
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 20,096,594 – 20,096,765 |
| Length | 171 |
| Max. P | 0.978666 |

| Location | 20,096,594 – 20,096,685 |
|---|---|
| Length | 91 |
| Sequences | 3 |
| Columns | 91 |
| Reading direction | forward |
| Mean pairwise identity | 97.07 |
| Mean single sequence MFE | -22.07 |
| Consensus MFE | -21.50 |
| Energy contribution | -21.83 |
| Covariance contribution | 0.33 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.65 |
| Structure conservation index | 0.97 |
| SVM decision value | 1.82 |
| SVM RNA-class probability | 0.978666 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
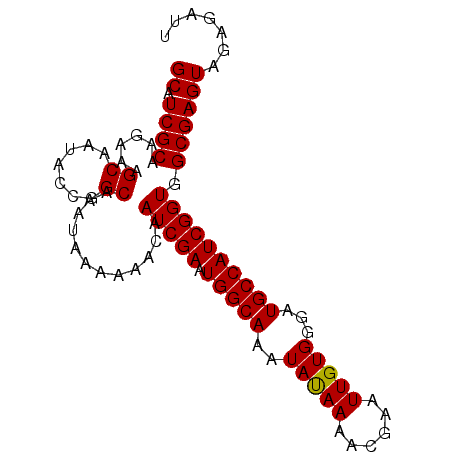
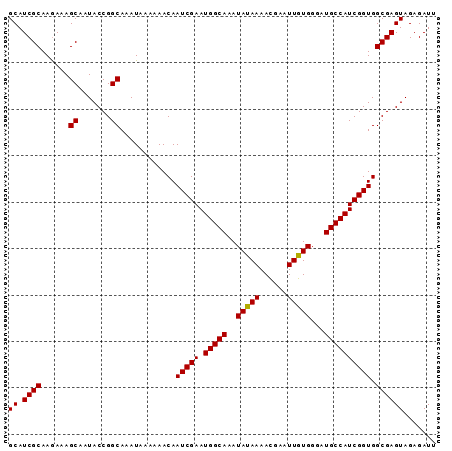
>X_DroMel_CAF1 20096594 91 + 22224390 AAAUGGCGCGAUGACGAGAGUGAGUUUUAAAAAUGCAACCAGUGCAGAAAAGGACGUGGUGAACAGGUCGAAAAAGCGAGCAUCGCAAGAA .......((((((((....))..(((((.....((((.....))))......(((.((.....)).)))...)))))...))))))..... ( -23.30) >DroSec_CAF1 6463 90 + 1 AAAUGGCGC-AUGACGAGAGUGAGUUUUAAAAAUGCAACCAGUGCAGAAAAAGACGUGGUGAACAGGUCGAAAAAGCGAGCAUCGCAAAAA .....(((.-(((((....))..(((((.....((((.....))))......(((.((.....)).)))...)))))...))))))..... ( -19.40) >DroSim_CAF1 6485 91 + 1 AAGUGGCGCGAUGACGAGAGUGAGUUUUAAAAAUGCAACCAGUGCAGAAAAAGACGUGGUGAACAGGUCGAAAAAGCGAGCAUCGCAAGAA .......((((((((....))..(((((.....((((.....))))......(((.((.....)).)))...)))))...))))))..... ( -23.50) >consensus AAAUGGCGCGAUGACGAGAGUGAGUUUUAAAAAUGCAACCAGUGCAGAAAAAGACGUGGUGAACAGGUCGAAAAAGCGAGCAUCGCAAGAA .......((((((((....))..(((((.....((((.....))))......(((.((.....)).)))...)))))...))))))..... (-21.50 = -21.83 + 0.33)



| Location | 20,096,673 – 20,096,765 |
|---|---|
| Length | 92 |
| Sequences | 3 |
| Columns | 92 |
| Reading direction | forward |
| Mean pairwise identity | 97.10 |
| Mean single sequence MFE | -20.73 |
| Consensus MFE | -20.95 |
| Energy contribution | -20.73 |
| Covariance contribution | -0.22 |
| Combinations/Pair | 1.04 |
| Mean z-score | -2.19 |
| Structure conservation index | 1.01 |
| SVM decision value | 1.55 |
| SVM RNA-class probability | 0.963103 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 20096673 92 + 22224390 GCAUCGCAAGAAAGCAAUACCGGCAAAUAAAAAACAAUCGAAUGGCAAAUACAAAAUGAAUUGUGGGAUGCCAUCGGUGGCGAGUAGAGAUG ((.((((......((.......))............(((((.(((((..(((((......)))))...)))))))))).))))))....... ( -22.00) >DroSec_CAF1 6541 92 + 1 GCAUCGCAAAAAAGCAAUACCGGCAAAUAAAAAACAAUCGAAUGGCAAAUAUAAAACGAAUUGUGGGAUGCCAUCGGUGGCGAGUAGAGAUU ((.((((......((.......))............(((((.(((((..(((((......)))))...)))))))))).))))))....... ( -20.10) >DroSim_CAF1 6564 92 + 1 GCAUCGCAAGAAAGCAAUACCGGCAAAUAAAAAACAAUCGAAUGGCAAAUAUAAAACGAAUUGUGGGAUGCCAUCGGUGGCGAGUAGAGAUU ((.((((......((.......))............(((((.(((((..(((((......)))))...)))))))))).))))))....... ( -20.10) >consensus GCAUCGCAAGAAAGCAAUACCGGCAAAUAAAAAACAAUCGAAUGGCAAAUAUAAAACGAAUUGUGGGAUGCCAUCGGUGGCGAGUAGAGAUU ((.((((......((.......))............(((((.(((((..(((((......)))))...)))))))))).))))))....... (-20.95 = -20.73 + -0.22)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:54:34 2006