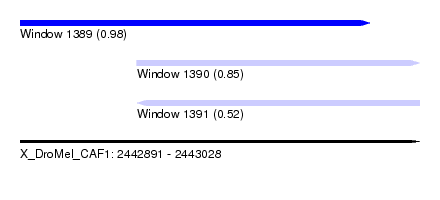

| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 2,442,891 – 2,443,028 |
| Length | 137 |
| Max. P | 0.980656 |

| Location | 2,442,891 – 2,443,011 |
|---|---|
| Length | 120 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 98.06 |
| Mean single sequence MFE | -42.83 |
| Consensus MFE | -41.03 |
| Energy contribution | -41.37 |
| Covariance contribution | 0.33 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.99 |
| Structure conservation index | 0.96 |
| SVM decision value | 1.87 |
| SVM RNA-class probability | 0.980656 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 2442891 120 + 22224390 UUAAACUUCCACUUCUAACCCUCAGCCGCUGAUAAACUAAUCAAAAUCUGGGGAGCAUACGAUGGCAAGUUCGAGAAGACCAUUUCGGGCCACAAGCUGGGCAUCAGCGAUGUGGCCUGG ...........(((((...((((((....((((......))))....))))))......(((........)))))))).......(((((((((.((((.....))))..))))))))). ( -39.90) >DroSec_CAF1 3802 120 + 1 UUAAACUUCCACUUCUACUCCUCAGCCGCUGAUAAACUAAUCAAAAUCUGGGGAGCCUACGAUGGCAAGUUCGAGAAGACCAUUUCGGGCCACAAGCUGGGCAUCAGCGAUGUGGCCUGG ...........(((((..(((((((....((((......))))....)))))))(((......))).......))))).......(((((((((.((((.....))))..))))))))). ( -44.60) >DroSim_CAF1 5955 120 + 1 UUAAACUUCCACUUCUAUCCCUCAGCCGCUGAUAAACUAAUCAAAAUCUGGGGAGCCUACGAUGGCAAGUUCGAGAAGACCAUUUCGGGCCACAAGCUGGGCAUCAGCGAUGUGGCCUGG ...........(((((.((((.(((....((((......))))....)))))))(((......))).......))))).......(((((((((.((((.....))))..))))))))). ( -44.00) >consensus UUAAACUUCCACUUCUAACCCUCAGCCGCUGAUAAACUAAUCAAAAUCUGGGGAGCCUACGAUGGCAAGUUCGAGAAGACCAUUUCGGGCCACAAGCUGGGCAUCAGCGAUGUGGCCUGG ...........(((((...((((((....((((......))))....)))))).(((......))).......))))).......(((((((((.((((.....))))..))))))))). (-41.03 = -41.37 + 0.33)

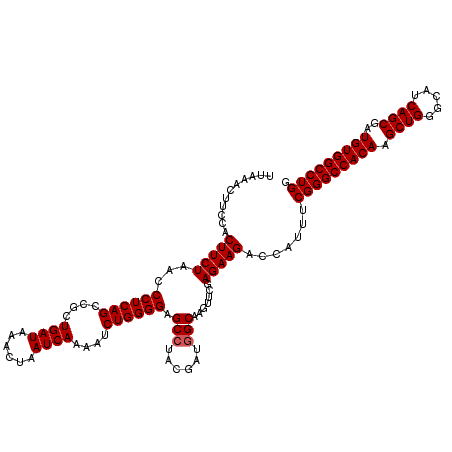
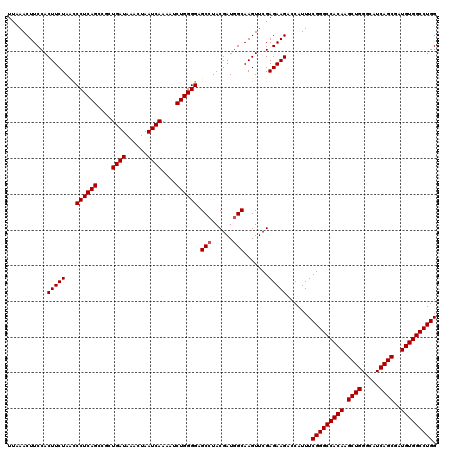
| Location | 2,442,931 – 2,443,028 |
|---|---|
| Length | 97 |
| Sequences | 3 |
| Columns | 97 |
| Reading direction | forward |
| Mean pairwise identity | 99.31 |
| Mean single sequence MFE | -36.57 |
| Consensus MFE | -35.50 |
| Energy contribution | -35.83 |
| Covariance contribution | 0.33 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.95 |
| Structure conservation index | 0.97 |
| SVM decision value | 0.77 |
| SVM RNA-class probability | 0.845234 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 2442931 97 + 22224390 UCAAAAUCUGGGGAGCAUACGAUGGCAAGUUCGAGAAGACCAUUUCGGGCCACAAGCUGGGCAUCAGCGAUGUGGCCUGGAGCUCAGACUCGCGACU ......((((((((((...(((........)))..........(((((((((((.((((.....))))..)))))))))))))))...)))).)).. ( -34.50) >DroSec_CAF1 3842 97 + 1 UCAAAAUCUGGGGAGCCUACGAUGGCAAGUUCGAGAAGACCAUUUCGGGCCACAAGCUGGGCAUCAGCGAUGUGGCCUGGAGCUCAGACUCGCGACU ......((((((..(((......)))................((((((((((((.((((.....))))..))))))))))))))))))......... ( -37.60) >DroSim_CAF1 5995 97 + 1 UCAAAAUCUGGGGAGCCUACGAUGGCAAGUUCGAGAAGACCAUUUCGGGCCACAAGCUGGGCAUCAGCGAUGUGGCCUGGAGCUCAGACUCGCGACU ......((((((..(((......)))................((((((((((((.((((.....))))..))))))))))))))))))......... ( -37.60) >consensus UCAAAAUCUGGGGAGCCUACGAUGGCAAGUUCGAGAAGACCAUUUCGGGCCACAAGCUGGGCAUCAGCGAUGUGGCCUGGAGCUCAGACUCGCGACU ......((((((..(((......)))................((((((((((((.((((.....))))..))))))))))))))))))......... (-35.50 = -35.83 + 0.33)

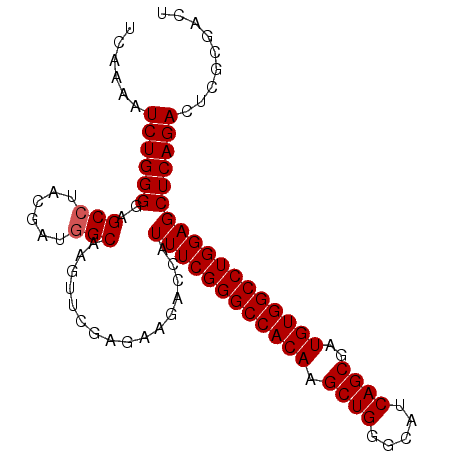
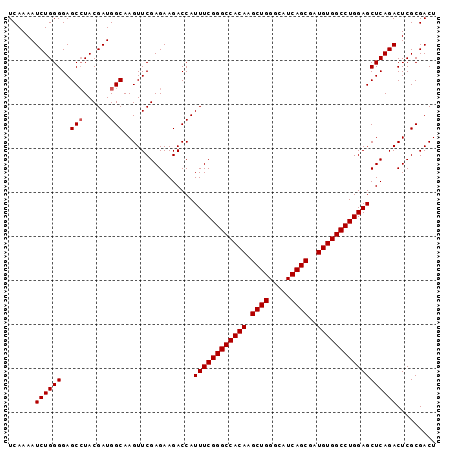
| Location | 2,442,931 – 2,443,028 |
|---|---|
| Length | 97 |
| Sequences | 3 |
| Columns | 97 |
| Reading direction | reverse |
| Mean pairwise identity | 99.31 |
| Mean single sequence MFE | -32.77 |
| Consensus MFE | -31.70 |
| Energy contribution | -31.70 |
| Covariance contribution | -0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.44 |
| Structure conservation index | 0.97 |
| SVM decision value | -0.02 |
| SVM RNA-class probability | 0.521083 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 2442931 97 - 22224390 AGUCGCGAGUCUGAGCUCCAGGCCACAUCGCUGAUGCCCAGCUUGUGGCCCGAAAUGGUCUUCUCGAACUUGCCAUCGUAUGCUCCCCAGAUUUUGA ....((((((.((((..((((((((((..((((.....)))).))))))).....)))....)))).))))))........................ ( -31.70) >DroSec_CAF1 3842 97 - 1 AGUCGCGAGUCUGAGCUCCAGGCCACAUCGCUGAUGCCCAGCUUGUGGCCCGAAAUGGUCUUCUCGAACUUGCCAUCGUAGGCUCCCCAGAUUUUGA ((((((((((.((((..((((((((((..((((.....)))).))))))).....)))....)))).)))))).......((....)).)))).... ( -33.30) >DroSim_CAF1 5995 97 - 1 AGUCGCGAGUCUGAGCUCCAGGCCACAUCGCUGAUGCCCAGCUUGUGGCCCGAAAUGGUCUUCUCGAACUUGCCAUCGUAGGCUCCCCAGAUUUUGA ((((((((((.((((..((((((((((..((((.....)))).))))))).....)))....)))).)))))).......((....)).)))).... ( -33.30) >consensus AGUCGCGAGUCUGAGCUCCAGGCCACAUCGCUGAUGCCCAGCUUGUGGCCCGAAAUGGUCUUCUCGAACUUGCCAUCGUAGGCUCCCCAGAUUUUGA ....((((((.((((..((((((((((..((((.....)))).))))))).....)))....)))).))))))........................ (-31.70 = -31.70 + -0.00)
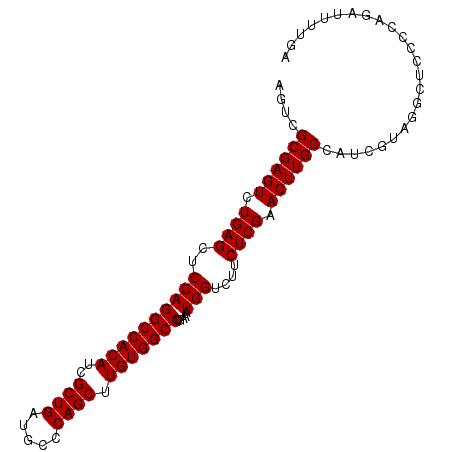


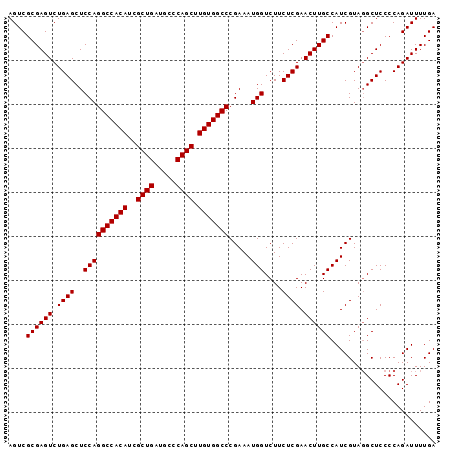
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:55:34 2006