
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 20,348,356 – 20,348,475 |
| Length | 119 |
| Max. P | 0.999993 |

| Location | 20,348,356 – 20,348,475 |
|---|---|
| Length | 119 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 69.41 |
| Mean single sequence MFE | -35.62 |
| Consensus MFE | -24.80 |
| Energy contribution | -30.18 |
| Covariance contribution | 5.38 |
| Combinations/Pair | 1.26 |
| Mean z-score | -3.79 |
| Structure conservation index | 0.70 |
| SVM decision value | 5.77 |
| SVM RNA-class probability | 0.999993 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 20348356 119 + 22224390 UCGAGAGAAGGCUCGACCAGACCAAAGAGAUAUGAUGAUAUGAUAUCAAAGUUUUUGGUUCGCCGAGCCGGAUGUUGCAUGAAAAAAAUUGUGCACUUCCUUGCAA-UUUUACCAUUCAA ((....)).((((((..(..(((((((((((((.((...)).)))))....))))))))..).))))))(((.((.(((..(......)..))))).)))......-............. ( -32.90) >DroSec_CAF1 16277 111 + 1 CCAAUAUCCGGCUCGAG-GGACCAAAAUGAUAUGAUGA-----UAGCAAAGUUUUCGGUCCGCCAAGCCGGAUGUUGCAUUAAA--AAUAGUGCAGUUCCUUGCAA-UUUUCCCAUUCAA .((((((((((((.(..-((((((((((..(((....)-----)).....))))).)))))..).)))))))))))).......--.....(((((....))))).-............. ( -33.80) >DroEre_CAF1 13050 100 + 1 CCAUAAUCCGGCUCGAG-GGACCAUAAAGAUAUGUUUA-----UAGCGUAGAUUUUGGUCCAUUGAGCCGG-----------UA--AAUAAUGCAGUUGCUUGCAA-UUUGCCCAUUAAU .........(((((((.-((((((.(((..(((((...-----..)))))..))))))))).)))))))((-----------((--(((..(((((....))))))-))))))....... ( -38.20) >DroYak_CAF1 16826 112 + 1 ACAAUAUCCGGCUCGAG-GGACCACCAAUAUAUGUUAA-----UGUGGUUGUCUUUGGUCCUUCGUGCCGCAUAUUGCAUUAAA--AACAGUGCAGUUCUUUGCAAAAUUUCCCAUUAAA .((((((.((((.((((-((((((.((((((((....)-----))).))))....)))))))))).)))).)))))).......--.....(((((....)))))............... ( -37.60) >consensus CCAAUAUCCGGCUCGAG_GGACCAAAAAGAUAUGAUGA_____UAGCAAAGUUUUUGGUCCGCCGAGCCGGAUGUUGCAUUAAA__AAUAGUGCAGUUCCUUGCAA_UUUUCCCAUUAAA .((((((((((((((...(((((((((...(((((((......)))))))..)))))))))..))))))))))))))..............(((((....)))))............... (-24.80 = -30.18 + 5.38)


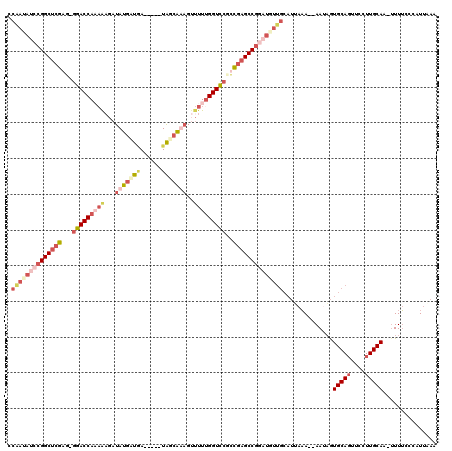
| Location | 20,348,356 – 20,348,475 |
|---|---|
| Length | 119 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 69.41 |
| Mean single sequence MFE | -32.88 |
| Consensus MFE | -20.00 |
| Energy contribution | -22.56 |
| Covariance contribution | 2.56 |
| Combinations/Pair | 1.17 |
| Mean z-score | -3.04 |
| Structure conservation index | 0.61 |
| SVM decision value | 4.82 |
| SVM RNA-class probability | 0.999953 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 20348356 119 - 22224390 UUGAAUGGUAAAA-UUGCAAGGAAGUGCACAAUUUUUUUCAUGCAACAUCCGGCUCGGCGAACCAAAAACUUUGAUAUCAUAUCAUCAUAUCUCUUUGGUCUGGUCGAGCCUUCUCUCGA ((((..((.....-(((((.(((((..........))))).)))))...))((((((((.(((((((.....((((........))))......)))))).).)))))))).....)))) ( -31.50) >DroSec_CAF1 16277 111 - 1 UUGAAUGGGAAAA-UUGCAAGGAACUGCACUAUU--UUUAAUGCAACAUCCGGCUUGGCGGACCGAAAACUUUGCUA-----UCAUCAUAUCAUUUUGGUCC-CUCGAGCCGGAUAUUGG ........(((((-(((((......))))..)))--)))....(((.(((((((((((.(((((((((....((.((-----(....))).)))))))))))-.))))))))))).))). ( -37.50) >DroEre_CAF1 13050 100 - 1 AUUAAUGGGCAAA-UUGCAAGCAACUGCAUUAUU--UA-----------CCGGCUCAAUGGACCAAAAUCUACGCUA-----UAAACAUAUCUUUAUGGUCC-CUCGAGCCGGAUUAUGG ........(((..-(((....))).)))......--..-----------(((((((...((((((.((.........-----...........)).))))))-...)))))))....... ( -26.05) >DroYak_CAF1 16826 112 - 1 UUUAAUGGGAAAUUUUGCAAAGAACUGCACUGUU--UUUAAUGCAAUAUGCGGCACGAAGGACCAAAGACAACCACA-----UUAACAUAUAUUGGUGGUCC-CUCGAGCCGGAUAUUGU .......((((((..((((......))))..)))--)))...(((((((.((((.(((.((((((....(((.....-----..........))).))))))-.))).)))).))))))) ( -36.46) >consensus UUGAAUGGGAAAA_UUGCAAGGAACUGCACUAUU__UUUAAUGCAACAUCCGGCUCGACGGACCAAAAACUACGAUA_____UCAACAUAUCUUUUUGGUCC_CUCGAGCCGGAUAUUGG ...............((((......))))..................((((((((((..(((((((((.........................)))))))))...))))))))))..... (-20.00 = -22.56 + 2.56)

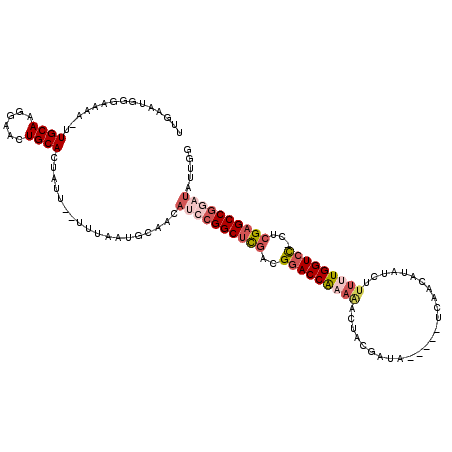
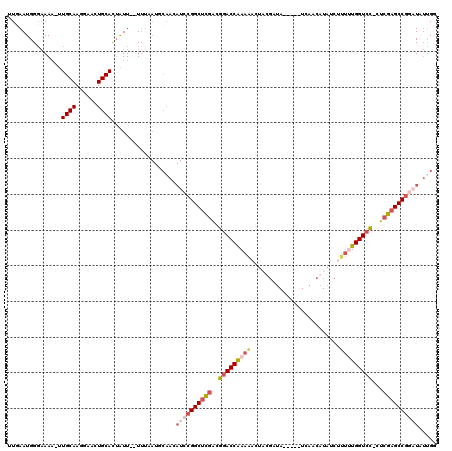
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:58:01 2006