
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 19,940,582 – 19,940,673 |
| Length | 91 |
| Max. P | 0.808443 |

| Location | 19,940,582 – 19,940,673 |
|---|---|
| Length | 91 |
| Sequences | 5 |
| Columns | 95 |
| Reading direction | reverse |
| Mean pairwise identity | 83.35 |
| Mean single sequence MFE | -23.64 |
| Consensus MFE | -12.92 |
| Energy contribution | -14.24 |
| Covariance contribution | 1.32 |
| Combinations/Pair | 1.11 |
| Mean z-score | -2.59 |
| Structure conservation index | 0.55 |
| SVM decision value | 0.64 |
| SVM RNA-class probability | 0.808443 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 19940582 91 - 22224390 UUGGCAAUGUCACGUAAGCAACGUGGCAGCAACAACGGCAACAACGGCAACAACUGUGGCAAACU--ACAAUGAACUG--AAUAGCGUUGCACAA (((....((((((((.....))))))))....)))..(((((...(....)...(((((....))--)))........--......))))).... ( -24.40) >DroSim_CAF1 46304 82 - 1 UUGGCAAUGUCACGUAAGCAACGUGGCAGCAACAACG---------GCAACAACUGCGGCAAACU--ACAAUGAACUG--AAUAGCGUUGCACAA .......((((((((.....))))))))(((((..(.---------(((.....))).)......--.((......))--......))))).... ( -19.90) >DroEre_CAF1 43879 91 - 1 UUGGCAAUGUCACGUAAGCAACGUGGCAGCAACAACUGCAACAACUGCAACAACUGCGGCAAGCU--ACAAUGAACUG--AAUAGCGUUGCACAA (((....((((((((.....))))))))....))).((((((..(((((.....)))))...(((--(((......))--..))))))))))... ( -27.70) >DroYak_CAF1 45559 82 - 1 UUGGCAAUGUCACGUAAGCAACGUGGCAGCAACAACUG---------CAACAACUGCGGCAAACU--ACAAUGAACUG--AAUAGCGUUGCACAA .......((((((((.....))))))))(((((..(((---------((.....)))))......--.((......))--......))))).... ( -22.00) >DroAna_CAF1 71547 84 - 1 UUGGCAAUGUCACGUAAGCAAC-----AAAAACA------UGGCCUGCAACCACUGCGGGCAACUCAACAAUGAACUGGGAACAGCGUUGCACAA .((((...)))).....(((((-----.......------..(((((((.....)))))))..............(((....))).))))).... ( -24.20) >consensus UUGGCAAUGUCACGUAAGCAACGUGGCAGCAACAACGG______C_GCAACAACUGCGGCAAACU__ACAAUGAACUG__AAUAGCGUUGCACAA .......((((((((.....))))))))(((((.............(((.....)))..................(((....))).))))).... (-12.92 = -14.24 + 1.32)

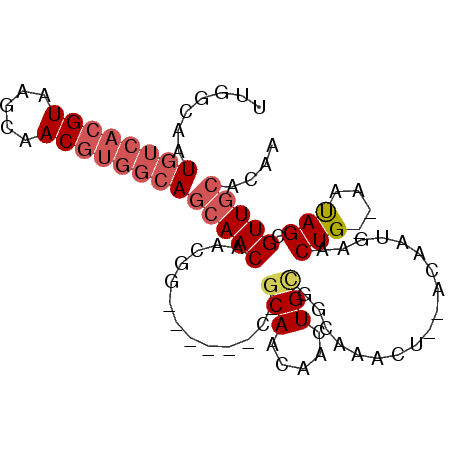

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:53:26 2006