
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 21,853,562 – 21,853,693 |
| Length | 131 |
| Max. P | 0.871070 |

| Location | 21,853,562 – 21,853,665 |
|---|---|
| Length | 103 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 73.68 |
| Mean single sequence MFE | -21.84 |
| Consensus MFE | -6.32 |
| Energy contribution | -7.48 |
| Covariance contribution | 1.16 |
| Combinations/Pair | 1.27 |
| Mean z-score | -1.72 |
| Structure conservation index | 0.29 |
| SVM decision value | 0.21 |
| SVM RNA-class probability | 0.634669 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 21853562 103 - 22224390 AGUG--CUCGCCUACUUUCGAGUA--UUCAUCAUUCA---------AGUGAUGCACUUGCAUUAUUCAUGCACACUCCAUCAGCUGCGA----CAUUUGAUUGCCGCUCUUGGCCCAUUA ((((--((((........))))))--))((((((...---------.))))))....(((((.....)))))....(((..(((.((((----(....).)))).)))..)))....... ( -24.90) >DroSec_CAF1 40541 103 - 1 AGUG--CUCGCCUACUUUUGAGUA--UUCAUCAUUCA---------AGUGAUGCACUUGCAUUAUUCAUGCACACUCCAUCAGCUGCGA----CAUUUGAUUGCCGCUCUUGGCCCAUUA .(((--(..........((((((.--......)))))---------)(((((((....)))))))....))))...(((..(((.((((----(....).)))).)))..)))....... ( -24.90) >DroEre_CAF1 16194 112 - 1 AGUG--CUCGCCUACUUUUGAGUA--UUCAUCAUUGAUCAUUCAUCAGUGAUGCAUUCGCAUUAUUCAUUCACACGCCAUCAGCUGCGA----CAUUUGAUUGCCGCUCUUGGCCCUUUU ((((--((((........))))))--))((((((((((.....))))))))))......................((((..(((.((((----(....).)))).)))..))))...... ( -29.90) >DroYak_CAF1 17589 103 - 1 AGUG--CUCGCCUAGUUUUGAGUA--UUCAUCAUUCA---------GGUGAUGCAUUCGCAUUAUUCAUUCACACACCAUCAGCUGCGA----CAUUUGAUAGCCUUUCUUGGCCCAUUA .(((--((((((((((..((....--..))..))).)---------))))).))))(((((.......................)))))----.........(((......)))...... ( -21.10) >DroAna_CAF1 10777 102 - 1 ACUAAACUCAUAUCAUUCUGAGUAGAGCUAUCAAUCA---------AAAGCAACUCACACAUAAUUCAUUCACACGCCAUCGACUCUAAACGCUUUCUGA--------CUCGUCUCAUU- .....(((((........))))).(((.......(((---------(((((........................................))))).)))--------.....)))...- ( -8.40) >consensus AGUG__CUCGCCUACUUUUGAGUA__UUCAUCAUUCA_________AGUGAUGCACUCGCAUUAUUCAUUCACACGCCAUCAGCUGCGA____CAUUUGAUUGCCGCUCUUGGCCCAUUA ......((((........))))........................((((((((....))))))))....................................(((......)))...... ( -6.32 = -7.48 + 1.16)

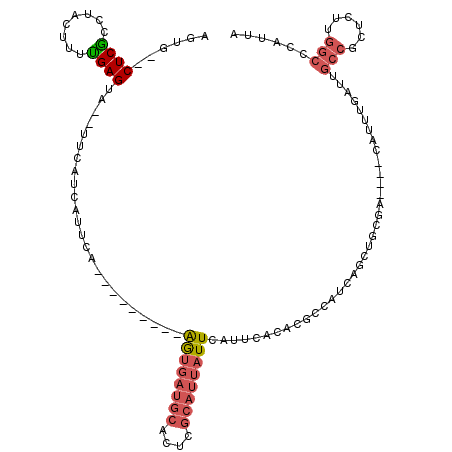

| Location | 21,853,598 – 21,853,693 |
|---|---|
| Length | 95 |
| Sequences | 5 |
| Columns | 113 |
| Reading direction | reverse |
| Mean pairwise identity | 72.26 |
| Mean single sequence MFE | -18.52 |
| Consensus MFE | -5.50 |
| Energy contribution | -6.50 |
| Covariance contribution | 1.00 |
| Combinations/Pair | 1.31 |
| Mean z-score | -1.93 |
| Structure conservation index | 0.30 |
| SVM decision value | 0.87 |
| SVM RNA-class probability | 0.871070 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 21853598 95 - 22224390 AAAUGU-----CUGCAUCCAUCGAGAUCCCUUCAGUG--CUCGCCUACUUUCGAGUA--UUCAUCAUUCA---------AGUGAUGCACUUGCAUUAUUCAUGCACACUCCAU ....((-----.(((((.......((.......((((--((((........))))))--))......)).---------((((((((....)))))))).))))).))..... ( -20.32) >DroSec_CAF1 40577 95 - 1 AAAUGA-----CUGCAUCCAUCGAGAACCCUUCAGUG--CUCGCCUACUUUUGAGUA--UUCAUCAUUCA---------AGUGAUGCACUUGCAUUAUUCAUGCACACUCCAU ......-----.(((((((..((((.((......)).--)))).......((((((.--......)))))---------)).))))))..(((((.....)))))........ ( -17.80) >DroEre_CAF1 16230 104 - 1 AAAUGG-----CUGCAUCCAGCGCUCUCCCUUCAGUG--CUCGCCUACUUUUGAGUA--UUCAUCAUUGAUCAUUCAUCAGUGAUGCAUUCGCAUUAUUCAUUCACACGCCAU ..((((-----(.((.....))((.........((((--((((........))))))--))((((((((((.....)))))))))).....))...............))))) ( -26.10) >DroYak_CAF1 17625 98 - 1 AAAUGC--AUUCUGCAUCCAUCACGCUCCCUUCAGUG--CUCGCCUAGUUUUGAGUA--UUCAUCAUUCA---------GGUGAUGCAUUCGCAUUAUUCAUUCACACACCAU .(((((--.....((.........)).......((((--((((((((((..((....--..))..))).)---------))))).))))).)))))................. ( -19.10) >DroAna_CAF1 10808 102 - 1 AGAUUCCGAUUCCGAUUCCG--AGACUCCAAUCACUAAACUCAUAUCAUUCUGAGUAGAGCUAUCAAUCA---------AAAGCAACUCACACAUAAUUCAUUCACACGCCAU .((((..((.((((....))--.)).)).)))).....(((((........)))))...(((........---------..)))............................. ( -9.30) >consensus AAAUGC_____CUGCAUCCAUCGAGCUCCCUUCAGUG__CUCGCCUACUUUUGAGUA__UUCAUCAUUCA_________AGUGAUGCACUCGCAUUAUUCAUUCACACGCCAU .............((((......................((((........))))........................((((((((....)))))))).))))......... ( -5.50 = -6.50 + 1.00)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 13:09:22 2006