Mean z-score | -2.27 |
| Structure conservation index | 0.76 |
| SVM decision value | 1.19 |
| SVM RNA-class probability | 0.928196 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
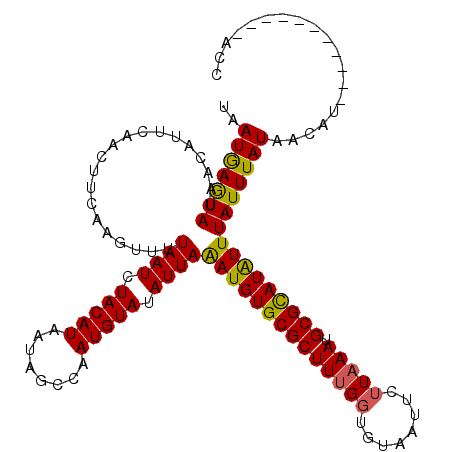
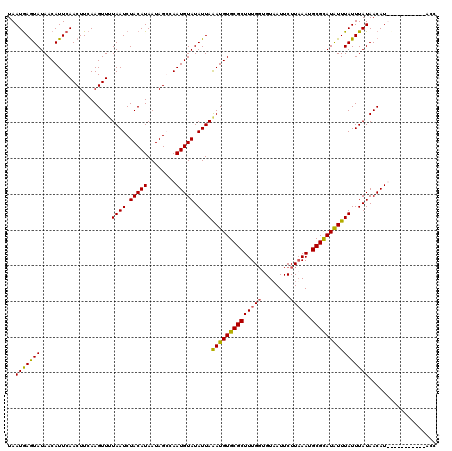
>X_DroMel_CAF1 2828589 120 - 22224390 UAAUGAGUAUAACAUUCAACUUCAAGUUUUAAUCUACAUAAUAGGCAAUGUAUAUUAAAUGUGCGCUUUGGUGUAAUUCUUUAAUGCGUAUAUUUAUUUAUAACAUUCGAAUGGUAUACC ......(((((.(((((........((........)).......(.((((((((.(((((((((((.((((.(.....).)))).)))))))))))..))).))))))))))).))))). ( -24.80) >DroSec_CAF1 216821 120 - 1 UAAUGAGUAUAACAUUCAACUUCAAGUUUUAAUCUACAUAAUAGCGAAUGUAUAUUAGAUGUGCGCUUUGGUGUAAUUCUUAAAUGCGCAUAUUUAUUUAUAACAUUUGAAUUGUAUGCC ...(((((.....)))))..................((((....((((((((((.((((((((((((((((........))))).)))))))))))..))).))))))).....)))).. ( -26.30) >DroSim_CAF1 189397 106 - 1 UAAUGAGUAUAACAUUCAACUUCAAGUUAUAAUCUACAUAAUAGCCAAUGUAUAUUAAAUGUGCGCUUUGGUGUAAUUCUUAAAUGCGCAUAUUUAUUUAUAACAU-------------- ...(((((.....))))).......(((((((..(((((........)))))...((((((((((((((((........))))).))))))))))).)))))))..-------------- ( -23.40) >DroEre_CAF1 236565 109 - 1 UAAUAAAUAUAACAUUCAACUUCAAGUUUUAAUCUACAUAAUAGGCAAUGUAUAUUAAAUGUGCGCUUUGCCGUUAUUCUUAAAUGCGCAUGUUUAUUUAUAACAU-----------ACU ..(((((((.(((..............((((((.(((((........))))).)))))).((((((...................)))))))))))))))).....-----------... ( -17.61) >consensus UAAUGAGUAUAACAUUCAACUUCAAGUUUUAAUCUACAUAAUAGCCAAUGUAUAUUAAAUGUGCGCUUUGGUGUAAUUCUUAAAUGCGCAUAUUUAUUUAUAACAU___________ACC ..(((((((....................((((.(((((........))))).))))((((((((((((((........))))).))))))))))))))))................... (-17.61 = -17.18 + -0.44)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:58:33 2006