
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 20,152,399 – 20,152,503 |
| Length | 104 |
| Max. P | 0.992075 |

| Location | 20,152,399 – 20,152,503 |
|---|---|
| Length | 104 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 82.03 |
| Mean single sequence MFE | -34.74 |
| Consensus MFE | -19.30 |
| Energy contribution | -20.38 |
| Covariance contribution | 1.08 |
| Combinations/Pair | 1.08 |
| Mean z-score | -3.10 |
| Structure conservation index | 0.56 |
| SVM decision value | 1.86 |
| SVM RNA-class probability | 0.980433 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 20152399 104 + 22224390 -------AUGCUGGC----GCAGGAGUGGAUGCAUGAGUUGAUGCCAACAACGUAUUAAUUUUGUGUAUCAAAAUAAACUUAUUGUUCUGGCAUUCUGCCCG-----CUCCAAGCCAGCC -------..((((((----...((((((((((((..(((((((((.......))))))).))..))))))...................(((.....)))))-----))))..)))))). ( -40.60) >DroSec_CAF1 7499 108 + 1 -------AUGCGGAUGCUGGCAGGAGUGGAUGCAUGAGUUGAUGCCAACAACGUAUUAAUUUUGUGUAUCAAAAUAAACUUAUUGUUCUGGCAUUCUGCCCG-----CUCCAAGCCAGCC -------........((((((.((((((((((((..(((((((((.......))))))).))..))))))...................(((.....)))))-----))))..)))))). ( -39.40) >DroSim_CAF1 12578 108 + 1 -------AUGCGGAUGCUGGCAGGAGUGGAUGCAUGAGUUGAUGCCAACAACGUAUUAAUUUUGUGUAUCAAAAUAAACUUAUUGUUCUGGCAUUCUGCCCG-----CUCCAAACCAGCC -------........(((((..((((((((((((..(((((((((.......))))))).))..))))))...................(((.....)))))-----))))...))))). ( -36.10) >DroEre_CAF1 3094 97 + 1 -----------------------AGCUGGAUGCAUGAGCUGAUGCCAACAACGUAUUAAUUUUGUGUAUCAAAAUAAACUUAUUGUUCUGGCAUUCAGCCCGGCCCGCUCCAAGCCAGCC -----------------------.(((((..((.((.((((((((((((((.(..((.((((((.....)))))).))..).))))..))))).))))).))....))......))))). ( -28.50) >DroYak_CAF1 5630 104 + 1 ACUGACAAGGUGGAUGCUGGCAGGAGUGGAUGCAUGAGUUGAUGCCAACAACGUAUUAAUUUUGUGUAUCAAAAUAAACUUAUUGUUCUGGCAUUCUGCCCG-----CU----------- ........((..(((((..((((((((.((((((..(((((((((.......))))))).))..)))))).......)))).)))..)..))).))..))..-----..----------- ( -29.10) >consensus _______AUGCGGAUGCUGGCAGGAGUGGAUGCAUGAGUUGAUGCCAACAACGUAUUAAUUUUGUGUAUCAAAAUAAACUUAUUGUUCUGGCAUUCUGCCCG_____CUCCAAGCCAGCC ..................(((((((...((((((..(((((((((.......))))))).))..)))))).............(((....)))))))))).................... (-19.30 = -20.38 + 1.08)
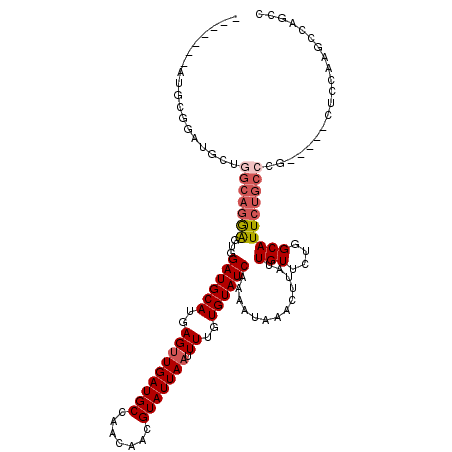


| Location | 20,152,399 – 20,152,503 |
|---|---|
| Length | 104 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 82.03 |
| Mean single sequence MFE | -35.20 |
| Consensus MFE | -17.30 |
| Energy contribution | -18.38 |
| Covariance contribution | 1.08 |
| Combinations/Pair | 1.03 |
| Mean z-score | -3.74 |
| Structure conservation index | 0.49 |
| SVM decision value | 2.31 |
| SVM RNA-class probability | 0.992075 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 20152399 104 - 22224390 GGCUGGCUUGGAG-----CGGGCAGAAUGCCAGAACAAUAAGUUUAUUUUGAUACACAAAAUUAAUACGUUGUUGGCAUCAACUCAUGCAUCCACUCCUGC----GCCAGCAU------- .((((((..((((-----.(((((((((((((..(((((..(((.((((((.....)))))).)))..)))))))))))....)).)))..)).))))...----))))))..------- ( -38.60) >DroSec_CAF1 7499 108 - 1 GGCUGGCUUGGAG-----CGGGCAGAAUGCCAGAACAAUAAGUUUAUUUUGAUACACAAAAUUAAUACGUUGUUGGCAUCAACUCAUGCAUCCACUCCUGCCAGCAUCCGCAU------- .((((((..((((-----.(((((((((((((..(((((..(((.((((((.....)))))).)))..)))))))))))....)).)))..)).)))).))))))........------- ( -39.00) >DroSim_CAF1 12578 108 - 1 GGCUGGUUUGGAG-----CGGGCAGAAUGCCAGAACAAUAAGUUUAUUUUGAUACACAAAAUUAAUACGUUGUUGGCAUCAACUCAUGCAUCCACUCCUGCCAGCAUCCGCAU------- .((((((..((((-----.(((((((((((((..(((((..(((.((((((.....)))))).)))..)))))))))))....)).)))..)).)))).))))))........------- ( -37.30) >DroEre_CAF1 3094 97 - 1 GGCUGGCUUGGAGCGGGCCGGGCUGAAUGCCAGAACAAUAAGUUUAUUUUGAUACACAAAAUUAAUACGUUGUUGGCAUCAGCUCAUGCAUCCAGCU----------------------- ((((.((.....)).))))(((((((.(((((..(((((..(((.((((((.....)))))).)))..)))))))))))))))))..((.....)).----------------------- ( -37.20) >DroYak_CAF1 5630 104 - 1 -----------AG-----CGGGCAGAAUGCCAGAACAAUAAGUUUAUUUUGAUACACAAAAUUAAUACGUUGUUGGCAUCAACUCAUGCAUCCACUCCUGCCAGCAUCCACCUUGUCAGU -----------.(-----(.(((((.((((((..(((((..(((.((((((.....)))))).)))..))))))))))).......((....))...))))).))............... ( -23.90) >consensus GGCUGGCUUGGAG_____CGGGCAGAAUGCCAGAACAAUAAGUUUAUUUUGAUACACAAAAUUAAUACGUUGUUGGCAUCAACUCAUGCAUCCACUCCUGCCAGCAUCCGCAU_______ ....((..((((.......(((..((.(((((..(((((..(((.((((((.....)))))).)))..))))))))))))..))).....))))..))...................... (-17.30 = -18.38 + 1.08)
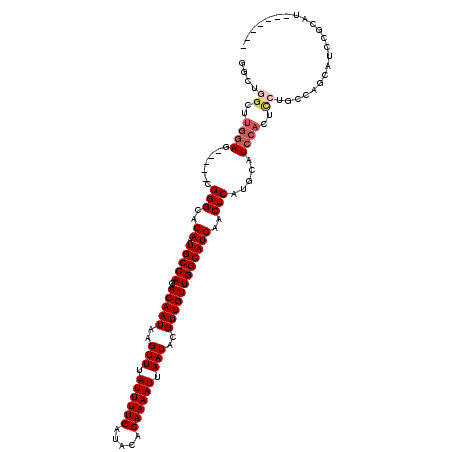


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:55:20 2006