
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 20,464,551 – 20,464,641 |
| Length | 90 |
| Max. P | 0.941963 |

| Location | 20,464,551 – 20,464,641 |
|---|---|
| Length | 90 |
| Sequences | 6 |
| Columns | 98 |
| Reading direction | forward |
| Mean pairwise identity | 78.12 |
| Mean single sequence MFE | -13.81 |
| Consensus MFE | -12.28 |
| Energy contribution | -12.45 |
| Covariance contribution | 0.17 |
| Combinations/Pair | 1.22 |
| Mean z-score | -1.81 |
| Structure conservation index | 0.89 |
| SVM decision value | 1.32 |
| SVM RNA-class probability | 0.941963 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 20464551 90 + 22224390 ------GCUCUUGCC--UUUUUUCGAGAUUAUUAUUAGCGCAACAUUAACGUCAAAUUACGCAUCUAUUUUGCGCACUUUAUCUCGAUCUCCCUAUAU ------((....)).--.....(((((((........((((((.((...(((......))).....)).)))))).....)))))))........... ( -16.92) >DroSec_CAF1 37918 92 + 1 ------CCUCUUGCCGUUUUUUUCGGGAUUAUUAUUAGCGCAACAUUAACGUCAAAUUACUCAUCUAUUUUGCGCACUUUAUCUCGAUCUCCCUAUAU ------................(((((((........((((((.((....................)).)))))).....)))))))........... ( -13.07) >DroSim_CAF1 43027 92 + 1 ------CCUCUUGCCGUUUUUUUCGGGAUUAUUAUUAGCGCAACAUUAACGUCAAAUUACGCAUCUAUUUUGCGCACUUUAUCUCGAUCUCCCUAUAU ------................(((((((........((((((.((...(((......))).....)).)))))).....)))))))........... ( -15.42) >DroWil_CAF1 56736 90 + 1 GAUCUACUUUAUGUGAU---CUGACAGAUUAUUAUUAGCGUAACAUUAACGUCAAAUUACGCAUCUAUUUUGCGCACUUUAUCUCUCUCUUCC----- ((((.((.....)))))---).((.((((........((((((.((...(((......))).....)).)))))).....)))).))......----- ( -12.72) >DroYak_CAF1 36762 89 + 1 ------CCUCGCCC---AUUUCUCGAGAUUAUUAUUAGCGCAACAUUAACGUCAAAUUACGCAUCUAUUUUGCGCACUUUAUCUCGAUCUCCCUAUAU ------........---.....(((((((........((((((.((...(((......))).....)).)))))).....)))))))........... ( -16.22) >DroPer_CAF1 59864 83 + 1 UACCCAUCUAAUGCCGG---CUGACAGAUUAUUAUUAGCGUAACAUUAACGUCAAAUUACGCAUCUAUUUUGCGCACUUUAUCC------------AU ........(((((...(---((((..........)))))....)))))...........((((.......))))..........------------.. ( -8.50) >consensus ______CCUCUUGCCGUUUUUUUCGAGAUUAUUAUUAGCGCAACAUUAACGUCAAAUUACGCAUCUAUUUUGCGCACUUUAUCUCGAUCUCCCUAUAU ......................(((((((........((((((.((...(((......))).....)).)))))).....)))))))........... (-12.28 = -12.45 + 0.17)


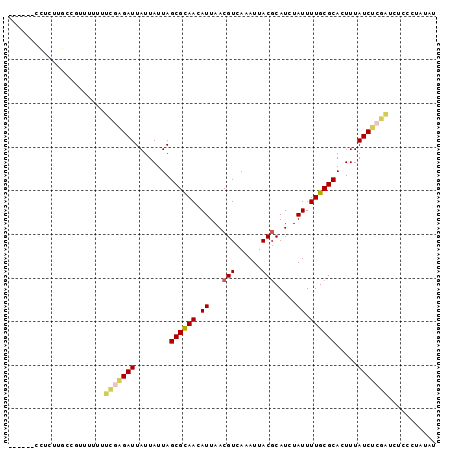
| Location | 20,464,551 – 20,464,641 |
|---|---|
| Length | 90 |
| Sequences | 6 |
| Columns | 98 |
| Reading direction | reverse |
| Mean pairwise identity | 78.12 |
| Mean single sequence MFE | -17.41 |
| Consensus MFE | -13.23 |
| Energy contribution | -14.20 |
| Covariance contribution | 0.97 |
| Combinations/Pair | 1.18 |
| Mean z-score | -1.33 |
| Structure conservation index | 0.76 |
| SVM decision value | 0.71 |
| SVM RNA-class probability | 0.830195 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 20464551 90 - 22224390 AUAUAGGGAGAUCGAGAUAAAGUGCGCAAAAUAGAUGCGUAAUUUGACGUUAAUGUUGCGCUAAUAAUAAUCUCGAAAAAA--GGCAAGAGC------ ...........(((((((.....((((((.......((((......)))).....))))))........))))))).....--.........------ ( -20.32) >DroSec_CAF1 37918 92 - 1 AUAUAGGGAGAUCGAGAUAAAGUGCGCAAAAUAGAUGAGUAAUUUGACGUUAAUGUUGCGCUAAUAAUAAUCCCGAAAAAAACGGCAAGAGG------ .....((((..............((((((.((.((((..........)))).)).)))))).........))))..................------ ( -14.70) >DroSim_CAF1 43027 92 - 1 AUAUAGGGAGAUCGAGAUAAAGUGCGCAAAAUAGAUGCGUAAUUUGACGUUAAUGUUGCGCUAAUAAUAAUCCCGAAAAAAACGGCAAGAGG------ .....((((..............((((((.......((((......)))).....)))))).........))))..................------ ( -17.30) >DroWil_CAF1 56736 90 - 1 -----GGAAGAGAGAGAUAAAGUGCGCAAAAUAGAUGCGUAAUUUGACGUUAAUGUUACGCUAAUAAUAAUCUGUCAG---AUCACAUAAAGUAGAUC -----......((.((((....((((((.......))))))...(((((....)))))...........)))).)).(---(((((.....)).)))) ( -15.80) >DroYak_CAF1 36762 89 - 1 AUAUAGGGAGAUCGAGAUAAAGUGCGCAAAAUAGAUGCGUAAUUUGACGUUAAUGUUGCGCUAAUAAUAAUCUCGAGAAAU---GGGCGAGG------ ...........(((((((.....((((((.......((((......)))).....))))))........))))))).....---........------ ( -20.32) >DroPer_CAF1 59864 83 - 1 AU------------GGAUAAAGUGCGCAAAAUAGAUGCGUAAUUUGACGUUAAUGUUACGCUAAUAAUAAUCUGUCAG---CCGGCAUUAGAUGGGUA ..------------...((((.((((((.......)))))).)))).((((((((((..(((.(((......))).))---).)))))).)))).... ( -16.00) >consensus AUAUAGGGAGAUCGAGAUAAAGUGCGCAAAAUAGAUGCGUAAUUUGACGUUAAUGUUGCGCUAAUAAUAAUCUCGAAAAAAACGGCAAGAGG______ ...........(((((((.....((((((.......((((......)))).....))))))........)))))))...................... (-13.23 = -14.20 + 0.97)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:59:17 2006