
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 20,418,095 – 20,418,236 |
| Length | 141 |
| Max. P | 0.999855 |

| Location | 20,418,095 – 20,418,197 |
|---|---|
| Length | 102 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 88.29 |
| Mean single sequence MFE | -28.23 |
| Consensus MFE | -24.32 |
| Energy contribution | -24.85 |
| Covariance contribution | 0.53 |
| Combinations/Pair | 1.03 |
| Mean z-score | -1.69 |
| Structure conservation index | 0.86 |
| SVM decision value | 0.37 |
| SVM RNA-class probability | 0.708144 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 20418095 102 + 22224390 -CGCGUUCACGUUGCACCC-------C----------CCCGCCAACACAUGUACGACAUGUAAAUUGUUUCAAUUUGCGGCAUUUGCCGUAAAGCUGCGCAAUUUUCGACAUGCCAUGGA -.(((.......)))....-------.----------....(((...(((((.(((.....(((((((.....((((((((....)))))))).....)))))))))))))))...))). ( -28.10) >DroSec_CAF1 7464 106 + 1 -CGCGCUCACGUUGCACCC-------CCCC------CCCCGCCAACACAUGUACGACAUGUAAAUUGUUUCAAUUUGCGGCAUUUGCCGUAAAGCUGCGCAAUUUUCGACAUGCCAUGGA -...((.......))....-------....------.....(((...(((((.(((.....(((((((.....((((((((....)))))))).....)))))))))))))))...))). ( -27.60) >DroAna_CAF1 8083 103 + 1 CCCCAUCCUCGCUUCAAUU-------C----------GCGGCCAACACAUGUACAACAUGUAAAUUGUUUCAAUUUGCGGCAUUUGCCGUAAAGCUGCGCAAUUUUCGACAUGCCAUGGA ..((((....((.......-------.----------))(((....(((((.....)))))(((((((.....((((((((....)))))))).....))))))).......))))))). ( -29.60) >DroSim_CAF1 3887 102 + 1 -CGCGCUCACGUUGCACCC-------C----------CCCGCCAACACAUGUACGACAUGUAAAUUGUUUCAAUUUGCGGCAUUUGCCGUAAAGCUGCGCAAUUUUCGACAUGCCAUGGA -...((.......))....-------.----------....(((...(((((.(((.....(((((((.....((((((((....)))))))).....)))))))))))))))...))). ( -27.60) >DroEre_CAF1 6737 119 + 1 -CGCGUUCACGUUGCACACCACCCACACCCCCCGUCCCCCGCCAACACAUGUAGGACAUGUAAAUUGUUUCAAUUUGCGGCAUUUGCCGUAAAGCUGCGCAAUUUUCGACAUGCCAUGGA -.(((.......)))...(((............((((..((........))..))))(((((((((((.....((((((((....)))))))).....)))))))...))))....))). ( -28.40) >DroYak_CAF1 3509 106 + 1 -CGCGUUCACGUUGCACAC-------ACCC------CCCCGCCAACACAUGUACGACAUGUAAAUUGUUUCAAUUUGCGGCAUUUGCCGUAAAGCUGCGCAAUUUUCGACAUGCCAUGGA -.(((.......)))....-------....------.....(((...(((((.(((.....(((((((.....((((((((....)))))))).....)))))))))))))))...))). ( -28.10) >consensus _CGCGUUCACGUUGCACCC_______C__________CCCGCCAACACAUGUACGACAUGUAAAUUGUUUCAAUUUGCGGCAUUUGCCGUAAAGCUGCGCAAUUUUCGACAUGCCAUGGA ..(((.......)))..........................(((...(((((.(((.....(((((((.....((((((((....)))))))).....)))))))))))))))...))). (-24.32 = -24.85 + 0.53)

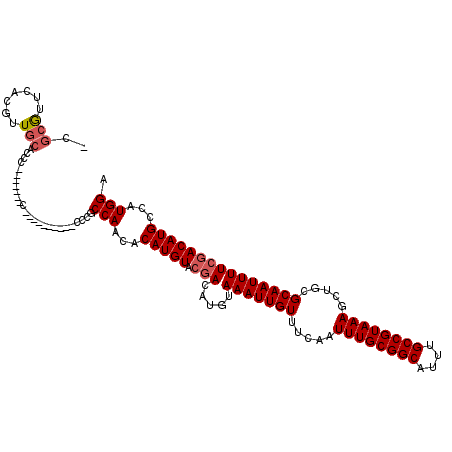

| Location | 20,418,117 – 20,418,236 |
|---|---|
| Length | 119 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 92.26 |
| Mean single sequence MFE | -33.15 |
| Consensus MFE | -31.24 |
| Energy contribution | -31.60 |
| Covariance contribution | 0.36 |
| Combinations/Pair | 1.03 |
| Mean z-score | -2.91 |
| Structure conservation index | 0.94 |
| SVM decision value | 4.27 |
| SVM RNA-class probability | 0.999855 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 20418117 119 + 22224390 GCCAACACAUGUACGACAUGUAAAUUGUUUCAAUUUGCGGCAUUUGCCGUAAAGCUGCGCAAUUUUCGACAUGCCAUGGAUACCCAUGGACCUGUCCGUUCGUCCACCAACAACCC-CAU ((.((((((((.....)))))(((((((.....((((((((....)))))))).....)))))))..((((.(((((((....)))))).).)))).))).)).............-... ( -35.50) >DroSec_CAF1 7490 119 + 1 GCCAACACAUGUACGACAUGUAAAUUGUUUCAAUUUGCGGCAUUUGCCGUAAAGCUGCGCAAUUUUCGACAUGCCAUGGAUACCCAUGGACCUGUCCGUCCGUCCGCCAACAACCC-CAU ((..(((((((.....)))))(((((((.....((((((((....)))))))).....)))))))..((((.(((((((....)))))).).)))).....))..)).........-... ( -36.00) >DroAna_CAF1 8106 115 + 1 GCCAACACAUGUACAACAUGUAAAUUGUUUCAAUUUGCGGCAUUUGCCGUAAAGCUGCGCAAUUUUCGACAUGCCAUGGAUACUCAUAGA---GUCCUUU-UUCGA-GUCCAACCCUUAU ......(((((.....)))))(((((((.....((((((((....)))))))).....)))))))..(((.((....((((.(.....).---))))...-..)).-))).......... ( -26.20) >DroSim_CAF1 3909 119 + 1 GCCAACACAUGUACGACAUGUAAAUUGUUUCAAUUUGCGGCAUUUGCCGUAAAGCUGCGCAAUUUUCGACAUGCCAUGGAUACCCAUGGACCUGUCCGUCCGUCCGCCAACAACCC-CAU ((..(((((((.....)))))(((((((.....((((((((....)))))))).....)))))))..((((.(((((((....)))))).).)))).....))..)).........-... ( -36.00) >DroEre_CAF1 6776 115 + 1 GCCAACACAUGUAGGACAUGUAAAUUGUUUCAAUUUGCGGCAUUUGCCGUAAAGCUGCGCAAUUUUCGACAUGCCAUGGAUACCCAUGGACUUGUCC----AUCCACCAACAACCC-CAU .........(((.((....(.(((((((.....((((((((....)))))))).....))))))).)((((..((((((....))))))...)))).----.....)).)))....-... ( -35.30) >DroYak_CAF1 3535 115 + 1 GCCAACACAUGUACGACAUGUAAAUUGUUUCAAUUUGCGGCAUUUGCCGUAAAGCUGCGCAAUUUUCGACAUGCCAUGGAUACCCAUGAACUUGUCC----GUCCACCAACAACCC-CAU ......(((((.....)))))(((((((.....((((((((....)))))))).....)))))))..((((...(((((....)))))....)))).----...............-... ( -29.90) >consensus GCCAACACAUGUACGACAUGUAAAUUGUUUCAAUUUGCGGCAUUUGCCGUAAAGCUGCGCAAUUUUCGACAUGCCAUGGAUACCCAUGGACCUGUCCGU__GUCCACCAACAACCC_CAU ......(((((.....)))))(((((((.....((((((((....)))))))).....)))))))..((((..((((((....))))))...))))........................ (-31.24 = -31.60 + 0.36)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:58:39 2006