
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 21,007,247 – 21,007,409 |
| Length | 162 |
| Max. P | 0.961800 |

| Location | 21,007,247 – 21,007,342 |
|---|---|
| Length | 95 |
| Sequences | 6 |
| Columns | 119 |
| Reading direction | reverse |
| Mean pairwise identity | 79.62 |
| Mean single sequence MFE | -22.92 |
| Consensus MFE | -14.52 |
| Energy contribution | -14.85 |
| Covariance contribution | 0.33 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.52 |
| Structure conservation index | 0.63 |
| SVM decision value | 0.20 |
| SVM RNA-class probability | 0.630200 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 21007247 95 - 22224390 AAAUUAUGCAUUCAUAAAGCAGU------UGCAUUCGGAGUGCC-------AGUGCCG-AGCAUGGCCAUAAAACACUCGCUGAAUGCAAUUUUAGCCACC----------UCCACCGC ...(((((....))))).(((((------(((((((((((((..-------.(((((.-.....)).)))....)))))..)))))))))))...))....----------........ ( -22.10) >DroSec_CAF1 89001 96 - 1 AAAUUAUGCAUUCAUAAAGCAGU------UGCAUUCGGAGUGCCG------AGUACCG-AGCAUGGCCAUAAAACACUCGCUGAAUGCAAUUUUAGCCACC----------UCCACCGC ......((((((((....((...------.))....((.((((((------.....))-.))))..)).............))))))))............----------........ ( -20.60) >DroSim_CAF1 104529 96 - 1 AAAUUAUGCAUUCAUAAAGCAGU------UGCAUUCGGAGUGCCG------AGUGCCG-AGCAUGGCCAUAAAACACUCGCUGAAUGCAAUUUUAGCCACC----------UCCACCGC ......((((((((....((...------.((((((((....)))------))))).(-((..((.........)))))))))))))))............----------........ ( -24.80) >DroEre_CAF1 29723 89 - 1 AAAUUAUGCAUUCAUAAAGCAGC------UGCAUUCGGAGUGC-------------CG-AACAUGGCCAUAAAACACUCGCUGAAUGCAAUUUUCGCCACC----------ACCACCAC ......((((((((....((...------.)).....((((((-------------(.-.....)))........))))..))))))))............----------........ ( -17.40) >DroYak_CAF1 35190 99 - 1 AAAUUAUGCAUUCAUAAAGCAGU------UGCAUUCGGAGUGC-------------CG-AACAUGGCCAUAAAACACUCGCUGAAUGCAAUUUUCGCCACCUCUUACUUCCACCUCCAC ...(((((....))))).(((((------((((((((((((((-------------(.-.....)))........))))..)))))))))))...))...................... ( -18.50) >DroAna_CAF1 41213 104 - 1 AAAUUAUGCAUUCAUAAAGCAGCUCCAGCUGCAGUCGCAGUCGCAGUUGCAGUUGCAGUCGCAGGACCAUAAAGCACUCGCUGAAUGCAAUUUUCAUU--C----------CCCAC--- ......(((((((.....((.(((.(((((((((..((....))..))))))))).))).))..........(((....)))))))))).........--.----------.....--- ( -34.10) >consensus AAAUUAUGCAUUCAUAAAGCAGU______UGCAUUCGGAGUGCC_______AGUGCCG_AGCAUGGCCAUAAAACACUCGCUGAAUGCAAUUUUAGCCACC__________UCCACCGC ......((((((((....(((........))).....(((((................................)))))..)))))))).............................. (-14.52 = -14.85 + 0.33)

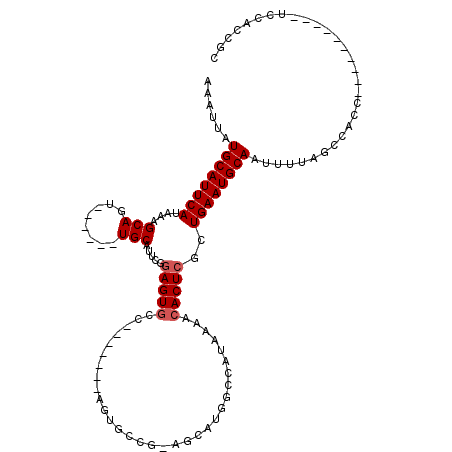

| Location | 21,007,309 – 21,007,409 |
|---|---|
| Length | 100 |
| Sequences | 6 |
| Columns | 111 |
| Reading direction | forward |
| Mean pairwise identity | 91.51 |
| Mean single sequence MFE | -31.23 |
| Consensus MFE | -25.88 |
| Energy contribution | -25.63 |
| Covariance contribution | -0.25 |
| Combinations/Pair | 1.10 |
| Mean z-score | -2.93 |
| Structure conservation index | 0.83 |
| SVM decision value | 1.53 |
| SVM RNA-class probability | 0.961800 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 21007309 100 + 22224390 UCCGAAUGCA------ACUGCUUUAUGAAUGCAUAAUUUUAU--UAUGCACAAAUUCUAUUUACAACGCGACACGUUGCGU---GAACGGUUCAUGAGGCGCAAAUGCGAG ......((((------..(((((((((((((((((((...))--)))))................((((((....))))))---.....))))))))))))....)))).. ( -29.30) >DroSec_CAF1 89064 100 + 1 UCCGAAUGCA------ACUGCUUUAUGAAUGCAUAAUUUUAU--UAUGCACAAAAUCUAUUUACAACGCGACACGUUGCGU---GAACGGUUCAUGAGGCGCAAAUGCGAG ......((((------..(((((((((((((((((((...))--)))))................((((((....))))))---.....))))))))))))....)))).. ( -29.30) >DroSim_CAF1 104592 100 + 1 UCCGAAUGCA------ACUGCUUUAUGAAUGCAUAAUUUUAU--UAUGCACAAAUUCUAUUUACAACGCGACACGUUGCGU---GAACGGUUCAUGAGGCGCAAAUGCGAG ......((((------..(((((((((((((((((((...))--)))))................((((((....))))))---.....))))))))))))....)))).. ( -29.30) >DroEre_CAF1 29779 100 + 1 UCCGAAUGCA------GCUGCUUUAUGAAUGCAUAAUUUUAU--UAUGCACAAAUUCUAUUUACAACGCGACACGUUGCGU---GAACGGUUCAUGAGGCGCAAGUGCGAG ......((((------((.((((((((((((((((((...))--)))))................((((((....))))))---.....)))))))))))))...)))).. ( -30.60) >DroYak_CAF1 35256 103 + 1 UCCGAAUGCA------ACUGCUUUAUGAAUGCAUAAUUUUAU--UAUGCACAAAUUCUAUUUACAACGCGACACGUUGCGUGGUGAACGGUUCAUGAGGCGCACAUGCGAG ......((((------..(((((((((((((((((((...))--))))))......(..(((((.((((((....)))))).)))))..))))))))))))....)))).. ( -33.70) >DroAna_CAF1 41278 108 + 1 UGCGACUGCAGCUGGAGCUGCUUUAUGAAUGCAUAAUUUUAUUUUAUGCACAAAUUCUAUUUACAACGCGACACGUUGCGU---GAACGGUUCAUAAAGUUGCUAUGCCCG .(((((((((((....))))).((((((((((((((.......))))))................((((((....))))))---.....))))))))))))))........ ( -35.20) >consensus UCCGAAUGCA______ACUGCUUUAUGAAUGCAUAAUUUUAU__UAUGCACAAAUUCUAUUUACAACGCGACACGUUGCGU___GAACGGUUCAUGAGGCGCAAAUGCGAG .......(((........(((((((((((((((((.........))))))......(..(((...((((((....))))))...)))..))))))))))))....)))... (-25.88 = -25.63 + -0.25)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 13:04:18 2006