
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 21,378,487 – 21,378,606 |
| Length | 119 |
| Max. P | 0.989818 |

| Location | 21,378,487 – 21,378,587 |
|---|---|
| Length | 100 |
| Sequences | 5 |
| Columns | 115 |
| Reading direction | forward |
| Mean pairwise identity | 75.39 |
| Mean single sequence MFE | -35.26 |
| Consensus MFE | -16.38 |
| Energy contribution | -16.22 |
| Covariance contribution | -0.16 |
| Combinations/Pair | 1.21 |
| Mean z-score | -2.17 |
| Structure conservation index | 0.46 |
| SVM decision value | 1.35 |
| SVM RNA-class probability | 0.945192 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 21378487 100 + 22224390 GGC---UGGUGGCUGGUGG--------UAGAAGGUCUG-G---GCGGACACUCUGGCCAAAGGACAACAGCAUUUCAUUUGUUGUCCUUCACAUUCCCUUCUCACACUUGGCACA ...---..(((.(.((((.--------.((((((..((-(---.(((.....))).)))(((((((((((........))))))))))).......))))))..)))).).))). ( -40.90) >DroSec_CAF1 24139 100 + 1 GGC---UGGUGGCUGAUGG--------UGGAAGGGCUG-G---GCGGACACUCUGGCCACAGGACAACAGCAUUUCAUUUGUUGUCCUUCACAUUCCCUUCUCACACUUGGCACA ...---..(((.(..(..(--------((((((((.((-(---.(((.....))).))).((((((((((........)))))))))).......)))))).)))..)..)))). ( -39.10) >DroEre_CAF1 15140 81 + 1 --------------------------CUGGGAGG-----G---CAGGACACUCUGGCCACAGGACAACAGCAUUUCAUUUGUUGUCCUUCACAUUCCCUUUUCACACUUGGCACA --------------------------..((((((-----.---((((....)))).))..((((((((((........))))))))))......))))................. ( -26.40) >DroYak_CAF1 15416 107 + 1 GGCGAGUGGUGGCUGGUGGCUGGUGGCUGGGAGG-----G---CAGGACACUCUGGCCACAGGACAACAGCAUUUCAUUUGUUGUCCUUCACGUUCCUUUUUCGCACUUGGCACA (.((((((.(((..((..(((((((((..((..(-----.---.....)..))..)))))((((((((((........)))))))))).)).))..))...))))))))).)... ( -41.10) >DroAna_CAF1 25429 108 + 1 ------UGGGAACUGGAAACUGGAAACUGGGCAU-CCUGGCACUAGGACACUCUGGCCACAGGACAACAGCAUUUCAUUUGUUGUCCUUCACAUUCUCUUUUCACACUUGGCACG ------.(((((.(((((..(((...(.(((..(-((((....)))))..))).).)))..(((((((((........)))))))))))).)))))))................. ( -28.80) >consensus GGC___UGGUGGCUGGUGG_______CUGGGAGG_C___G___CAGGACACUCUGGCCACAGGACAACAGCAUUUCAUUUGUUGUCCUUCACAUUCCCUUUUCACACUUGGCACA ............................((((((...........((.(.....).))..((((((((((........))))))))))........))))))............. (-16.38 = -16.22 + -0.16)


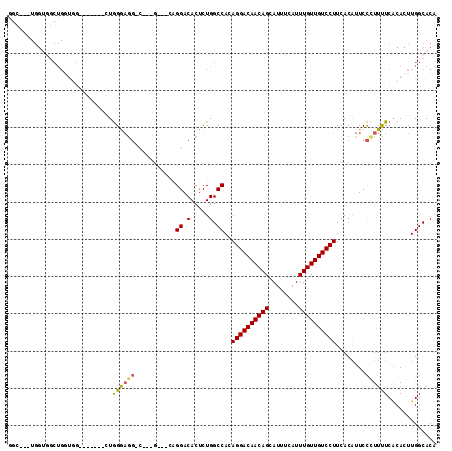
| Location | 21,378,512 – 21,378,606 |
|---|---|
| Length | 94 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 77.51 |
| Mean single sequence MFE | -27.50 |
| Consensus MFE | -18.09 |
| Energy contribution | -19.07 |
| Covariance contribution | 0.97 |
| Combinations/Pair | 1.18 |
| Mean z-score | -3.00 |
| Structure conservation index | 0.66 |
| SVM decision value | 2.18 |
| SVM RNA-class probability | 0.989818 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 21378512 94 + 22224390 UG-G---GCGGACACUCUGGCCAAAGGACAACAGCAUUUCAUUUGUUGUCCUUCACAUUCCCUUCUCACACUUGGCACA---UAAACAAAGGGAAUGUCCC------------------- ((-(---.(((.....))).)))(((((((((((........))))))))))).((((((((((...............---......))))))))))...------------------- ( -32.20) >DroVir_CAF1 7498 79 + 1 ---G---AAUGA--------------GAC--AAACAUUUCAUUUGUUGUCCUUCACAUUCUCUUUUCACACUUUGCGCACCGAAAACAAAGCGAAUGCCCC------------------- ---(---((.(.--------------(((--((.((.......))))))))))).(((((.((((......((((.....))))...)))).)))))....------------------- ( -10.80) >DroSec_CAF1 24164 94 + 1 UG-G---GCGGACACUCUGGCCACAGGACAACAGCAUUUCAUUUGUUGUCCUUCACAUUCCCUUCUCACACUUGGCACA---UAAACAAAGGGAAUGUCCC------------------- ((-(---.(((.....))).))).((((((((((........))))))))))..((((((((((...............---......))))))))))...------------------- ( -31.00) >DroEre_CAF1 15148 92 + 1 ---G---CAGGACACUCUGGCCACAGGACAACAGCAUUUCAUUUGUUGUCCUUCACAUUCCCUUUUCACACUUGGCACA---UAAACAAAGGGAAUGUCCU------------------- ---.---.(((((...........((((((((((........))))))))))....(((((((((..............---.....))))))))))))))------------------- ( -27.61) >DroYak_CAF1 15450 92 + 1 ---G---CAGGACACUCUGGCCACAGGACAACAGCAUUUCAUUUGUUGUCCUUCACGUUCCUUUUUCGCACUUGGCACA---UAAACAAAGGGAAUGUCCU------------------- ---.---.((((((((((.((((.((((((((((........))))))))))...((.........))....))))...---.......))))..))))))------------------- ( -25.40) >DroAna_CAF1 25458 117 + 1 CUGGCACUAGGACACUCUGGCCACAGGACAACAGCAUUUCAUUUGUUGUCCUUCACAUUCUCUUUUCACACUUGGCACG---UAAACAAAGGGAAUGGCCCUCGGUCUCGGUCUCGGUCC ((((.((..((((.....((((..((((((((((........))))))))))....(((((((((....((.......)---)....)))))))))))))....))))..)).))))... ( -38.00) >consensus ___G___CAGGACACUCUGGCCACAGGACAACAGCAUUUCAUUUGUUGUCCUUCACAUUCCCUUUUCACACUUGGCACA___UAAACAAAGGGAAUGUCCC___________________ .........((.............((((((((((........))))))))))...((((((((((......................)))))))))).)).................... (-18.09 = -19.07 + 0.97)

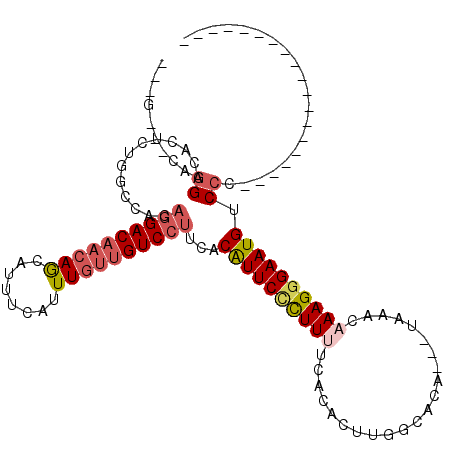

| Location | 21,378,512 – 21,378,606 |
|---|---|
| Length | 94 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 77.51 |
| Mean single sequence MFE | -30.10 |
| Consensus MFE | -17.98 |
| Energy contribution | -19.20 |
| Covariance contribution | 1.22 |
| Combinations/Pair | 1.16 |
| Mean z-score | -1.98 |
| Structure conservation index | 0.60 |
| SVM decision value | 1.02 |
| SVM RNA-class probability | 0.902450 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 21378512 94 - 22224390 -------------------GGGACAUUCCCUUUGUUUA---UGUGCCAAGUGUGAGAAGGGAAUGUGAAGGACAACAAAUGAAAUGCUGUUGUCCUUUGGCCAGAGUGUCCGC---C-CA -------------------((((((((((((((..(((---..(.....)..))))))))))))))(((((((((((..........)))))))))))((.(.....).)).)---)-). ( -35.30) >DroVir_CAF1 7498 79 - 1 -------------------GGGGCAUUCGCUUUGUUUUCGGUGCGCAAAGUGUGAAAAGAGAAUGUGAAGGACAACAAAUGAAAUGUUU--GUC--------------UCAUU---C--- -------------------(((((.(((((...((((((....(((.....)))....)))))))))))(((((..........)))))--)))--------------))...---.--- ( -17.50) >DroSec_CAF1 24164 94 - 1 -------------------GGGACAUUCCCUUUGUUUA---UGUGCCAAGUGUGAGAAGGGAAUGUGAAGGACAACAAAUGAAAUGCUGUUGUCCUGUGGCCAGAGUGUCCGC---C-CA -------------------((((((((((((((..(((---..(.....)..))))))))))))))..(((((((((..........)))))))))..((.(.....).)).)---)-). ( -33.20) >DroEre_CAF1 15148 92 - 1 -------------------AGGACAUUCCCUUUGUUUA---UGUGCCAAGUGUGAAAAGGGAAUGUGAAGGACAACAAAUGAAAUGCUGUUGUCCUGUGGCCAGAGUGUCCUG---C--- -------------------((((((((((((((.((((---..(.....)..)))))))))...((.((((((((((..........))))))))).).))..))))))))).---.--- ( -31.10) >DroYak_CAF1 15450 92 - 1 -------------------AGGACAUUCCCUUUGUUUA---UGUGCCAAGUGCGAAAAAGGAACGUGAAGGACAACAAAUGAAAUGCUGUUGUCCUGUGGCCAGAGUGUCCUG---C--- -------------------((((((((((((((.....---((..(...)..))..)))))...((.((((((((((..........))))))))).).))..))))))))).---.--- ( -30.70) >DroAna_CAF1 25458 117 - 1 GGACCGAGACCGAGACCGAGGGCCAUUCCCUUUGUUUA---CGUGCCAAGUGUGAAAAGAGAAUGUGAAGGACAACAAAUGAAAUGCUGUUGUCCUGUGGCCAGAGUGUCCUAGUGCCAG (((((.....((....))..((((((...((((.((((---((.......)))))).)))).......(((((((((..........)))))))))))))))...).))))......... ( -32.80) >consensus ___________________GGGACAUUCCCUUUGUUUA___UGUGCCAAGUGUGAAAAGGGAAUGUGAAGGACAACAAAUGAAAUGCUGUUGUCCUGUGGCCAGAGUGUCCUC___C___ ....................(((((((((((((..........(((.....))).)))))))))))..(((((((((..........))))))))).............))......... (-17.98 = -19.20 + 1.22)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 13:08:17 2006