
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 19,992,624 – 19,992,715 |
| Length | 91 |
| Max. P | 0.697109 |

| Location | 19,992,624 – 19,992,715 |
|---|---|
| Length | 91 |
| Sequences | 5 |
| Columns | 93 |
| Reading direction | forward |
| Mean pairwise identity | 80.22 |
| Mean single sequence MFE | -20.56 |
| Consensus MFE | -13.60 |
| Energy contribution | -15.32 |
| Covariance contribution | 1.72 |
| Combinations/Pair | 1.14 |
| Mean z-score | -1.75 |
| Structure conservation index | 0.66 |
| SVM decision value | 0.34 |
| SVM RNA-class probability | 0.697109 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
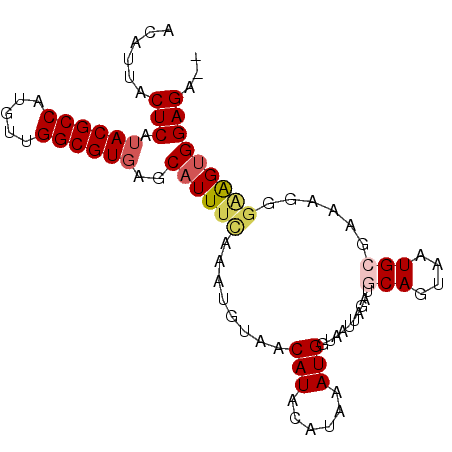
>X_DroMel_CAF1 19992624 91 + 22224390 ACAUUACUCAUACGCCAUGUUGGCGUUAGCAUUUUAAAUGUAACAUACAUUAAUGGUAAUCAGAUGCAGUAAUGCGAAAGGGAGGUGGAGA-- .(((..(((..(((((.....)))))((.((((...((((((...)))))))))).))......((((....))))...)))..)))....-- ( -21.40) >DroSec_CAF1 30133 93 + 1 ACAUUACUCAUACGCCAUGUUGGCGUGAGCAUUUCAAAUGUAACAUACAUAAAUGGUAGCUAGAUUCAGUAAUGCGAAAGGAAAGUGGAGACA .(((((((..((((((.....))))))((((((....(((((...)))))....))).)))......)))))))(....).....((....)) ( -21.50) >DroSim_CAF1 23275 93 + 1 ACAUUACUCAUACGCCAUGUUGGCGUGAGCAUUUCAAAUGUAACAUACAUAAAUGGUAGCUAGAUGCAGCAAUGCGAAAGGGAAGUGGAGACA ......(((.((((((.....))))))..((((((..(((((...)))))..............((((....)))).....)))))))))... ( -23.80) >DroEre_CAF1 25009 87 + 1 ACAUUACUCAUACGCCAUGUUGGCGUGAACAUUUCAAAUGUAACAU----AAAUGGUAAUUAGAUGCAAUAAUGCCAAACGGCGGUGGAGA-- ......(((...((((...(((((((..((((.....)))).....----.....(((......)))....)))))))..))))...))).-- ( -22.60) >DroYak_CAF1 23342 72 + 1 ACAUUACUCAUACGCCAUGUUGGCGUGAACAUUUCAAAUGUAACAU----AAAUGACAAUUAGAGA---------------GGAGAGGAGA-- ......(((.((((((.....))))))....((((((.(((.....----.....))).)).))))---------------...)))....-- ( -13.50) >consensus ACAUUACUCAUACGCCAUGUUGGCGUGAGCAUUUCAAAUGUAACAUACAUAAAUGGUAAUUAGAUGCAGUAAUGCGAAAGGGAAGUGGAGA__ ......(((.((((((.....))))))..((((((........(((......)))..........(((....)))......)))))))))... (-13.60 = -15.32 + 1.72)


| Location | 19,992,624 – 19,992,715 |
|---|---|
| Length | 91 |
| Sequences | 5 |
| Columns | 93 |
| Reading direction | reverse |
| Mean pairwise identity | 80.22 |
| Mean single sequence MFE | -17.58 |
| Consensus MFE | -11.62 |
| Energy contribution | -13.18 |
| Covariance contribution | 1.56 |
| Combinations/Pair | 1.10 |
| Mean z-score | -1.58 |
| Structure conservation index | 0.66 |
| SVM decision value | 0.00 |
| SVM RNA-class probability | 0.535087 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
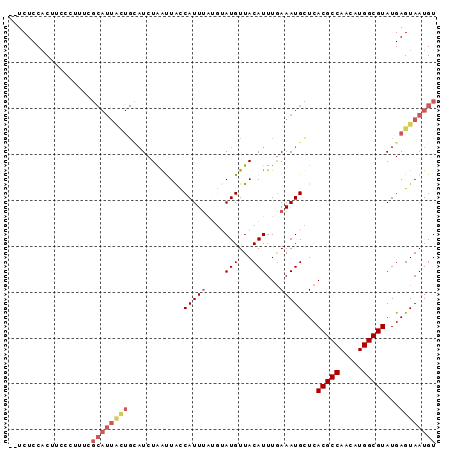
>X_DroMel_CAF1 19992624 91 - 22224390 --UCUCCACCUCCCUUUCGCAUUACUGCAUCUGAUUACCAUUAAUGUAUGUUACAUUUAAAAUGCUAACGCCAACAUGGCGUAUGAGUAAUGU --................((((((((.(((........((((((((((...))))))...))))...(((((.....)))))))))))))))) ( -19.00) >DroSec_CAF1 30133 93 - 1 UGUCUCCACUUUCCUUUCGCAUUACUGAAUCUAGCUACCAUUUAUGUAUGUUACAUUUGAAAUGCUCACGCCAACAUGGCGUAUGAGUAAUGU ..................((((((((.(....(((...((...(((((...))))).))....))).(((((.....))))).).)))))))) ( -19.60) >DroSim_CAF1 23275 93 - 1 UGUCUCCACUUCCCUUUCGCAUUGCUGCAUCUAGCUACCAUUUAUGUAUGUUACAUUUGAAAUGCUCACGCCAACAUGGCGUAUGAGUAAUGU ..................((((((((.(((..(((...((...(((((...))))).))....))).(((((.....)))))))))))))))) ( -20.20) >DroEre_CAF1 25009 87 - 1 --UCUCCACCGCCGUUUGGCAUUAUUGCAUCUAAUUACCAUUU----AUGUUACAUUUGAAAUGUUCACGCCAACAUGGCGUAUGAGUAAUGU --.(((...(((((((((((......((((..(((....))).----)))).((((.....))))....))))).))))))...)))...... ( -18.40) >DroYak_CAF1 23342 72 - 1 --UCUCCUCUCC---------------UCUCUAAUUGUCAUUU----AUGUUACAUUUGAAAUGUUCACGCCAACAUGGCGUAUGAGUAAUGU --..........---------------................----(((((((....((.....))(((((.....)))))....))))))) ( -10.70) >consensus __UCUCCACUUCCCUUUCGCAUUACUGCAUCUAAUUACCAUUUAUGUAUGUUACAUUUGAAAUGCUCACGCCAACAUGGCGUAUGAGUAAUGU ..................((((((((............(((((....(((...)))...)))))...(((((.....)))))...)))))))) (-11.62 = -13.18 + 1.56)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:53:55 2006