
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 20,369,016 – 20,369,111 |
| Length | 95 |
| Max. P | 0.867831 |

| Location | 20,369,016 – 20,369,111 |
|---|---|
| Length | 95 |
| Sequences | 4 |
| Columns | 95 |
| Reading direction | forward |
| Mean pairwise identity | 82.22 |
| Mean single sequence MFE | -35.85 |
| Consensus MFE | -26.54 |
| Energy contribution | -26.85 |
| Covariance contribution | 0.31 |
| Combinations/Pair | 1.19 |
| Mean z-score | -1.81 |
| Structure conservation index | 0.74 |
| SVM decision value | 0.58 |
| SVM RNA-class probability | 0.790275 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 20369016 95 + 22224390 UUAUUUUCUCUGCAUGCACGUGUGGUGGUGUUGGCGGUGCGGCCUUGUGUGUGUGAGCUUGGCAUGCCGGGCAGCACACCCACACUACCAUUUGC ...............(((...((((((((((.((..((((.((((.((((((.........)))))).)))).))))..)))))))))))).))) ( -40.00) >DroSim_CAF1 16900 95 + 1 AUAUUUUCUUUGCGUGCACGUGUGUUGGUGUUGGCGGUGCGGCCUUGUGUGUGUGAGCUUGGCAUGCCGGGCAGCACACCCACACCACCAUUGGC ...............((....(((.((((((.((..((((.((((.((((((.........)))))).)))).))))..)))))))).)))..)) ( -35.10) >DroEre_CAF1 15884 78 + 1 UUGUUAUCUCUGCGUGUGCGUGUGGUGG---------------CCUGUGUGUGUGCGCUUGACAUGCCGGGCAGCACACCCACACCAC--UUUGC ..........((.(((((.(((((.((.---------------((((.((((((.......)))))))))))).))))).))))))).--..... ( -30.10) >DroYak_CAF1 6255 93 + 1 AUAUUUUCUCUGCGUGCACGUGUGGUGGUGUUGGUGGUGCGGCCUUGUGUGUGUGAGCCUGGCAUGCCGGGCAGCACACCCACACCAC--AUUGC ...............(((.((((((((.....(((......))).((.((((((..((((((....)))))).)))))).))))))))--))))) ( -38.20) >consensus AUAUUUUCUCUGCGUGCACGUGUGGUGGUGUUGGCGGUGCGGCCUUGUGUGUGUGAGCUUGGCAUGCCGGGCAGCACACCCACACCAC__UUUGC ...............(((...((((((.....(((......))).((.((((((..((((((....)))))).)))))).))))))))....))) (-26.54 = -26.85 + 0.31)


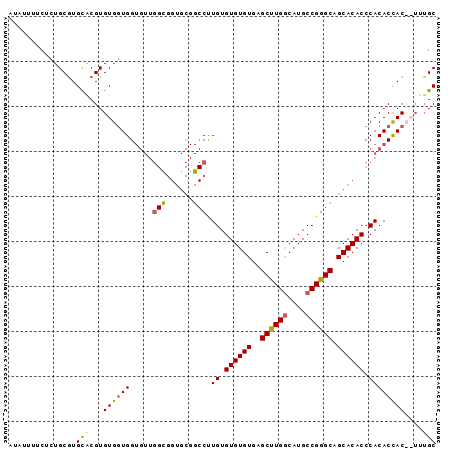
| Location | 20,369,016 – 20,369,111 |
|---|---|
| Length | 95 |
| Sequences | 4 |
| Columns | 95 |
| Reading direction | reverse |
| Mean pairwise identity | 82.22 |
| Mean single sequence MFE | -29.88 |
| Consensus MFE | -21.42 |
| Energy contribution | -21.55 |
| Covariance contribution | 0.13 |
| Combinations/Pair | 1.17 |
| Mean z-score | -1.98 |
| Structure conservation index | 0.72 |
| SVM decision value | 0.86 |
| SVM RNA-class probability | 0.867831 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
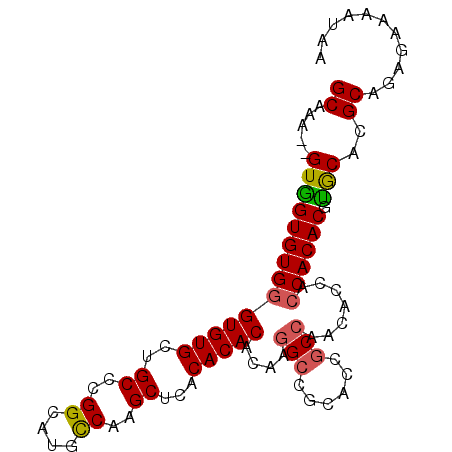
>X_DroMel_CAF1 20369016 95 - 22224390 GCAAAUGGUAGUGUGGGUGUGCUGCCCGGCAUGCCAAGCUCACACACACAAGGCCGCACCGCCAACACCACCACACGUGCAUGCAGAGAAAAUAA (((....((((((((((((((..((..((....))..)).)))))......(((......))).......)))))).))).)))........... ( -30.10) >DroSim_CAF1 16900 95 - 1 GCCAAUGGUGGUGUGGGUGUGCUGCCCGGCAUGCCAAGCUCACACACACAAGGCCGCACCGCCAACACCAACACACGUGCACGCAAAGAAAAUAU ((...((((((((((((((((((....))))))))..(((...........))))))))))))).(((........)))...))........... ( -32.40) >DroEre_CAF1 15884 78 - 1 GCAAA--GUGGUGUGGGUGUGCUGCCCGGCAUGUCAAGCGCACACACACAGG---------------CCACCACACGCACACGCAGAGAUAACAA .....--(((((.((.((((((.((..((....))..))))))))...)).)---------------)))).....((....))........... ( -24.30) >DroYak_CAF1 6255 93 - 1 GCAAU--GUGGUGUGGGUGUGCUGCCCGGCAUGCCAGGCUCACACACACAAGGCCGCACCACCAACACCACCACACGUGCACGCAGAGAAAAUAU ((...--((((((((((((((..(((.((....)).))).))))).(....).)))))))))...(((........)))...))........... ( -32.70) >consensus GCAAA__GUGGUGUGGGUGUGCUGCCCGGCAUGCCAAGCUCACACACACAAGGCCGCACCGCCAACACCACCACACGUGCACGCAGAGAAAAUAA ((.....((((((((((((((..((..((....))..))...)))))....(((......))).......)))))).)))..))........... (-21.42 = -21.55 + 0.13)

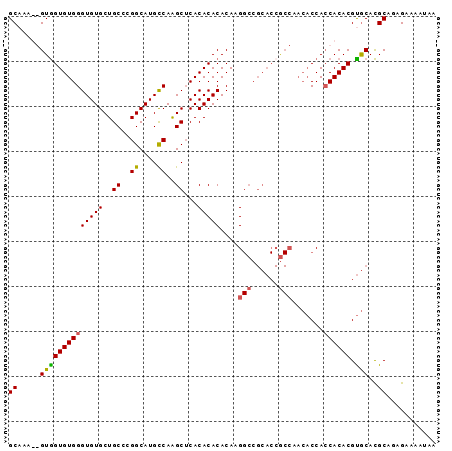
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:58:13 2006