
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 2,776,449 – 2,776,553 |
| Length | 104 |
| Max. P | 0.685899 |

| Location | 2,776,449 – 2,776,553 |
|---|---|
| Length | 104 |
| Sequences | 6 |
| Columns | 118 |
| Reading direction | forward |
| Mean pairwise identity | 77.99 |
| Mean single sequence MFE | -30.90 |
| Consensus MFE | -19.44 |
| Energy contribution | -19.08 |
| Covariance contribution | -0.36 |
| Combinations/Pair | 1.17 |
| Mean z-score | -1.48 |
| Structure conservation index | 0.63 |
| SVM decision value | 0.32 |
| SVM RNA-class probability | 0.685899 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 2776449 104 + 22224390 CGAACGGCAAACGGAGUG--GG----GAUAC--GUAGCCA------GGAUGCGACACGCAUUUGUUGGAUUAAUUGUGUGCUGGCUGGCAGUUUGUCAACCGCAAGGUAAAUAUUUUU .....(((((((..(((.--((----.((((--(...(((------(((((((...))))))..))))......))))).)).)))....))))))).(((....))).......... ( -33.20) >DroVir_CAF1 238370 107 + 1 ---AC--AGAUACA----CAGCUACAGAUAC--GGCGUCAGGAUGAGGAUGCGCCACGCAUUUGCUGGAUUAAUUGUGUGUUGGCUGGCAGUUUGUCAACCGCAAGGUAAAUAUUUUU ---((--((((...----((((((.(.((((--((((((........))))).(((.((....)))))......))))).)))))))...))))))..(((....))).......... ( -32.00) >DroPse_CAF1 236993 95 + 1 ---------------GAG--GA----GAUAC--AUGGCCAGGAUGAGGAUGCGACACGCAUUUGUUGGAUUAAUUGUGUGCUGGCUGGCAGUUUGUCAACCGCAAGGUAAAUAUUUUU ---------------((.--((----(....--.(((((((.(((..(((.(((((......))))).))).....))).)))))))....))).)).(((....))).......... ( -27.00) >DroGri_CAF1 243417 102 + 1 ----C--AGAGAUA----CCGC----GAUAC--GCUGCCAGGAUGAGGAUGCGCCACGCAUUUGCUGGAUUAAUUGUGUGCUGGCUGGCAGUUUGUCAACCGCAAGGUAAAUAUUUUU ----.--(((((((----....----((((.--((((((((..(.((.((((((((.((....)))))......))))).)).))))))))).)))).(((....)))...))))))) ( -33.60) >DroEre_CAF1 181593 102 + 1 ----CGGCGAACGGCGUA--GG----GACACAGGGAGCCA------GGAUGCGACACGCAUUUGUUGGAUUAAUUGUGUGCUGGCUGGCAGUUUGUCAACCGCAAGGUAAAUAUUUUU ----.(((((((.((((.--((----.((((((..(((((------(((((((...))))))..)))).))..)))))).)).))..)).))))))).(((....))).......... ( -36.30) >DroAna_CAF1 276919 91 + 1 ----------ACGGAGCA--GA----GA-----GGAGUCA------GGAUGCGACACGCAUUUGUUGGAUUAAUUGUGUGCUGGCUGGCAGUUUGUCAACCGCAAGGUAAAUAUUUUU ----------.(((.(((--..----((-----..((((.------(((((((...)))))))....))))..)).))).)))..((((.....))))(((....))).......... ( -23.30) >consensus ____C___GAACGG_G____GA____GAUAC__GGAGCCA______GGAUGCGACACGCAUUUGUUGGAUUAAUUGUGUGCUGGCUGGCAGUUUGUCAACCGCAAGGUAAAUAUUUUU ..........................((((......((((......(((((((...)))))))((..(((.......)).)..))))))....)))).(((....))).......... (-19.44 = -19.08 + -0.36)

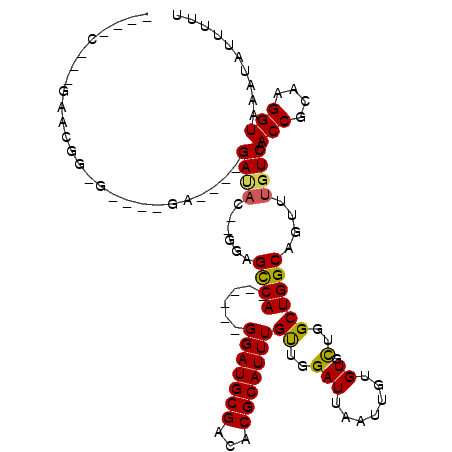
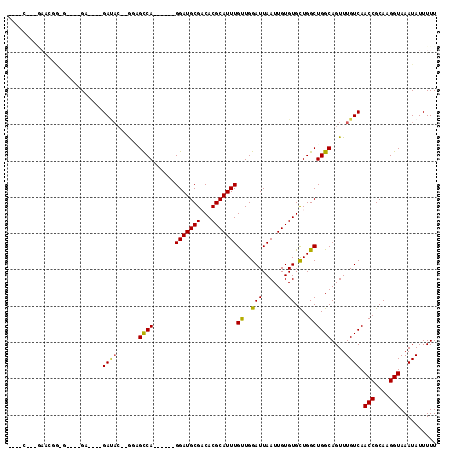
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:58:22 2006