| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 2,237,173 – 2,237,278 |
| Length | 105 |
| Max. P | 0.986289 |

| Location | 2,237,173 – 2,237,278 |
|---|---|
| Length | 105 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 86.79 |
| Mean single sequence MFE | -43.95 |
| Consensus MFE | -40.83 |
| Energy contribution | -41.22 |
| Covariance contribution | 0.39 |
| Combinations/Pair | 1.06 |
| Mean z-score | -2.24 |
| Structure conservation index | 0.93 |
| SVM decision value | 2.04 |
| SVM RNA-class probability | 0.986289 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 2237173 105 - 22224390 CACGAUGCCCGAGGCUGAUGUUCUGAAUUUGAUACGAUCGCAGGGCUCGUCCUUGACGUUGGAGAUAUUCAGAAGGUCAGGCGCGGGCGCCACAACAAUCACGU---C------------ ...(.((((((.(.(((((.((((((((......(((.(((((((.....))))).))))).....)))))))).))))).).)))))).).............---.------------ ( -41.30) >DroVir_CAF1 305 114 - 1 CACGAUGCCCGAGGCUGAUGUUCUGAAUUUGAUACGAUCGCAGGGCUCGUCCUUGACGUUGGAGAUAUUCAGAAGGUCAGGCGCGGGCGCCAACAUGGCCAGCAAUGCCGCCAA------ ...(.((((((.(.(((((.((((((((......(((.(((((((.....))))).))))).....)))))))).))))).).)))))).)....((((((....))..)))).------ ( -47.00) >DroPse_CAF1 416 114 - 1 CACGAUGCCCGAGGCUGAUGUUCUGAAUUUGAUACGAUCGCAGGGCUCGACCUUGACGUUGGAGAUAUUCAGAAGGUCAGGCGCGGGCGCCACCAUGGCCAUAGACACAGCCAC------ ...(.((((((.(.(((((.((((((((......(((.(((((((.....))))).))))).....)))))))).))))).).)))))).)....((((..........)))).------ ( -45.50) >DroWil_CAF1 351 107 - 1 CACGAUGCCCGAGGCUGAUGUUCUGAAUUUGAUACGAUCGCAGGGCUCGUCCUUGACGUUGGAGAUAUUCAGAAGGUCAGGAUCGGGUGCCAAUAUGGCCACACACC------------- ......((((((..(((((.((((((((......(((.(((((((.....))))).))))).....)))))))).)))))..))))))(((.....)))........------------- ( -41.90) >DroMoj_CAF1 341 120 - 1 UACGAUGCCCGAGGCUGAUGUUCUGAAUUUGAUACGAUCGCAGGGCUCGUCCUUAACGUUGGAGAUAUUCAGAAGGUCAGGCGCGGGCGCCAACAUGGCCACCAAUGCCACCAAUGCCAC ......(((((.(.(((((.((((((((......(((.((.((((....))))...))))).....)))))))).))))).).)))))((.....((((.......)))).....))... ( -42.50) >DroPer_CAF1 392 114 - 1 CACGAUGCCCGAGGCUGAUGUUCUGAAUUUGAUACGAUCGCAGGGCUCGACCUUGACGUUGGAGAUAUUCAGAAGGUCAGGCGCGGGCGCCACCAUGGCCAUAGACACAGCCAC------ ...(.((((((.(.(((((.((((((((......(((.(((((((.....))))).))))).....)))))))).))))).).)))))).)....((((..........)))).------ ( -45.50) >consensus CACGAUGCCCGAGGCUGAUGUUCUGAAUUUGAUACGAUCGCAGGGCUCGUCCUUGACGUUGGAGAUAUUCAGAAGGUCAGGCGCGGGCGCCAACAUGGCCACAAACAC_GCCA_______ ......(((((.(.(((((.((((((((......(((.(((((((.....))))).))))).....)))))))).))))).).)))))(((.....)))..................... (-40.83 = -41.22 + 0.39)

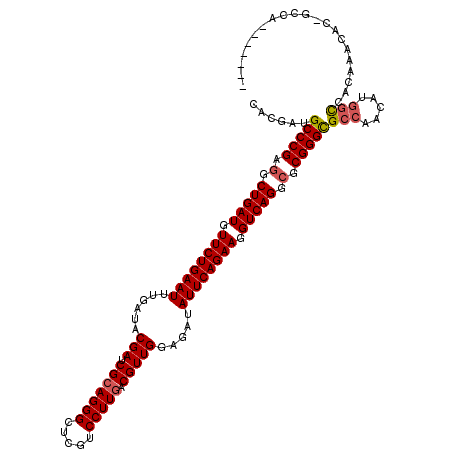
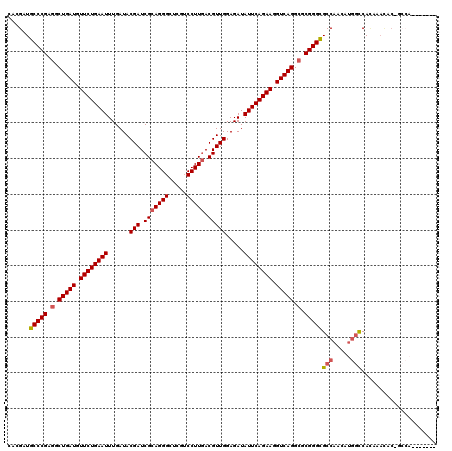
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:54:08 2006