
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 21,014,566 – 21,014,755 |
| Length | 189 |
| Max. P | 0.999265 |

| Location | 21,014,566 – 21,014,684 |
|---|---|
| Length | 118 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 93.47 |
| Mean single sequence MFE | -37.18 |
| Consensus MFE | -32.64 |
| Energy contribution | -32.80 |
| Covariance contribution | 0.16 |
| Combinations/Pair | 1.03 |
| Mean z-score | -2.50 |
| Structure conservation index | 0.88 |
| SVM decision value | 3.47 |
| SVM RNA-class probability | 0.999265 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
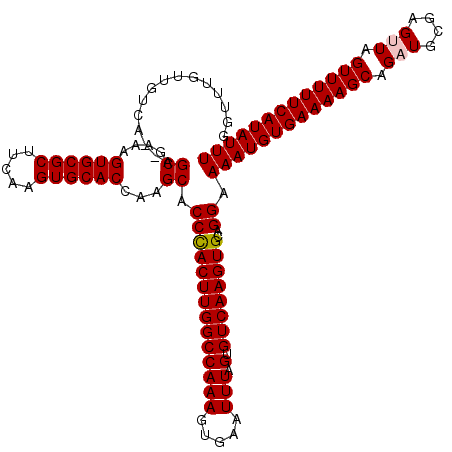
>X_DroMel_CAF1 21014566 118 - 22224390 GGUUUGUUGUCA--AGCGAAAGUGCGCUUCAAGUGCACCAAGCACCCACUUGGCCAAAGUGAAUUUAGUGUCAAGUGCAGGAAAAUGUGAAAAGCAGAUGCGAGUUAGUUUUUCAUAUUU .((((((((...--..)))..((((((.....))))))))))).((((((((((((((.....))).).))))))))..)).(((((((((((((.(((....))).))))))))))))) ( -35.40) >DroVir_CAF1 36369 120 - 1 CUGUUGUUGUCGGCAGCGAAAGUGCGCUUGAAGUGCACGGAGCACCUACUUGGCCAAAGUGAAUUUAGUGUCAAGUGCAGGAAAAUGUGAAAAGCAGCUACGAGUUAGUUUUUCAUAUUU ((.((((((....)))))).))(((((((((.((((.....)))).(((((.....))))).........)))))))))...((((((((((((((((.....))).))))))))))))) ( -38.70) >DroSec_CAF1 1388 118 - 1 GGUUUGUUGUCA--AGCGAAAGUGCGCUUCAAGUGCACCAAGCACCCACUUGGCCAAAGUGAAUUUAGUGUCAAGUGCAGGAAAAUGUGAAAAGCAGAUGCGAGUUAGUUUUUCAUAUUU .((((((((...--..)))..((((((.....))))))))))).((((((((((((((.....))).).))))))))..)).(((((((((((((.(((....))).))))))))))))) ( -35.40) >DroSim_CAF1 1054 118 - 1 GGUUUGUUGUCA--AGCGAAAGUGCGCUUCAAGUGCACCAAGCACCCACUUGGCCAAAGUGAAUUUAGUGUCAAGUGCAGGAAAAUGUGAAAAGCAGAUGCGAGUUAGUUUUUCAUAUUU .((((((((...--..)))..((((((.....))))))))))).((((((((((((((.....))).).))))))))..)).(((((((((((((.(((....))).))))))))))))) ( -35.40) >DroMoj_CAF1 145136 120 - 1 UCGUUGUUGUUGGCAGCGAAAGUGCGCUUGAAGUGCACGGAGCACCUACUUGGCCAAAGUGAAUUUAGUGUCAAGUGCAGGAAAAUGUGAAAAGCAGCUACGAGUUAGUUUUUCAUAUUU (((((((.....)))))))...(((((((((.((((.....)))).(((((.....))))).........)))))))))...((((((((((((((((.....))).))))))))))))) ( -41.00) >consensus GGUUUGUUGUCA__AGCGAAAGUGCGCUUCAAGUGCACCAAGCACCCACUUGGCCAAAGUGAAUUUAGUGUCAAGUGCAGGAAAAUGUGAAAAGCAGAUGCGAGUUAGUUUUUCAUAUUU ...............((....((((((.....))))))...)).((((((((((((((.....))).).))))))))..)).(((((((((((((.(((....))).))))))))))))) (-32.64 = -32.80 + 0.16)


| Location | 21,014,646 – 21,014,755 |
|---|---|
| Length | 109 |
| Sequences | 3 |
| Columns | 109 |
| Reading direction | forward |
| Mean pairwise identity | 95.72 |
| Mean single sequence MFE | -23.20 |
| Consensus MFE | -22.59 |
| Energy contribution | -22.37 |
| Covariance contribution | -0.22 |
| Combinations/Pair | 1.04 |
| Mean z-score | -1.85 |
| Structure conservation index | 0.97 |
| SVM decision value | 0.77 |
| SVM RNA-class probability | 0.845058 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 21014646 109 + 22224390 UGGUGCACUUGAAGCGCACUUUCGCUUGACAACAAACCCACCGUCAGCAUCGCAUCCAUAUCCACAUCCAUGGAGCGGCAUCCAUGCAGCAACAUCCACAUCCACAUCC ((.((((...((.((........(((.(((............))))))..(((.(((((..........)))))))))).))..)))).)).................. ( -23.20) >DroSec_CAF1 1468 103 + 1 UGGUGCACUUGAAGCGCACUUUCGCUUGACAACAAACCCACCGUCAGCAUCGCAUCCAUAUCCACAUCCAUGGAGCGGCAUCCACGCAGCAACAUCC------ACAUCC .(((((.....(((((......)))))(((............))).)))))((.(((((..........)))))(((.......))).)).......------...... ( -23.20) >DroSim_CAF1 1134 109 + 1 UGGUGCACUUGAAGCGCACUUUCGCUUGACAACAAACCCACCGUCAGCAUCGCAUCCAUAUCCACAUCCAUGGAGCGGCAUCCACGCAGCAACAUCCACAUCCACAUCC .(((((.....(((((......)))))(((............))).)))))((.(((((..........)))))(((.......))).))................... ( -23.20) >consensus UGGUGCACUUGAAGCGCACUUUCGCUUGACAACAAACCCACCGUCAGCAUCGCAUCCAUAUCCACAUCCAUGGAGCGGCAUCCACGCAGCAACAUCCACAUCCACAUCC .(((((.....(((((......)))))(((............))).)))))((.(((((..........)))))(((.......))).))................... (-22.59 = -22.37 + -0.22)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 13:04:27 2006