
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 20,879,245 – 20,879,387 |
| Length | 142 |
| Max. P | 0.998216 |

| Location | 20,879,245 – 20,879,364 |
|---|---|
| Length | 119 |
| Sequences | 3 |
| Columns | 119 |
| Reading direction | reverse |
| Mean pairwise identity | 83.00 |
| Mean single sequence MFE | -40.03 |
| Consensus MFE | -29.29 |
| Energy contribution | -29.63 |
| Covariance contribution | 0.35 |
| Combinations/Pair | 1.16 |
| Mean z-score | -1.69 |
| Structure conservation index | 0.73 |
| SVM decision value | 0.09 |
| SVM RNA-class probability | 0.576837 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 20879245 119 - 22224390 UCGGUGGAUCCAAUGUACAUAUGGAUACAUAUGUAUGUAUGUGUGCAUUAGGCGGGGAUUGCUCGAACGCGCUGGUGACGUCGCAGGAAUUCAAGGCAUUUGCAUCCCGCUGUGGAUUC .....((((((((((((((((((.((((....)))).)))))))))))).(((((((...........(((.((....)).)))...........((....)).)))))))..)))))) ( -46.50) >DroSec_CAF1 6866 113 - 1 UCGGUGGAUCCAAAGUGUAUAUGGAU----GUGUGCAUAUGUGUGCAUUAGGCGGGGAUUGCUCGA--GCGCUGGUGACGUCGCAGGAAUUCACGGCAUUUGCAUCCCGCUGUGGAUUC .....(((((((.((((.....((((----((((((((....))))))...((.((((((.((...--(((.((....)).))))).))))).).))....)))))))))).))))))) ( -40.40) >DroEre_CAF1 6897 115 - 1 GCGGUGGAUCCAAAGUACACUUGUAUA--UAGAUAUGUAUGUGUGCAUCAAGCGGGGAUUACUCGA--CUGCUGGUGACGUCGCAGGAAUUCAAGGCAUGUGCAUUUCUCGGUGGAUUC .....(((((((..((((....)))).--.(((.((((((((((........((((.....)))).--((((((....))..)))).........)))))))))).)))...))))))) ( -33.20) >consensus UCGGUGGAUCCAAAGUACAUAUGGAUA__UAUGUAUGUAUGUGUGCAUUAGGCGGGGAUUGCUCGA__GCGCUGGUGACGUCGCAGGAAUUCAAGGCAUUUGCAUCCCGCUGUGGAUUC .....(((((((..(((((((((.(((......))).))))))))).....(((((((((.((.....(((.((....)).))))).)))))...((....))...))))..))))))) (-29.29 = -29.63 + 0.35)



| Location | 20,879,285 – 20,879,387 |
|---|---|
| Length | 102 |
| Sequences | 3 |
| Columns | 102 |
| Reading direction | reverse |
| Mean pairwise identity | 74.17 |
| Mean single sequence MFE | -31.17 |
| Consensus MFE | -19.10 |
| Energy contribution | -19.33 |
| Covariance contribution | 0.23 |
| Combinations/Pair | 1.18 |
| Mean z-score | -2.47 |
| Structure conservation index | 0.61 |
| SVM decision value | 3.04 |
| SVM RNA-class probability | 0.998216 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 20879285 102 - 22224390 GACUCACUGGUGAUAAACCAGAAUCGGUGGAUCCAAUGUACAUAUGGAUACAUAUGUAUGUAUGUGUGCAUUAGGCGGGGAUUGCUCGAACGCGCUGGUGAC ......(((((.....))))).(((((((...((((((((((((((.((((....)))).)))))))))))).))((((.....))))....)))))))... ( -37.60) >DroSec_CAF1 6906 81 - 1 ---------------AACCAGAAUCGGUGGAUCCAAAGUGUAUAUGGAU----GUGUGCAUAUGUGUGCAUUAGGCGGGGAUUGCUCGA--GCGCUGGUGAC ---------------.(((((..((((..(((((..((((((((((.((----(....))).))))))))))......)))))..))))--...)))))... ( -26.70) >DroEre_CAF1 6937 98 - 1 GACUCACUGGUGAUAAACCAGAAGCGGUGGAUCCAAAGUACACUUGUAUA--UAGAUAUGUAUGUGUGCAUCAAGCGGGGAUUACUCGA--CUGCUGGUGAC ...((((((((.....)))...((((((.........((((((..(((((--(....))))))))))))......((((.....)))))--)))))))))). ( -29.20) >consensus GACUCACUGGUGAUAAACCAGAAUCGGUGGAUCCAAAGUACAUAUGGAUA__UAUGUAUGUAUGUGUGCAUUAGGCGGGGAUUGCUCGA__GCGCUGGUGAC ................(((((..((((..(((((...(((((((((.(((......))).))))))))).((....)))))))..)))).....)))))... (-19.10 = -19.33 + 0.23)


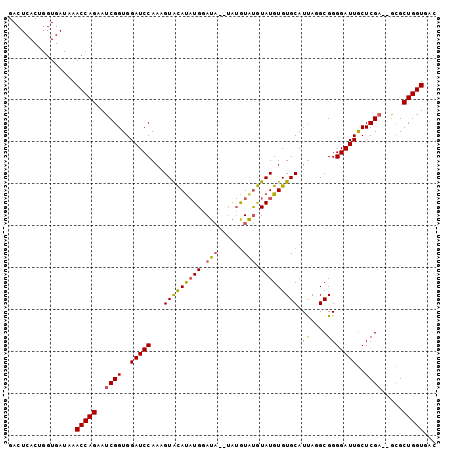
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 13:03:05 2006