
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 20,940,688 – 20,940,867 |
| Length | 179 |
| Max. P | 0.702022 |

| Location | 20,940,688 – 20,940,787 |
|---|---|
| Length | 99 |
| Sequences | 4 |
| Columns | 107 |
| Reading direction | forward |
| Mean pairwise identity | 88.67 |
| Mean single sequence MFE | -29.20 |
| Consensus MFE | -21.38 |
| Energy contribution | -22.88 |
| Covariance contribution | 1.50 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.81 |
| Structure conservation index | 0.73 |
| SVM decision value | 0.35 |
| SVM RNA-class probability | 0.702022 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 20940688 99 + 22224390 UUUCAUCCGCACUCUGCUCAAAUGGACAGCUGGUCGCCAUAACGCCAUUA--------CGAUCUCGGGCACCUUCACCUCGAAUUCGUCAAGCUCCUUUUUGGGCCA ...............(((((((.(((..(((((.((......))))...(--------(((..(((((........)).)))..))))..))))))..))))))).. ( -24.60) >DroSec_CAF1 12735 99 + 1 UUUCAUCCGCACUCGGCUCAAAUGGACAGCUGGUCGCCAUAACGCCAUUA--------CGAUCUCGGGCACCUUCACCUCGAAUACGUCGAGCUCCUGUUUGGGCCA ..............(((((((((((..((((((.((......))))....--------((((.(((((........)).)))....)))))))).))))))))))). ( -28.90) >DroEre_CAF1 11067 107 + 1 UUUCAUCCGCGCUCGGCUCAAAUGGACAGCUGGGCGCCAUAACGCCACAACGCCAUGACGAUCUCUGGCACCUUCACCUCGAAUACGUCUAGCUCCUGUUUGGGCCA ..............(((((((((((..(((((((((...............((((.((.....))))))...(((.....)))..))))))))).))))))))))). ( -35.30) >DroYak_CAF1 12264 99 + 1 UUUCAUCCACACUCGGCUCAAAUGGACAGCUGGGCGCCAUAACACCACUA--------CGACCUCGGAAACCUUCACCUCGAAUACGUCGAGCUCCUGUUUGGGCCA ..............(((((((((((..((((.((((..............--------.......(....).(((.....)))..)))).)))).))))))))))). ( -28.00) >consensus UUUCAUCCGCACUCGGCUCAAAUGGACAGCUGGGCGCCAUAACGCCACUA________CGAUCUCGGGCACCUUCACCUCGAAUACGUCGAGCUCCUGUUUGGGCCA ..............(((((((((((..((((.((((.......(((....................)))...(((.....)))..)))).)))).))))))))))). (-21.38 = -22.88 + 1.50)



| Location | 20,940,747 – 20,940,867 |
|---|---|
| Length | 120 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 89.82 |
| Mean single sequence MFE | -30.43 |
| Consensus MFE | -25.30 |
| Energy contribution | -26.55 |
| Covariance contribution | 1.25 |
| Combinations/Pair | 1.09 |
| Mean z-score | -1.36 |
| Structure conservation index | 0.83 |
| SVM decision value | -0.05 |
| SVM RNA-class probability | 0.508158 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
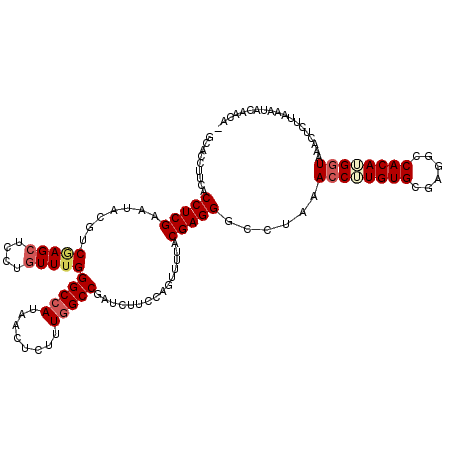
>X_DroMel_CAF1 20940747 120 + 22224390 GCACCUUCACCUCGAAUUCGUCAAGCUCCUUUUUGGGCCAUAACUCUUUGGCCGAUCUUUCAGUUUUACGAGGGCCUAAACCUUGUGCAAGGCCACAAGGUAAACUCUUAAAUGCAGCAU (((......(((((....)((.(((((......(.(((((........))))).)......))))).))))))......((((((((......))))))))...........)))..... ( -32.90) >DroSec_CAF1 12794 116 + 1 GCACCUUCACCUCGAAUACGUCGAGCUCCUGUUUGGGCCAUAACUCUUUGGCCGAUCUUUCAGUUUUACGAGGGCCUAAACCUUGUGCGAGGCCACAUGGUAAACUCUAAAAUACA---- ((((......(((((.....))))).....(((((((((....(((...(((.((....)).)))....))))))))))))...))))((((((....)))...))).........---- ( -31.20) >DroEre_CAF1 11134 119 + 1 GCACCUUCACCUCGAAUACGUCUAGCUCCUGUUUGGGCCAUAACUCUUUCGCCGAUCUUCCAGUUUUACGAGGGCCUAAACCGUGUGCGAGGUCACAUGCUAAACUCUUAAACUCAACA- (((.....((((((.(((((..........(((((((((....(((....((.(......).)).....))))))))))))))))).))))))....)))...................- ( -28.40) >DroYak_CAF1 12323 119 + 1 AAACCUUCACCUCGAAUACGUCGAGCUCCUGUUUGGGCCAUAACUCUUUGGCCGAUCUUCCAGUUUUACGAGGGUCUAAACCUUGUGCGCGGCCACAUGGUAAACUCUUAAAUCCAACG- ..((((((..(((((.....)))))...(((..(.(((((........))))).).....)))......))))))....(((.((((......)))).)))..................- ( -29.20) >consensus GCACCUUCACCUCGAAUACGUCGAGCUCCUGUUUGGGCCAUAACUCUUUGGCCGAUCUUCCAGUUUUACGAGGGCCUAAACCUUGUGCGAGGCCACAUGGUAAACUCUUAAAUACAACA_ .........(((((.......(((((....)))))(((((........)))))...............)))))......((((((((......))))))))................... (-25.30 = -26.55 + 1.25)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 13:04:00 2006