
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 21,968,142 – 21,968,241 |
| Length | 99 |
| Max. P | 0.668916 |

| Location | 21,968,142 – 21,968,241 |
|---|---|
| Length | 99 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 86.15 |
| Mean single sequence MFE | -14.60 |
| Consensus MFE | -10.88 |
| Energy contribution | -10.88 |
| Covariance contribution | 0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.48 |
| Structure conservation index | 0.74 |
| SVM decision value | 0.08 |
| SVM RNA-class probability | 0.571859 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 21968142 99 + 22224390 UCCACAAAC--------CAUUUUCCC---------ACUUUUCCACGCCCACCUGUCAGUCUACGCGUCAGUUUUUCAUUCUCACUUAUUUUGCCCU----GGCAGGCAUUCAUCAGCGUU .........--------.........---------........((((...((((((((.....(((........................))).))----)))))).........)))). ( -12.36) >DroSec_CAF1 33558 99 + 1 UCCACAAAC--------CAUUUUCCC---------ACUUUUCCACGCCCACCUGUCAGUCUACCCGUCAGUUUUUCAUUCUCACUUAUUGUGCCCU----GGCAGGCAUUCAUCAGCGUU .........--------.........---------........((((...((((((((..(((..((.(((...........))).)).)))..))----)))))).........)))). ( -15.00) >DroSim_CAF1 20315 98 + 1 UCCACAAAC--------CAUUUUCCC---------ACUU-UCCACGCCCACCUGUCAGUCUACCCGUCAGUUUUUCAUUCUCACUUAUUGUGCCCU----GGCAGGCAUUCAUCAGCGUU .........--------.........---------....-...((((...((((((((..(((..((.(((...........))).)).)))..))----)))))).........)))). ( -15.00) >DroYak_CAF1 23073 119 + 1 GCCACACCCUAUUUCCCCAUUUUCCCACUUUUCCCACUUUUCCACGCCCACCUGUCAGUCUACCCGUCAGUUAUUCAUUCUCACUUUU-UUGCCCUGGCUGGCAGGCAUUCAUCAGCGUU ...........................................((((...((((((((((............................-.......)))))))))).........)))). ( -16.05) >consensus UCCACAAAC________CAUUUUCCC_________ACUUUUCCACGCCCACCUGUCAGUCUACCCGUCAGUUUUUCAUUCUCACUUAUUGUGCCCU____GGCAGGCAUUCAUCAGCGUU ...........................................((((.....((((.........((.((.........)).))......((((......)))))))).......)))). (-10.88 = -10.88 + 0.00)

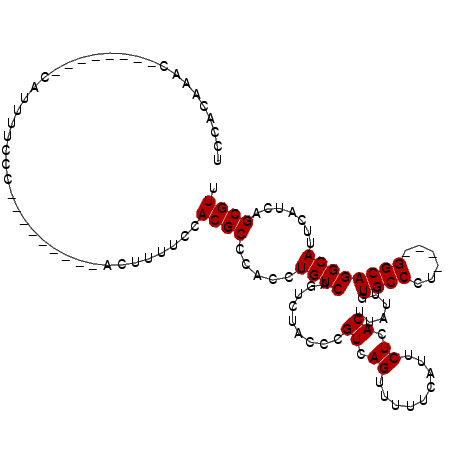
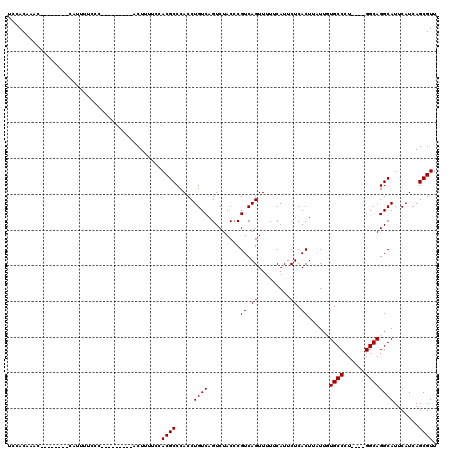
| Location | 21,968,142 – 21,968,241 |
|---|---|
| Length | 99 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 86.15 |
| Mean single sequence MFE | -23.67 |
| Consensus MFE | -19.08 |
| Energy contribution | -19.32 |
| Covariance contribution | 0.25 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.59 |
| Structure conservation index | 0.81 |
| SVM decision value | 0.29 |
| SVM RNA-class probability | 0.668916 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 21968142 99 - 22224390 AACGCUGAUGAAUGCCUGCC----AGGGCAAAAUAAGUGAGAAUGAAAAACUGACGCGUAGACUGACAGGUGGGCGUGGAAAAGU---------GGGAAAAUG--------GUUUGUGGA ..((((.....(((((..((----............((.((.........)).))..((......)).))..))))).....)))---------)........--------......... ( -21.30) >DroSec_CAF1 33558 99 - 1 AACGCUGAUGAAUGCCUGCC----AGGGCACAAUAAGUGAGAAUGAAAAACUGACGGGUAGACUGACAGGUGGGCGUGGAAAAGU---------GGGAAAAUG--------GUUUGUGGA ..((((.....(((((..((----(...(((.....)))....)).....(((.(((.....))).))))..))))).....)))---------)........--------......... ( -24.20) >DroSim_CAF1 20315 98 - 1 AACGCUGAUGAAUGCCUGCC----AGGGCACAAUAAGUGAGAAUGAAAAACUGACGGGUAGACUGACAGGUGGGCGUGGA-AAGU---------GGGAAAAUG--------GUUUGUGGA ..((((.....(((((..((----(...(((.....)))....)).....(((.(((.....))).))))..)))))...-.)))---------)........--------......... ( -23.80) >DroYak_CAF1 23073 119 - 1 AACGCUGAUGAAUGCCUGCCAGCCAGGGCAA-AAAAGUGAGAAUGAAUAACUGACGGGUAGACUGACAGGUGGGCGUGGAAAAGUGGGAAAAGUGGGAAAAUGGGGAAAUAGGGUGUGGC ..((((.....(((((..((.(.(((.((..-...(((...........))).....))...))).).))..))))).....)))).................................. ( -25.40) >consensus AACGCUGAUGAAUGCCUGCC____AGGGCAAAAUAAGUGAGAAUGAAAAACUGACGGGUAGACUGACAGGUGGGCGUGGAAAAGU_________GGGAAAAUG________GUUUGUGGA .(((((.((...(((((........)))))....................(((.(((.....))).))))).)))))........................................... (-19.08 = -19.32 + 0.25)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 13:10:07 2006