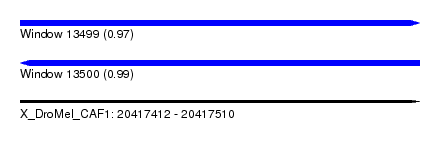
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 20,417,412 – 20,417,510 |
| Length | 98 |
| Max. P | 0.992550 |

| Location | 20,417,412 – 20,417,510 |
|---|---|
| Length | 98 |
| Sequences | 5 |
| Columns | 98 |
| Reading direction | forward |
| Mean pairwise identity | 90.46 |
| Mean single sequence MFE | -20.71 |
| Consensus MFE | -18.62 |
| Energy contribution | -18.26 |
| Covariance contribution | -0.36 |
| Combinations/Pair | 1.14 |
| Mean z-score | -1.68 |
| Structure conservation index | 0.90 |
| SVM decision value | 1.58 |
| SVM RNA-class probability | 0.965122 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 20417412 98 + 22224390 AAAAAAAAAAAACUGAAGGAAGUAGGUGAGAUAAGCGGUAAAAGGGGGUUAAAACUGGGACAGACUGGACUCUCAAAUUAAGUUGAGGAGCCAGCUGU ...........(((......)))............((((..............))))..((((.((((..((((((......))))))..)))))))) ( -18.34) >DroSec_CAF1 6778 92 + 1 ------AAAAAACUGAAGGAAGUAGGUGAGAUAAGCGGUAAAAGGGGGUUAAAACUGGGCCAGACCGGACUCUCAAAUUAAGUUGAGGAGCCAGCUGU ------.....(((......)))...........(((((....((.((((.......))))...))((..((((((......))))))..)).))))) ( -19.30) >DroSim_CAF1 3193 93 + 1 -----AAAAAAACUGAAGGAAGUAGGUGAGAUAAGCGGUAAAAGGGGGUUAAAACUGGGCCAGACCGGACUCUCAAAUUAAGUUGAGGAGCCAGCUGU -----......(((......)))...........(((((....((.((((.......))))...))((..((((((......))))))..)).))))) ( -19.30) >DroEre_CAF1 6085 89 + 1 ----A-----AGCUGGAGGAAGUAAGUGAGAUAAGCGGCAAGAGGGGGUUACAACUGGGCCAGACCGGACUCUCAAAUUAAGUUGAGGAUCCAGCUGU ----.-----(((((((((......((.......))(((...((..(....)..))..)))...))....((((((......)))))).))))))).. ( -25.00) >DroYak_CAF1 2863 94 + 1 ----AAAAGAAACUGGAGGAAGUAGGUGAGAUAAGCGGCAAAAGGGGGUUAAAACAGGGCCAGACCGGACUCUCAAAUUAAGUUGAGGAGCCAGCUGU ----.......(((......)))...........(((((....((.((((.......))))...))((..((((((......))))))..)).))))) ( -21.60) >consensus ____AAAAAAAACUGAAGGAAGUAGGUGAGAUAAGCGGUAAAAGGGGGUUAAAACUGGGCCAGACCGGACUCUCAAAUUAAGUUGAGGAGCCAGCUGU ...........(((......)))...........(((((....((.((((.......))))...))((..((((((......))))))..)).))))) (-18.62 = -18.26 + -0.36)


| Location | 20,417,412 – 20,417,510 |
|---|---|
| Length | 98 |
| Sequences | 5 |
| Columns | 98 |
| Reading direction | reverse |
| Mean pairwise identity | 90.46 |
| Mean single sequence MFE | -16.10 |
| Consensus MFE | -13.86 |
| Energy contribution | -13.90 |
| Covariance contribution | 0.04 |
| Combinations/Pair | 1.07 |
| Mean z-score | -2.14 |
| Structure conservation index | 0.86 |
| SVM decision value | 2.34 |
| SVM RNA-class probability | 0.992550 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
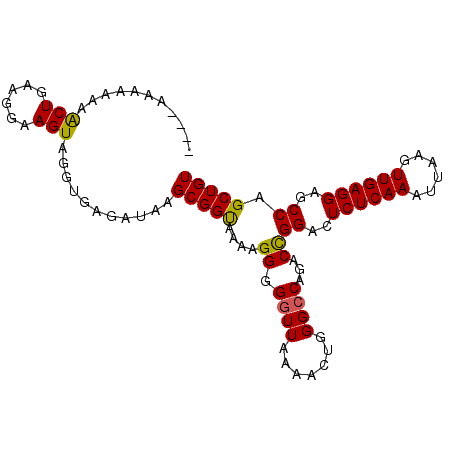
>X_DroMel_CAF1 20417412 98 - 22224390 ACAGCUGGCUCCUCAACUUAAUUUGAGAGUCCAGUCUGUCCCAGUUUUAACCCCCUUUUACCGCUUAUCUCACCUACUUCCUUCAGUUUUUUUUUUUU (((((((((((.((((......))))))).)))).))))........................................................... ( -15.70) >DroSec_CAF1 6778 92 - 1 ACAGCUGGCUCCUCAACUUAAUUUGAGAGUCCGGUCUGGCCCAGUUUUAACCCCCUUUUACCGCUUAUCUCACCUACUUCCUUCAGUUUUUU------ .((((((((((.((((......))))))).)))).)))......................................................------ ( -15.00) >DroSim_CAF1 3193 93 - 1 ACAGCUGGCUCCUCAACUUAAUUUGAGAGUCCGGUCUGGCCCAGUUUUAACCCCCUUUUACCGCUUAUCUCACCUACUUCCUUCAGUUUUUUU----- .((((((((((.((((......))))))).)))).))).......................................................----- ( -15.00) >DroEre_CAF1 6085 89 - 1 ACAGCUGGAUCCUCAACUUAAUUUGAGAGUCCGGUCUGGCCCAGUUGUAACCCCCUCUUGCCGCUUAUCUCACUUACUUCCUCCAGCU-----U---- .((((((((..(((((......)))))..))))).)))....(((.((((.......)))).))).......................-----.---- ( -19.80) >DroYak_CAF1 2863 94 - 1 ACAGCUGGCUCCUCAACUUAAUUUGAGAGUCCGGUCUGGCCCUGUUUUAACCCCCUUUUGCCGCUUAUCUCACCUACUUCCUCCAGUUUCUUUU---- .((((((((((.((((......))))))).)))).)))........................................................---- ( -15.00) >consensus ACAGCUGGCUCCUCAACUUAAUUUGAGAGUCCGGUCUGGCCCAGUUUUAACCCCCUUUUACCGCUUAUCUCACCUACUUCCUUCAGUUUUUUUU____ .((((((((((.((((......))))))).)))).)))............................................................ (-13.86 = -13.90 + 0.04)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:58:37 2006