
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 21,050,950 – 21,051,070 |
| Length | 120 |
| Max. P | 0.752461 |

| Location | 21,050,950 – 21,051,070 |
|---|---|
| Length | 120 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 85.14 |
| Mean single sequence MFE | -33.80 |
| Consensus MFE | -25.29 |
| Energy contribution | -25.07 |
| Covariance contribution | -0.22 |
| Combinations/Pair | 1.04 |
| Mean z-score | -2.16 |
| Structure conservation index | 0.75 |
| SVM decision value | 0.48 |
| SVM RNA-class probability | 0.752461 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
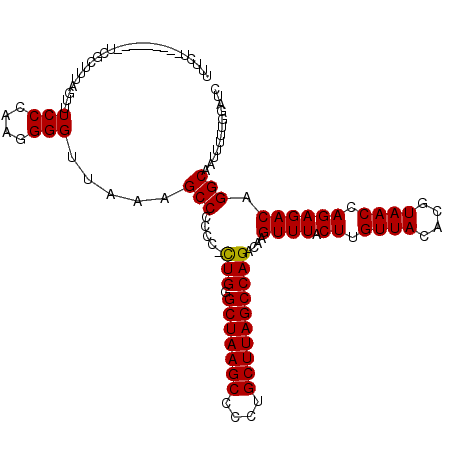
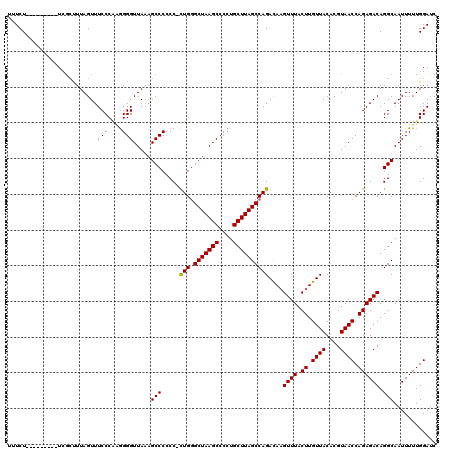
>X_DroMel_CAF1 21050950 120 + 22224390 UGUCUUGGAUUUCUUCACUUUAGUUUCCCCAAGGGUUAUAGCCCCCCCUUGGGCUAAGCCCCUGCUUAGCCAGACAAGUUUACUUGUUACACGUAACCAGAGACAGGCAAUUUUUGGAUC ((((((((.....................((((((..........))))))((((((((....)))))))).((((((....))))))........))).)))))............... ( -34.80) >DroEre_CAF1 21994 108 + 1 -UUCU---------UCGCUUUAGUUUCCCAGGGGGUUAAAGCCC-CC-CUGCGCUAAGCCCCUGCUUAGCCAGAUAAGUUUACUUGUUACACGUAACCAGAGACAGGCAAUUUUUGGAUC -....---------..((((..(((((...((((((....))))-))-(((.(((((((....))))))))))(((((....)))))............)))))))))............ ( -34.20) >DroYak_CAF1 24322 110 + 1 UUUCU---------UGGACUUAGUUUCCCGCGGGGUUAAAGCCCCCC-CUGGGCUAAGCCCCUGCUUAGCCAGACAUGUUUACUUGUUACACGUAACCAGAGACAGGCAAUUUUUGGAUC .....---------...........(((...(((((....))))).(-(((((((((((....))))))))...........((.((((....)))).))...))))........))).. ( -32.40) >consensus UUUCU_________UCGCUUUAGUUUCCCAAGGGGUUAAAGCCCCCC_CUGGGCUAAGCCCCUGCUUAGCCAGACAAGUUUACUUGUUACACGUAACCAGAGACAGGCAAUUUUUGGAUC .........................(((....))).....(((.....(((.(((((((....))))))))))....((((.((.((((....)))).)))))).)))............ (-25.29 = -25.07 + -0.22)

| Location | 21,050,950 – 21,051,070 |
|---|---|
| Length | 120 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 85.14 |
| Mean single sequence MFE | -34.97 |
| Consensus MFE | -25.97 |
| Energy contribution | -26.63 |
| Covariance contribution | 0.67 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.79 |
| Structure conservation index | 0.74 |
| SVM decision value | 0.01 |
| SVM RNA-class probability | 0.537765 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
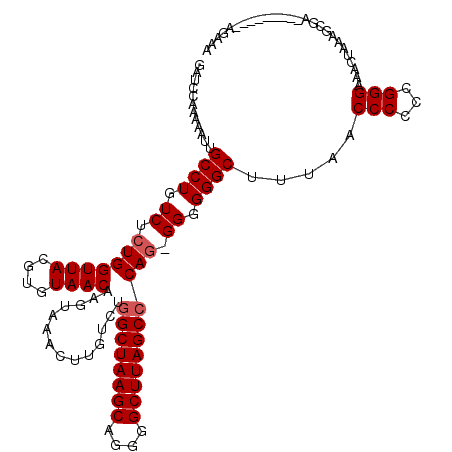
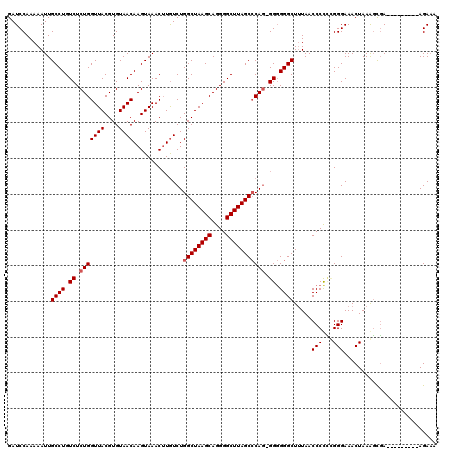
>X_DroMel_CAF1 21050950 120 - 22224390 GAUCCAAAAAUUGCCUGUCUCUGGUUACGUGUAACAAGUAAACUUGUCUGGCUAAGCAGGGGCUUAGCCCAAGGGGGGGCUAUAACCCUUGGGGAAACUAAAGUGAAGAAAUCCAAGACA ................((((.(((((((.....(((((....)))))..((((((((....))))))))((((((..(....)..))))))((....))...))))......))))))). ( -35.60) >DroEre_CAF1 21994 108 - 1 GAUCCAAAAAUUGCCUGUCUCUGGUUACGUGUAACAAGUAAACUUAUCUGGCUAAGCAGGGGCUUAGCGCAG-GG-GGGCUUUAACCCCCUGGGAAACUAAAGCGA---------AGAA- ..(((.......((..(((((((.(((.((.....(((....))).....))))).)))))))...)).(((-((-((.......))))))))))...........---------....- ( -33.50) >DroYak_CAF1 24322 110 - 1 GAUCCAAAAAUUGCCUGUCUCUGGUUACGUGUAACAAGUAAACAUGUCUGGCUAAGCAGGGGCUUAGCCCAG-GGGGGGCUUUAACCCCGCGGGAAACUAAGUCCA---------AGAAA ..............(((.((.(((((((((((.........))))))..))))))))))(((((((((((.(-.((((.......)))).))))...)))))))).---------..... ( -35.80) >consensus GAUCCAAAAAUUGCCUGUCUCUGGUUACGUGUAACAAGUAAACUUGUCUGGCUAAGCAGGGGCUUAGCCCAG_GGGGGGCUUUAACCCCCCGGGAAACUAAAGCGA_________AGAAA ............((((.((.(((((((....))))..............((((((((....))))))))))).)).)))).....(((...))).......................... (-25.97 = -26.63 + 0.67)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 13:05:35 2006