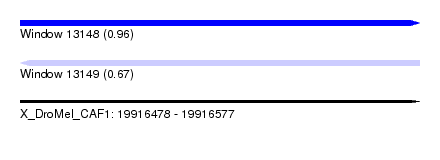

| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 19,916,478 – 19,916,577 |
| Length | 99 |
| Max. P | 0.963128 |

| Location | 19,916,478 – 19,916,577 |
|---|---|
| Length | 99 |
| Sequences | 4 |
| Columns | 108 |
| Reading direction | forward |
| Mean pairwise identity | 93.72 |
| Mean single sequence MFE | -38.80 |
| Consensus MFE | -34.38 |
| Energy contribution | -36.88 |
| Covariance contribution | 2.50 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.35 |
| Structure conservation index | 0.89 |
| SVM decision value | 1.55 |
| SVM RNA-class probability | 0.963128 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 19916478 99 + 22224390 GGCUCAAUCAAGGCGAUGUGUCGCCGGUCUGAUUGGAAGCA---------GCAGCUGAGAAGCCGCAAGGAUCGUGCCUGGGUGAGCCAUAACACAUAGUGGUGACAC ((((((..((.(((((....)))))(((.((((((....).---------((.(((....))).))...))))).)))))..))))))..........(((....))) ( -38.70) >DroSec_CAF1 30551 99 + 1 GGCUCAAUCAAGGCGAUGUGUCGCCGGUCUGAUUGGAAACA---------GCAGCUGAGAAGCCGCAAGGAUCGUGCCUGGGUGAGCCAUAACACAAAGUGGUGACAC ((((((..((.(((((....)))))(((.((((((....).---------((.(((....))).))...))))).)))))..))))))..........(((....))) ( -41.20) >DroEre_CAF1 28042 99 + 1 GGCUCAAUCAAGGCGAUGUGUCGCCCGUCUGAUUGGAAGCA---------GCAGCUGAGAAGCCGCAAGGAUCGUGCCUGGGUGAGCCAUAACACAAAGUGGUAACAC ((((((..((.(((.(((.((((((((......)))..)).---------((.(((....))).))...)))))))))))..))))))..........(((....))) ( -32.40) >DroYak_CAF1 29828 108 + 1 GGCUCAAUCAAGGCGAUGUGUCGCCGGUCUGAUUGGAAGCAGCAGCAGCAGCAGCUGAGAAGCCGCAAGGAUCGUGCCUGGGUGAGCCAUAACACAAAGUGGUGACAC ((((((..((.(((((....)))))(((.((((......((((.((....)).)))).....((....)))))).)))))..))))))..........(((....))) ( -42.90) >consensus GGCUCAAUCAAGGCGAUGUGUCGCCGGUCUGAUUGGAAGCA_________GCAGCUGAGAAGCCGCAAGGAUCGUGCCUGGGUGAGCCAUAACACAAAGUGGUGACAC ((((((..((.(((((....)))))(((.((((((....)..........((.(((....))).))...))))).)))))..))))))..........(((....))) (-34.38 = -36.88 + 2.50)


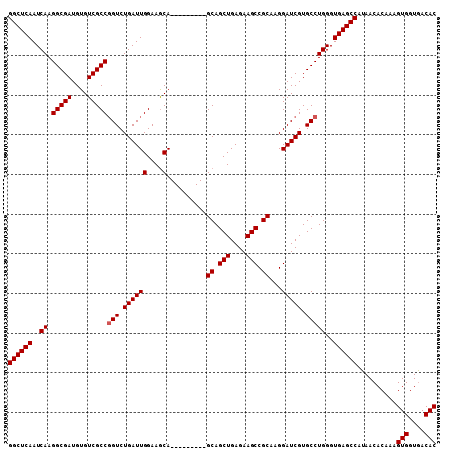
| Location | 19,916,478 – 19,916,577 |
|---|---|
| Length | 99 |
| Sequences | 4 |
| Columns | 108 |
| Reading direction | reverse |
| Mean pairwise identity | 93.72 |
| Mean single sequence MFE | -31.98 |
| Consensus MFE | -30.16 |
| Energy contribution | -29.78 |
| Covariance contribution | -0.38 |
| Combinations/Pair | 1.06 |
| Mean z-score | -1.43 |
| Structure conservation index | 0.94 |
| SVM decision value | 0.28 |
| SVM RNA-class probability | 0.666240 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 19916478 99 - 22224390 GUGUCACCACUAUGUGUUAUGGCUCACCCAGGCACGAUCCUUGCGGCUUCUCAGCUGC---------UGCUUCCAAUCAGACCGGCGACACAUCGCCUUGAUUGAGCC (((((..(((...)))...(((.....)))))))).......((((((....))))))---------.(((..(((((((...(((((....))))))))))))))). ( -30.50) >DroSec_CAF1 30551 99 - 1 GUGUCACCACUUUGUGUUAUGGCUCACCCAGGCACGAUCCUUGCGGCUUCUCAGCUGC---------UGUUUCCAAUCAGACCGGCGACACAUCGCCUUGAUUGAGCC (.((((..((.....))..)))).).....(((..((..(..((((((....))))))---------.)..))(((((((...(((((....)))))))))))).))) ( -30.70) >DroEre_CAF1 28042 99 - 1 GUGUUACCACUUUGUGUUAUGGCUCACCCAGGCACGAUCCUUGCGGCUUCUCAGCUGC---------UGCUUCCAAUCAGACGGGCGACACAUCGCCUUGAUUGAGCC (((....))).(((((((.(((.....)))))))))).....((((((....))))))---------.(((..((((((...((((((....))))))))))))))). ( -32.10) >DroYak_CAF1 29828 108 - 1 GUGUCACCACUUUGUGUUAUGGCUCACCCAGGCACGAUCCUUGCGGCUUCUCAGCUGCUGCUGCUGCUGCUUCCAAUCAGACCGGCGACACAUCGCCUUGAUUGAGCC (((....))).(((((((.(((.....)))))))))).....(((((....((((....))))..)))))...(((((((...(((((....)))))))))))).... ( -34.60) >consensus GUGUCACCACUUUGUGUUAUGGCUCACCCAGGCACGAUCCUUGCGGCUUCUCAGCUGC_________UGCUUCCAAUCAGACCGGCGACACAUCGCCUUGAUUGAGCC (((.(((......))).)))(((((....(((((........((((((....)))))).........)))))..((((((...(((((....)))))))))))))))) (-30.16 = -29.78 + -0.38)

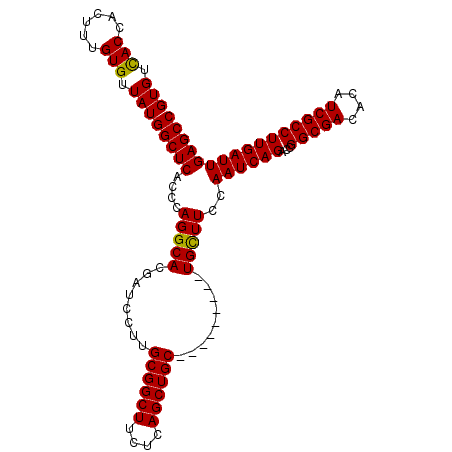

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:53:12 2006