
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 20,586,162 – 20,586,254 |
| Length | 92 |
| Max. P | 0.970651 |

| Location | 20,586,162 – 20,586,254 |
|---|---|
| Length | 92 |
| Sequences | 6 |
| Columns | 112 |
| Reading direction | forward |
| Mean pairwise identity | 78.39 |
| Mean single sequence MFE | -23.75 |
| Consensus MFE | -20.37 |
| Energy contribution | -20.70 |
| Covariance contribution | 0.33 |
| Combinations/Pair | 1.05 |
| Mean z-score | -2.12 |
| Structure conservation index | 0.86 |
| SVM decision value | 1.51 |
| SVM RNA-class probability | 0.960481 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
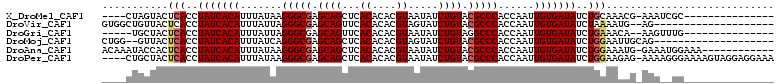
>X_DroMel_CAF1 20586162 92 + 22224390 ----CUAGUACUCACCUAUCACAUUUAUAAGGGCGAGCAGCUCACACACGUAAUAUCUGUACGCCCACCAAUUGUGAUAUCUGCAAACG-AAAUCGC--------------- ----...((...((..(((((((.......(((((.((((...((....)).....)))).)))))......)))))))..))...)).-.......--------------- ( -21.32) >DroVir_CAF1 204851 90 + 1 GUGGCUGUUACUCACCUAUCACAUUUAUUAGGGCGAGCAGUUCACACACGUAGUAUCUGUACGCCCACCAAUUGUGAUAUCUAAAAUG--AG-------------------- ((((.......)))).(((((((.......(((((.((((..(((....)).)...)))).)))))......))))))).........--..-------------------- ( -22.52) >DroGri_CAF1 184684 90 + 1 -----UGCUACUCACCUAUCACAUUUAUUAGGGCGAGCAGUUCACACACGUAAUAUCUGUAGGCCCACCAAUUGUGAUAUCUGAAACA--AAGUUUG--------------- -----......(((..(((((((.......((((..((((...((....)).....))))..))))......)))))))..)))....--.......--------------- ( -20.32) >DroMoj_CAF1 254499 90 + 1 CUGG--GUUACUCACCUAUCACAUUUAUCAGGGCGAGCAGCUCACACACGUAGUAUCUGUACGCCCACCAAUUGUGAUAUCUGGAAUUGCAG-------------------- (((.--(....(((..(((((((.......(((((.((((..(((....)).)...)))).)))))......)))))))..)))...).)))-------------------- ( -23.92) >DroAna_CAF1 162919 99 + 1 ACAAAUACCACUCACCUAUCACAUUUAUAAGGGCGAGCAGCUCACACACGUAAUAUCUGUACGCCCACCAAUUGUGAUAUCUGGAAAUG-GAAAUGGAAA------------ .......(((.(((..(((((((.......(((((.((((...((....)).....)))).)))))......)))))))..)))...))-).........------------ ( -24.32) >DroPer_CAF1 189261 107 + 1 ----CUGCUACUCACCUAUCACAUUUAUAAGGGCGAGCAGCUCACACACGUAAUAUCUGUACGCCCACCAAUUGUGAUAUCUGGAAGAG-AAAAGGGAAAAGUAGGAGGAAA ----((.(((((..(((((((((.......(((((.((((...((....)).....)))).)))))......)))))).(((.....))-)..)))....))))).)).... ( -30.12) >consensus ____CUGCUACUCACCUAUCACAUUUAUAAGGGCGAGCAGCUCACACACGUAAUAUCUGUACGCCCACCAAUUGUGAUAUCUGGAAAAG_AAA__GG_______________ ...........(((..(((((((.......(((((.((((...((....)).....)))).)))))......)))))))..)))............................ (-20.37 = -20.70 + 0.33)


| Location | 20,586,162 – 20,586,254 |
|---|---|
| Length | 92 |
| Sequences | 6 |
| Columns | 112 |
| Reading direction | reverse |
| Mean pairwise identity | 78.39 |
| Mean single sequence MFE | -29.27 |
| Consensus MFE | -25.19 |
| Energy contribution | -25.55 |
| Covariance contribution | 0.36 |
| Combinations/Pair | 1.04 |
| Mean z-score | -2.44 |
| Structure conservation index | 0.86 |
| SVM decision value | 1.66 |
| SVM RNA-class probability | 0.970651 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
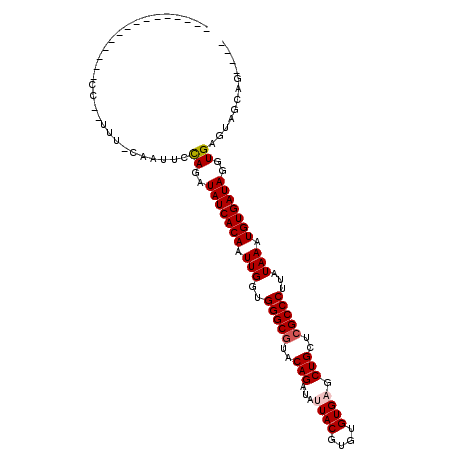
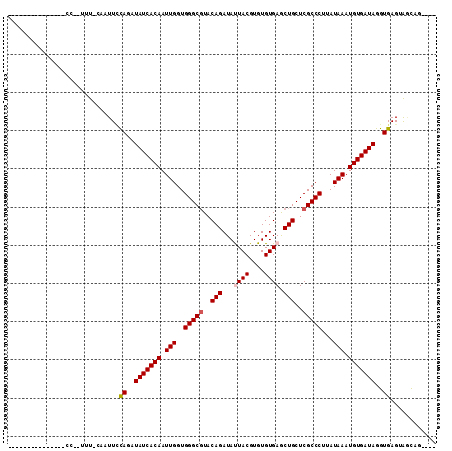
>X_DroMel_CAF1 20586162 92 - 22224390 ---------------GCGAUUU-CGUUUGCAGAUAUCACAAUUGGUGGGCGUACAGAUAUUACGUGUGUGAGCUGCUCGCCCUUAUAAAUGUGAUAGGUGAGUACUAG---- ---------------(((....-)))(..(...(((((((.(((..(((((..(((...((((....)))).)))..)))))...))).))))))).)..).......---- ( -29.60) >DroVir_CAF1 204851 90 - 1 --------------------CU--CAUUUUAGAUAUCACAAUUGGUGGGCGUACAGAUACUACGUGUGUGAACUGCUCGCCCUAAUAAAUGUGAUAGGUGAGUAACAGCCAC --------------------((--((((.....(((((((.(((..(((((..(((((((.....))))...)))..)))))...))).))))))))))))).......... ( -28.00) >DroGri_CAF1 184684 90 - 1 ---------------CAAACUU--UGUUUCAGAUAUCACAAUUGGUGGGCCUACAGAUAUUACGUGUGUGAACUGCUCGCCCUAAUAAAUGUGAUAGGUGAGUAGCA----- ---------------.......--(((((((..(((((((.(((..((((...(((...((((....)))).)))...))))...))).)))))))..)))..))))----- ( -23.80) >DroMoj_CAF1 254499 90 - 1 --------------------CUGCAAUUCCAGAUAUCACAAUUGGUGGGCGUACAGAUACUACGUGUGUGAGCUGCUCGCCCUGAUAAAUGUGAUAGGUGAGUAAC--CCAG --------------------(((......))).(((((((.(((..(((((..(((((((.....))))...)))..)))))...))).)))))))(((.....))--)... ( -27.40) >DroAna_CAF1 162919 99 - 1 ------------UUUCCAUUUC-CAUUUCCAGAUAUCACAAUUGGUGGGCGUACAGAUAUUACGUGUGUGAGCUGCUCGCCCUUAUAAAUGUGAUAGGUGAGUGGUAUUUGU ------------....((..((-((((..(...(((((((.(((..(((((..(((...((((....)))).)))..)))))...))).))))))).)..))))).)..)). ( -33.50) >DroPer_CAF1 189261 107 - 1 UUUCCUCCUACUUUUCCCUUUU-CUCUUCCAGAUAUCACAAUUGGUGGGCGUACAGAUAUUACGUGUGUGAGCUGCUCGCCCUUAUAAAUGUGAUAGGUGAGUAGCAG---- ....((.((((((..((....(-((.....)))(((((((.(((..(((((..(((...((((....)))).)))..)))))...))).))))))))).)))))).))---- ( -33.30) >consensus _______________CC__UUU_CAAUUCCAGAUAUCACAAUUGGUGGGCGUACAGAUAUUACGUGUGUGAGCUGCUCGCCCUUAUAAAUGUGAUAGGUGAGUAGCAG____ .............................((..(((((((.(((..(((((..(((...((((....)))).)))..)))))...))).)))))))..))............ (-25.19 = -25.55 + 0.36)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 13:00:00 2006