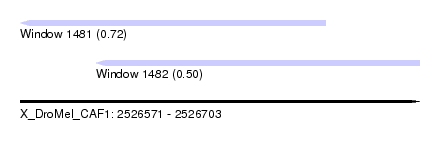

| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 2,526,571 – 2,526,703 |
| Length | 132 |
| Max. P | 0.720662 |

| Location | 2,526,571 – 2,526,672 |
|---|---|
| Length | 101 |
| Sequences | 4 |
| Columns | 118 |
| Reading direction | reverse |
| Mean pairwise identity | 74.85 |
| Mean single sequence MFE | -18.20 |
| Consensus MFE | -12.10 |
| Energy contribution | -12.10 |
| Covariance contribution | -0.00 |
| Combinations/Pair | 1.07 |
| Mean z-score | -1.60 |
| Structure conservation index | 0.66 |
| SVM decision value | 0.40 |
| SVM RNA-class probability | 0.720662 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
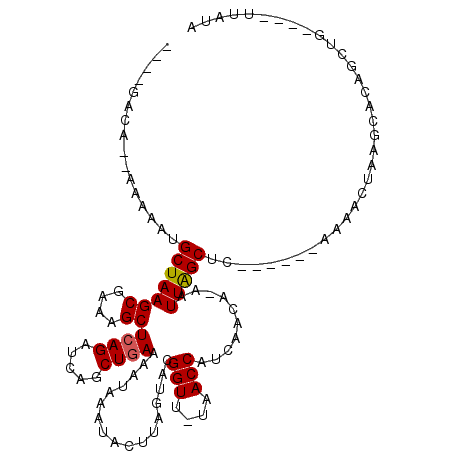
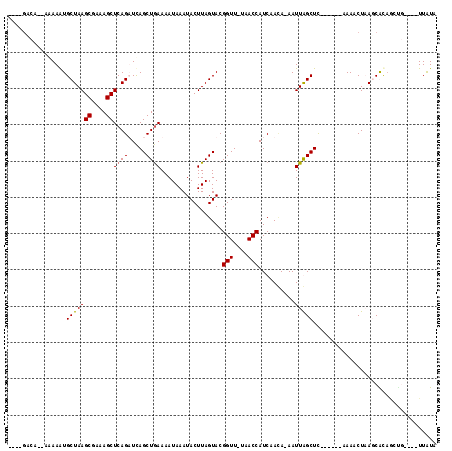
>X_DroMel_CAF1 2526571 101 - 22224390 AUAACACA--UAAAAUGCUAAGCGAAAGCUCAGUUUAGCUAAAAAUAAAUACUUAGUACGGUUUUAACCAUCAACAAAAUUGGCUC------AAAACUCAUCAU---------UUAUA ........--..(((((...(((....))).(((((((((((.......(((...))).(((....)))..........)))))).------.)))))...)))---------))... ( -14.40) >DroSec_CAF1 1843 101 - 1 ----GUCA--AAAAGUGCUAAGCUAAAGCUCAGUUCAGCUGAAAAUAAAUACUUAGUACGGUU-UAACCAUCAACAUAAUUGGCUU------AAAACUCAGCACAGCUG----UUAUU ----((((--(...((((((((.......((((.....)))).........))))))))((..-...))..........)))))..------......((((...))))----..... ( -20.89) >DroEre_CAF1 2227 100 - 1 ----G------UAAAUGCUAAGCGCAAGCUCAGAUCAGCUGAAAAUAAAUACUUAGUACGGUU-UAACCAUCAAUA-AAUUAGCUC------AAAACUAAGCGCAACUCUUAUUUAUA ----(------((((((((.(((....))).))....((....................((..-...)).......-.....(((.------.......)))))......))))))). ( -15.00) >DroYak_CAF1 2251 112 - 1 ----GACGAUAGAAAUGCUAAGCGAAAGCUCAGAUCAGCUGAAAAUAAAUACUUAGUACGGUU-UAACCAUCAAGA-AAUUAGCUCCAAAACAAAGCUAAACGCAGGCGUCAGUUGUA ----((((...((..((((((((....))((((.....))))..........)))))).((..-...)).))....-..((((((.........)))))).......))))....... ( -22.50) >consensus ____GACA__AAAAAUGCUAAGCGAAAGCUCAGAUCAGCUGAAAAUAAAUACUUAGUACGGUU_UAACCAUCAACA_AAUUAGCUC______AAAACUAAGCACAGCUG____UUAUA ................(((((((....))((((.....)))).................(((....)))..........))))).................................. (-12.10 = -12.10 + -0.00)

| Location | 2,526,596 – 2,526,703 |
|---|---|
| Length | 107 |
| Sequences | 4 |
| Columns | 109 |
| Reading direction | reverse |
| Mean pairwise identity | 81.24 |
| Mean single sequence MFE | -18.31 |
| Consensus MFE | -12.18 |
| Energy contribution | -12.43 |
| Covariance contribution | 0.25 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.56 |
| Structure conservation index | 0.66 |
| SVM decision value | -0.07 |
| SVM RNA-class probability | 0.500000 |
| Prediction | OTHER |
Download alignment: ClustalW | MAF
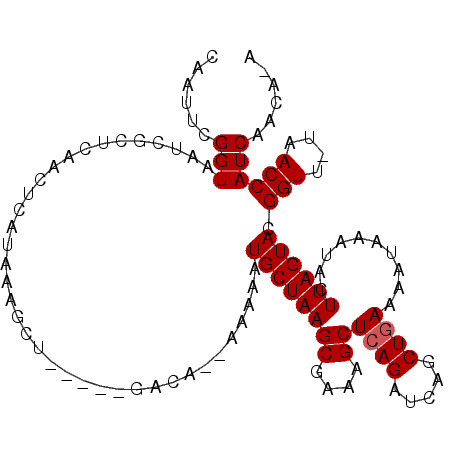
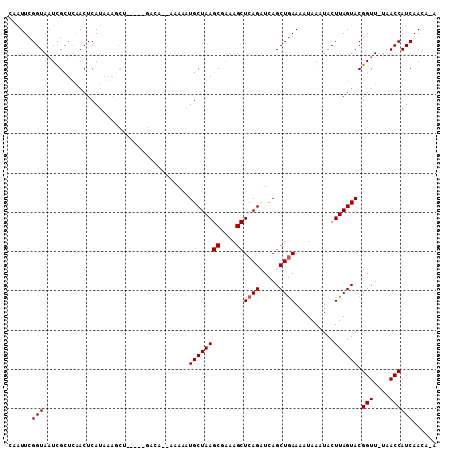
>X_DroMel_CAF1 2526596 107 - 22224390 CAGUUCGGUAAUCGCUUAACUCAUAAACUCAAUAACACA--UAAAAUGCUAAGCGAAAGCUCAGUUUAGCUAAAAAUAAAUACUUAGUACGGUUUUAACCAUCAACAAA ......((((((((.........................--......(((.(((....))).)))...(((((..........))))).)))))...)))......... ( -14.90) >DroSec_CAF1 1873 101 - 1 CAAUUCGGUAAUCGCUUUACUCAUAAAACG-----GUCA--AAAAGUGCUAAGCUAAAGCUCAGUUCAGCUGAAAAUAAAUACUUAGUACGGUU-UAACCAUCAACAUA ......(((((.....)))))........(-----((.(--((..((((((((.......((((.....)))).........))))))))..))-).)))......... ( -20.29) >DroEre_CAF1 2261 96 - 1 CAAUUCGGUAUUUGUGCAACUCAUAAAGCU-----G------UAAAUGCUAAGCGCAAGCUCAGAUCAGCUGAAAAUAAAUACUUAGUACGGUU-UAACCAUCAAUA-A ..(((.(((((((((...........((((-----(------......((.(((....))).))..)))))....))))))))).)))..((..-...)).......-. ( -20.86) >DroYak_CAF1 2291 102 - 1 CAAUUCGGUAAUUGCUCAACUCAUAAGGCU-----GACGAUAGAAAUGCUAAGCGAAAGCUCAGAUCAGCUGAAAAUAAAUACUUAGUACGGUU-UAACCAUCAAGA-A ......(((.((((.(((.((.....)).)-----)))))).....((((((((....))((((.....))))..........)))))).....-..))).......-. ( -17.20) >consensus CAAUUCGGUAAUCGCUCAACUCAUAAAGCU_____GACA__AAAAAUGCUAAGCGAAAGCUCAGAUCAGCUGAAAAUAAAUACUUAGUACGGUU_UAACCAUCAACA_A ......(((.....................................((((((((....))((((.....))))..........)))))).(((....))))))...... (-12.18 = -12.43 + 0.25)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:56:57 2006