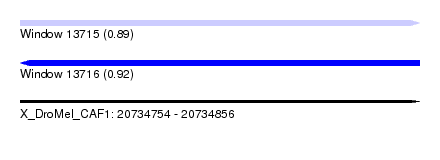

| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 20,734,754 – 20,734,856 |
| Length | 102 |
| Max. P | 0.916182 |

| Location | 20,734,754 – 20,734,856 |
|---|---|
| Length | 102 |
| Sequences | 3 |
| Columns | 102 |
| Reading direction | forward |
| Mean pairwise identity | 94.77 |
| Mean single sequence MFE | -31.90 |
| Consensus MFE | -28.27 |
| Energy contribution | -28.17 |
| Covariance contribution | -0.11 |
| Combinations/Pair | 1.07 |
| Mean z-score | -2.50 |
| Structure conservation index | 0.89 |
| SVM decision value | 0.95 |
| SVM RNA-class probability | 0.888673 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 20734754 102 + 22224390 ACCGCCGGCGCUGCGGAGUUGCAACUUCUGGAGAGAUUCGGUUCUCUCCAAAACUGCAUUGGCUACGAACUCUGUCUAAAACCAGUGACAAUUUUCGAAAUU ...(((((.((...(((((....)))))((((((((......)))))))).....)).)))))..((((...((((..........))))...))))..... ( -31.30) >DroSec_CAF1 42403 102 + 1 ACCGCCGGCGCUGCGGAGUUGCAACUUCUGGAGAGAUUCGGUUCUCUCCAAAACUGCAUUGGCUACGAACUCUGUCUAAAACUAAUGACAAUUUUCGAAAUU ...(((((.((...(((((....)))))((((((((......)))))))).....)).)))))..((((...((((..........))))...))))..... ( -31.30) >DroYak_CAF1 43106 102 + 1 AACGUCGGCGCUGCGGAGUUGCAACUUCUGGAGAGAUUCGGUUCUCUCCAAAACUGCAUUGCCUACGAACUUUGUCUAAAACUAGUGGCAAUUUUUGAAAUU ..(((.((((.((((((((....)))))((((((((......)))))))).....))).)))).)))....(((((..........)))))........... ( -33.10) >consensus ACCGCCGGCGCUGCGGAGUUGCAACUUCUGGAGAGAUUCGGUUCUCUCCAAAACUGCAUUGGCUACGAACUCUGUCUAAAACUAGUGACAAUUUUCGAAAUU ...(((((.((...(((((....)))))((((((((......)))))))).....)).)))))..((((...((((..........))))...))))..... (-28.27 = -28.17 + -0.11)



| Location | 20,734,754 – 20,734,856 |
|---|---|
| Length | 102 |
| Sequences | 3 |
| Columns | 102 |
| Reading direction | reverse |
| Mean pairwise identity | 94.77 |
| Mean single sequence MFE | -34.23 |
| Consensus MFE | -29.45 |
| Energy contribution | -30.23 |
| Covariance contribution | 0.78 |
| Combinations/Pair | 1.03 |
| Mean z-score | -2.95 |
| Structure conservation index | 0.86 |
| SVM decision value | 1.11 |
| SVM RNA-class probability | 0.916182 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 20734754 102 - 22224390 AAUUUCGAAAAUUGUCACUGGUUUUAGACAGAGUUCGUAGCCAAUGCAGUUUUGGAGAGAACCGAAUCUCUCCAGAAGUUGCAACUCCGCAGCGCCGGCGGU .....((((..(((((..........)))))..))))..(((...((..(((((((((((......)))))))))))(((((......))))))).)))... ( -35.50) >DroSec_CAF1 42403 102 - 1 AAUUUCGAAAAUUGUCAUUAGUUUUAGACAGAGUUCGUAGCCAAUGCAGUUUUGGAGAGAACCGAAUCUCUCCAGAAGUUGCAACUCCGCAGCGCCGGCGGU .....((((..(((((..........)))))..))))..(((...((..(((((((((((......)))))))))))(((((......))))))).)))... ( -35.50) >DroYak_CAF1 43106 102 - 1 AAUUUCAAAAAUUGCCACUAGUUUUAGACAAAGUUCGUAGGCAAUGCAGUUUUGGAGAGAACCGAAUCUCUCCAGAAGUUGCAACUCCGCAGCGCCGACGUU ....((.(((((((....))))))).)).......(((.(((.......(((((((((((......)))))))))))(((((......)))))))).))).. ( -31.70) >consensus AAUUUCGAAAAUUGUCACUAGUUUUAGACAGAGUUCGUAGCCAAUGCAGUUUUGGAGAGAACCGAAUCUCUCCAGAAGUUGCAACUCCGCAGCGCCGGCGGU .....((((..(((((..........)))))..))))..(((...((..(((((((((((......)))))))))))(((((......))))))).)))... (-29.45 = -30.23 + 0.78)

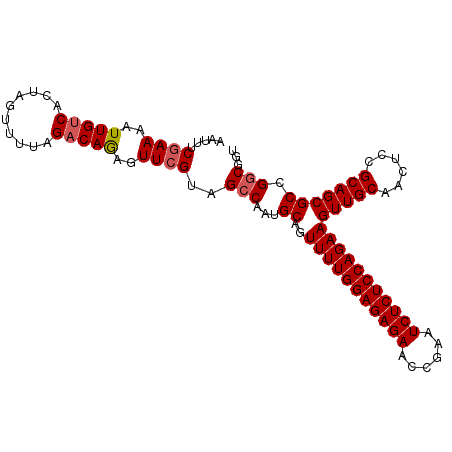

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 13:01:51 2006