| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 20,709,117 – 20,709,213 |
| Length | 96 |
| Max. P | 0.999345 |

| Location | 20,709,117 – 20,709,213 |
|---|---|
| Length | 96 |
| Sequences | 3 |
| Columns | 102 |
| Reading direction | forward |
| Mean pairwise identity | 79.67 |
| Mean single sequence MFE | -24.00 |
| Consensus MFE | -18.09 |
| Energy contribution | -18.43 |
| Covariance contribution | 0.34 |
| Combinations/Pair | 1.11 |
| Mean z-score | -3.80 |
| Structure conservation index | 0.75 |
| SVM decision value | 3.53 |
| SVM RNA-class probability | 0.999345 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 20709117 96 + 22224390 GUACAUAUAUAUGUGGUAUUUAUUUGCUUAUGGGAGCUGCAAAUGCCUGCUGCAUUUCUUGAUUUUCUGCUGCUCCAAUAAAUAAAUAAAUACUCA------ .(((((....)))))(((((((((((.((((.(((((.((((((((.....)))))...........))).))))).)))).)))))))))))...------ ( -28.10) >DroSec_CAF1 16798 102 + 1 UUUCAUGUAUAUGUAAUAUUUAUUUGCUUAUGGGAGCUCCAAGUGCCUGCUGCAUUUCUUGCUUUUUUGCUGCUCCAAUAAAUAAAUAAAUAUUUGUGAGUA .((((((.......((((((((((((.((((.(((((...((((((.....))))))...((......)).))))).)))).)))))))))))))))))).. ( -25.31) >DroYak_CAF1 16668 88 + 1 -----UAUAUAUUUGAUAUUUAUUUGCUUAUUGGCGCCCCAAAUGCCUGCUGCAUUCCUUGCUCUU---CUGCUCCAAUAAAUAAAUAAAUACUCA------ -----........(((((((((((((.(((((((.((.......(....).(((.....)))....---..)).))))))).)))))))))).)))------ ( -18.60) >consensus _U_CAUAUAUAUGUGAUAUUUAUUUGCUUAUGGGAGCUCCAAAUGCCUGCUGCAUUUCUUGCUUUU_UGCUGCUCCAAUAAAUAAAUAAAUACUCA______ ..............((((((((((((.((((.(((((...((((((.....))))))..............))))).)))).))))))))))))........ (-18.09 = -18.43 + 0.34)


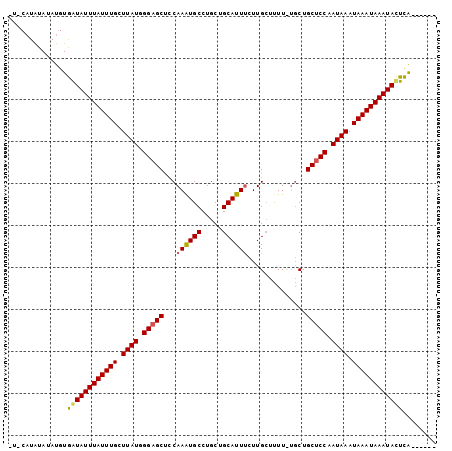
| Location | 20,709,117 – 20,709,213 |
|---|---|
| Length | 96 |
| Sequences | 3 |
| Columns | 102 |
| Reading direction | reverse |
| Mean pairwise identity | 79.67 |
| Mean single sequence MFE | -24.47 |
| Consensus MFE | -19.80 |
| Energy contribution | -19.80 |
| Covariance contribution | 0.00 |
| Combinations/Pair | 1.11 |
| Mean z-score | -3.97 |
| Structure conservation index | 0.81 |
| SVM decision value | 3.45 |
| SVM RNA-class probability | 0.999234 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 20709117 96 - 22224390 ------UGAGUAUUUAUUUAUUUAUUGGAGCAGCAGAAAAUCAAGAAAUGCAGCAGGCAUUUGCAGCUCCCAUAAGCAAAUAAAUACCACAUAUAUAUGUAC ------.(.((((((((((.(((((.(((((.((.(.....)...((((((.....)))))))).))))).))))).)))))))))))((((....)))).. ( -27.50) >DroSec_CAF1 16798 102 - 1 UACUCACAAAUAUUUAUUUAUUUAUUGGAGCAGCAAAAAAGCAAGAAAUGCAGCAGGCACUUGGAGCUCCCAUAAGCAAAUAAAUAUUACAUAUACAUGAAA ........(((((((((((.(((((.(((((.((......))......(((.....)))......))))).))))).))))))))))).............. ( -23.50) >DroYak_CAF1 16668 88 - 1 ------UGAGUAUUUAUUUAUUUAUUGGAGCAG---AAGAGCAAGGAAUGCAGCAGGCAUUUGGGGCGCCAAUAAGCAAAUAAAUAUCAAAUAUAUA----- ------(((.(((((((((.((((((((.((.(---.....)...((((((.....))))))...)).)))))))).))))))))))))........----- ( -22.40) >consensus ______UGAGUAUUUAUUUAUUUAUUGGAGCAGCA_AAAAGCAAGAAAUGCAGCAGGCAUUUGGAGCUCCCAUAAGCAAAUAAAUAUCACAUAUAUAUG_A_ .........((((((((((.(((((.(((((.(........)...((((((.....))))))...))))).))))).))))))))))............... (-19.80 = -19.80 + 0.00)

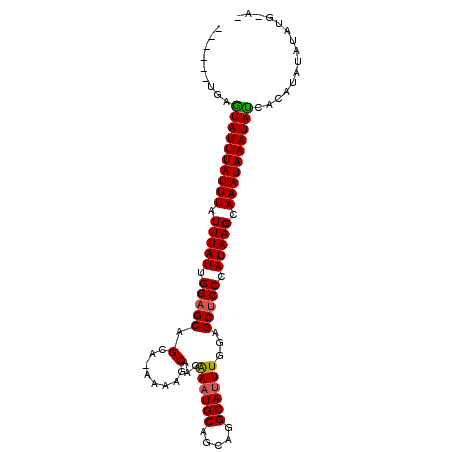

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 13:01:44 2006