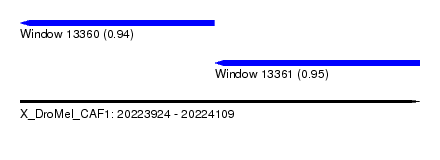
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 20,223,924 – 20,224,109 |
| Length | 185 |
| Max. P | 0.951497 |

| Location | 20,223,924 – 20,224,014 |
|---|---|
| Length | 90 |
| Sequences | 5 |
| Columns | 97 |
| Reading direction | reverse |
| Mean pairwise identity | 72.16 |
| Mean single sequence MFE | -12.68 |
| Consensus MFE | -9.88 |
| Energy contribution | -9.88 |
| Covariance contribution | -0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.07 |
| Structure conservation index | 0.78 |
| SVM decision value | 1.25 |
| SVM RNA-class probability | 0.936398 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 20223924 90 - 22224390 -----AAUUAUUAAUUUAACAUUUUUUUACAUACAUUUUUUACAAAUACAUAUACUAUGAC--CCAUUUCGCAGGUGGCAAAGGCCACAUACGCCAC -----........................................................--.......((..(((((....)))))....))... ( -9.40) >DroSec_CAF1 10791 88 - 1 -----AAUUAUUAAUUUAACAUUUUAAGUCAAAUAUUUUU--CAAAUACAC--ACUUUGACCCCCAUUUUGCAGGUGGCAAAGGCCACAUACGCCAC -----......................((((((.......--.........--..)))))).........((..(((((....)))))....))... ( -11.77) >DroSim_CAF1 10036 88 - 1 -----AAUUAUUAAUUCAACAUUUUAAGUCAAAUAUUUUU--CAAAUACAC--ACUUUGACCCCCAUUUUGCAGGUGGCAAAGGCCACAUACGCCAC -----......................((((((.......--.........--..)))))).........((..(((((....)))))....))... ( -11.77) >DroEre_CAF1 10819 84 - 1 UGAUAAAUAGUGAAUU--AAAUUGUAAUUUAAAUACUCCG--CUGAUAC------G---AGACUCAAUUCGCAGGUGGCCAAGGCCACAUACGCCAC .........(((((((--...(((((..............--....)))------)---).....)))))))..(((((....)))))......... ( -16.87) >DroYak_CAF1 10434 85 - 1 CUAUAAAUAACUAAUU--ACAUUUAAAAUGAAGCACUUCG--CUGAUAC------UA--AGACUCAUUUCGCAGGUGGCCAAGGCCACAUACGCCAC ................--.......(((((((((.....)--)).....------..--....)))))).((..(((((....)))))....))... ( -13.60) >consensus _____AAUUAUUAAUU_AACAUUUUAAGUCAAAUAUUUUU__CAAAUACA___ACUAUGACACCCAUUUCGCAGGUGGCAAAGGCCACAUACGCCAC ......................................................................((..(((((....)))))....))... ( -9.88 = -9.88 + -0.00)


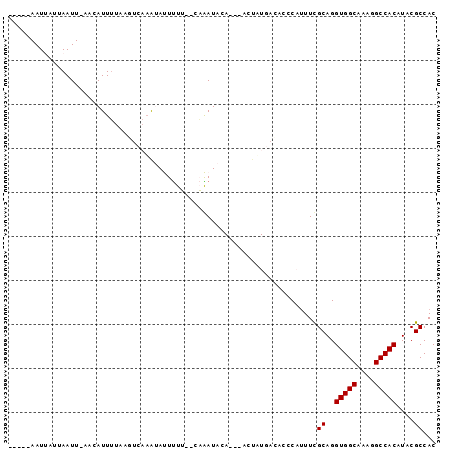
| Location | 20,224,014 – 20,224,109 |
|---|---|
| Length | 95 |
| Sequences | 3 |
| Columns | 95 |
| Reading direction | reverse |
| Mean pairwise identity | 92.98 |
| Mean single sequence MFE | -21.33 |
| Consensus MFE | -21.51 |
| Energy contribution | -20.63 |
| Covariance contribution | -0.88 |
| Combinations/Pair | 1.19 |
| Mean z-score | -1.91 |
| Structure conservation index | 1.01 |
| SVM decision value | 1.41 |
| SVM RNA-class probability | 0.951497 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 20224014 95 - 22224390 UGAUUCUAUUGAACGCACACAAGAUAAAAUGAACCAUCGCUUUGGAGCUGUUGAUGGACACAAAAUAUUAUUUUCAGCUCCAAAUUGAAAUUCAA .............................((((...(((.((((((((((..(((((..........)))))..)))))))))).)))..)))). ( -20.60) >DroSec_CAF1 10879 95 - 1 UGAUUCUAUUGAACGCACACAAGAUUACAUGAGCCAUCGCUUUGGAGCUGUUGAUGAACACAAAAGAACGUUUUCAGCUCCAAAAUGAAAUUCGA ........(((((.((.((..........)).))..(((.((((((((((..((((..(......)..))))..)))))))))).)))..))))) ( -21.70) >DroSim_CAF1 10124 95 - 1 UGAUUCUAUUGAACGCACACAAGAUUACAUGAGCCAUCGCUUUGGAGCUGUUGAUGAACACAAAAGAACGUUUUCAGCUCCAAAAUGAAAUUCGA ........(((((.((.((..........)).))..(((.((((((((((..((((..(......)..))))..)))))))))).)))..))))) ( -21.70) >consensus UGAUUCUAUUGAACGCACACAAGAUUACAUGAGCCAUCGCUUUGGAGCUGUUGAUGAACACAAAAGAACGUUUUCAGCUCCAAAAUGAAAUUCGA .............................((((...(((.((((((((((..((((............))))..)))))))))).)))..)))). (-21.51 = -20.63 + -0.88)


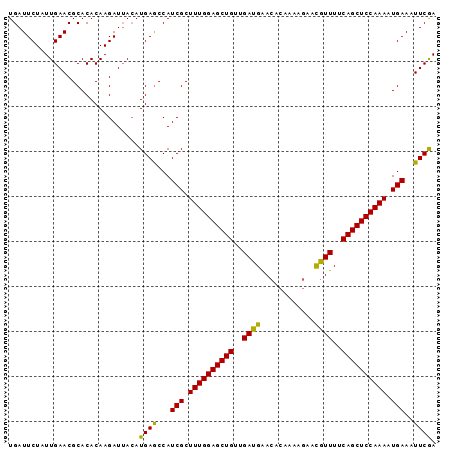
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:56:30 2006