
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 21,402,539 – 21,402,647 |
| Length | 108 |
| Max. P | 0.660734 |

| Location | 21,402,539 – 21,402,647 |
|---|---|
| Length | 108 |
| Sequences | 4 |
| Columns | 114 |
| Reading direction | forward |
| Mean pairwise identity | 89.48 |
| Mean single sequence MFE | -26.43 |
| Consensus MFE | -17.72 |
| Energy contribution | -18.97 |
| Covariance contribution | 1.25 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.44 |
| Structure conservation index | 0.67 |
| SVM decision value | 0.27 |
| SVM RNA-class probability | 0.660734 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 21402539 108 + 22224390 AUCGAAAUACAUACGUCGGCAUUAUCCGAC-GACUAAAGCCGUCUUUCAA----GCCC-CACCGAUGGAAUUUUACUUGGCAAAUUUCAUCCAAGUUAGUGCUCGGCAAUAACA .............((((((......)))))-)......((((.......(----(((.-.((.((((((((((........))))))))))...))..).)))))))....... ( -24.61) >DroSec_CAF1 52832 105 + 1 AUCGAAAU----ACGUCGGCAUUAUCCGACUUACUAAAGCCGCCUUUCAA----GCAC-CACCGAUGGAAUUUAACUUUGCAAAUUUCAUCCAAGUUAGUGCUCGGCAAUAACA ........----..(((((......)))))........((((.......(----((((-.((.((((((((((........))))))))))...))..)))))))))....... ( -25.71) >DroSim_CAF1 20449 105 + 1 AUCGAAAU----ACGUCGGCAUUAUCCGACUUACUAAAGCCGCCUUUCAA----GCAC-CACCGAUGGAAUUUUACUUUGCAAAUUUCAUCCAAGUUAGUGCUCGGCAAUAACA ........----..(((((......)))))........((((.......(----((((-.((.((((((((((........))))))))))...))..)))))))))....... ( -25.51) >DroYak_CAF1 41241 109 + 1 AUCGAAAU----ACGUCGGCCUUAUCCGCC-GACUAAAGCCGCCUUUCAGCCCAGCUCCGACCACUGGAAUUUAACUUGGCAAAUUUCAUCCAAGUUAGUGCUCGGCAAUAACA ........----..((((((.......)))-)))...............(((.(((((((.....))))..(((((((((..........))))))))).))).)))....... ( -29.90) >consensus AUCGAAAU____ACGUCGGCAUUAUCCGAC_GACUAAAGCCGCCUUUCAA____GCAC_CACCGAUGGAAUUUAACUUGGCAAAUUUCAUCCAAGUUAGUGCUCGGCAAUAACA ..............(((((......)))))........((((............((((..((.((((((((((........))))))))))...))..)))).))))....... (-17.72 = -18.97 + 1.25)

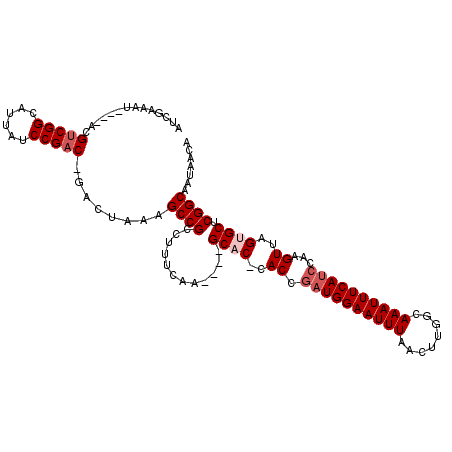
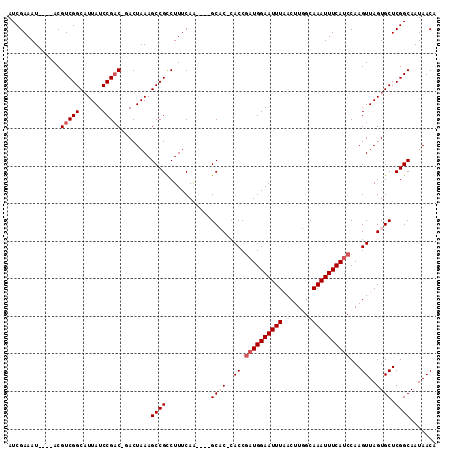
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 13:08:34 2006