
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 21,290,752 – 21,290,852 |
| Length | 100 |
| Max. P | 0.987009 |

| Location | 21,290,752 – 21,290,852 |
|---|---|
| Length | 100 |
| Sequences | 4 |
| Columns | 102 |
| Reading direction | forward |
| Mean pairwise identity | 85.83 |
| Mean single sequence MFE | -29.98 |
| Consensus MFE | -26.27 |
| Energy contribution | -26.52 |
| Covariance contribution | 0.25 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.96 |
| Structure conservation index | 0.88 |
| SVM decision value | 1.15 |
| SVM RNA-class probability | 0.922359 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 21290752 100 + 22224390 ACACAC-CCACACAUGGCAUGGUGUUAGG-GGUGUUGGGGCCUCACCAACACACACAUAGCCACGCAUGAAUGCCUGCAAAGUAUGCAAAGUUUAUUCAUGA ......-.....((((((.((((......-.(((((((.......))))))).......)))).))((((((...((((.....))))..)))))).)))). ( -28.84) >DroSec_CAF1 31112 102 + 1 ACACACCCCACACAUGGCAUGGUGUUAGGGGGUGUUGGGGCCUCACCAACACACACAUAGCCACGCAUGAAUGCCUGCAAAGUAUGCAAAGUUUAUUCAUGA .....((((((((........))))..))))(((((((.......))))))).............((((((((..((((.....)))).....)))))))). ( -30.90) >DroEre_CAF1 30324 84 + 1 ---------A----GGG---GGUGUUGG--GGUGUUGGGGCUGCACCAACACACACACAGCCGCGCAUGAAUGCCUGCAAAGUAUGCAAAGUUUAUUCAUGA ---------.----(.(---(.(((((.--.(((((((.......))))))).)).))).)).).((((((((..((((.....)))).....)))))))). ( -29.50) >DroYak_CAF1 32743 94 + 1 -------ACAGACAUGGCAUGGUGUUGGG-GGUGUUGGGGCUGCACCAACACACACAUAGCCACGCAUGAAUGCCUGCAAAGUAUGCAAAGUUUAUUCAUGA -------.......(((((((((((.(..-(((((.......)))))..))))).))).))))..((((((((..((((.....)))).....)))))))). ( -30.70) >consensus _______CCACACAUGGCAUGGUGUUAGG_GGUGUUGGGGCCGCACCAACACACACAUAGCCACGCAUGAAUGCCUGCAAAGUAUGCAAAGUUUAUUCAUGA ...............(((...((((......(((((((.......)))))))))))...)))...((((((((..((((.....)))).....)))))))). (-26.27 = -26.52 + 0.25)

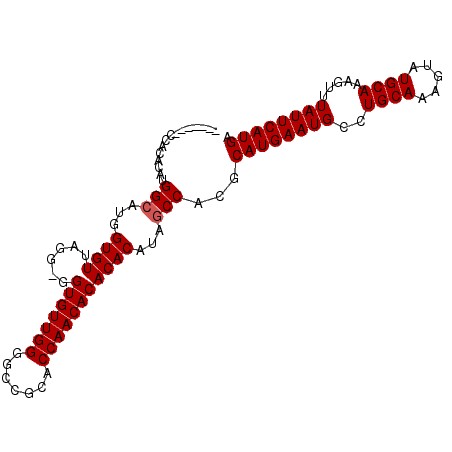

| Location | 21,290,752 – 21,290,852 |
|---|---|
| Length | 100 |
| Sequences | 4 |
| Columns | 102 |
| Reading direction | reverse |
| Mean pairwise identity | 85.83 |
| Mean single sequence MFE | -32.40 |
| Consensus MFE | -26.50 |
| Energy contribution | -27.50 |
| Covariance contribution | 1.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.71 |
| Structure conservation index | 0.82 |
| SVM decision value | 2.06 |
| SVM RNA-class probability | 0.987009 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 21290752 100 - 22224390 UCAUGAAUAAACUUUGCAUACUUUGCAGGCAUUCAUGCGUGGCUAUGUGUGUGUUGGUGAGGCCCCAACACC-CCUAACACCAUGCCAUGUGUGG-GUGUGU .............(((((.....)))))(((((((((((((((...(((((((((((.(....)))))))).-....))))...)))))))))))-)))).. ( -39.00) >DroSec_CAF1 31112 102 - 1 UCAUGAAUAAACUUUGCAUACUUUGCAGGCAUUCAUGCGUGGCUAUGUGUGUGUUGGUGAGGCCCCAACACCCCCUAACACCAUGCCAUGUGUGGGGUGUGU .............(((((.....)))))(((((((((((((((...(((((((((((.(....))))))))......))))...))))))))).)))))).. ( -34.80) >DroEre_CAF1 30324 84 - 1 UCAUGAAUAAACUUUGCAUACUUUGCAGGCAUUCAUGCGCGGCUGUGUGUGUGUUGGUGCAGCCCCAACACC--CCAACACC---CCC----U--------- .(((((((....((((((.....)))))).)))))))...((..((((..(((((((.......))))))).--...)))).---.))----.--------- ( -25.10) >DroYak_CAF1 32743 94 - 1 UCAUGAAUAAACUUUGCAUACUUUGCAGGCAUUCAUGCGUGGCUAUGUGUGUGUUGGUGCAGCCCCAACACC-CCCAACACCAUGCCAUGUCUGU------- .(((((((....((((((.....)))))).)))))))((((((...(((((((((((.......))))))).-....))))...)))))).....------- ( -30.70) >consensus UCAUGAAUAAACUUUGCAUACUUUGCAGGCAUUCAUGCGUGGCUAUGUGUGUGUUGGUGAAGCCCCAACACC_CCCAACACCAUGCCAUGUGUGG_______ .(((((((....((((((.....)))))).)))))))((((((...(((((((((((.......)))))))......))))...))))))............ (-26.50 = -27.50 + 1.00)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 13:07:42 2006