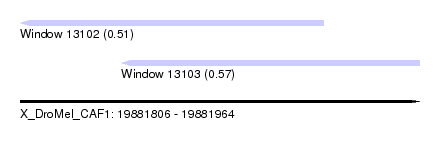

| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 19,881,806 – 19,881,964 |
| Length | 158 |
| Max. P | 0.569450 |

| Location | 19,881,806 – 19,881,926 |
|---|---|
| Length | 120 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 87.78 |
| Mean single sequence MFE | -29.89 |
| Consensus MFE | -20.90 |
| Energy contribution | -21.78 |
| Covariance contribution | 0.88 |
| Combinations/Pair | 1.16 |
| Mean z-score | -1.45 |
| Structure conservation index | 0.70 |
| SVM decision value | -0.05 |
| SVM RNA-class probability | 0.506137 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
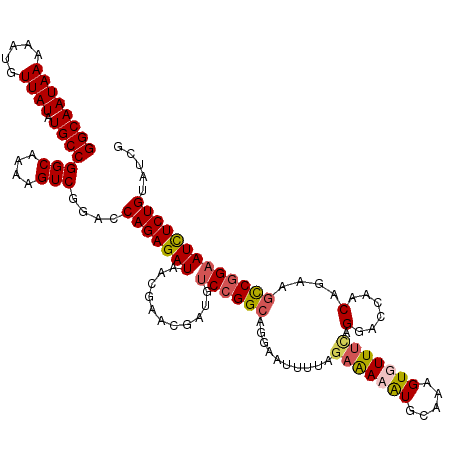
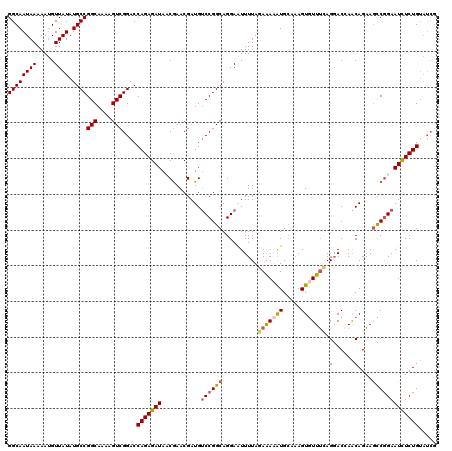
>X_DroMel_CAF1 19881806 120 - 22224390 GGCAAUAAAAAUGUUAUAUGCCGGCAAAAGUCGGACCAGAGAUACCGAACGAUGUCCGGCAGGAGUUUUAGAAAUAUGCAAAGUGUUUCAGGACGAACAGAAACCGGAAUCUCUGUAUCG ((((((((.....)))).))))(((....)))((..(((((((.(((.......(((....)))(((((.(((((((.....))))))))))))..........))).)))))))..)). ( -33.90) >DroSec_CAF1 35563 120 - 1 GGCAAUAAAAAUGUUAUAUGCCGGCAAAAGUCGGACCAGAGAUACCGAACGAUGUCCGGCAGGAAUUUUAGAAAUAUGCAAAGUGUUUCAGGACGAACAGAAGCCGGAAUCUCUGUAUCG ((((((((.....)))).))))(((....)))((..(((((((..(....)...((((((..........(((((((.....))))))).(......)....)))))))))))))..)). ( -32.00) >DroEre_CAF1 12900 114 - 1 GGCAAUAAAAAGGUUAUAUGCCGGCAAAAGUCGGACCAGAGAUAACGAACGAUGGCCGGCAGGAAUUUUAGAAAAGUGA--AGUGUUUUAGGACCAGCAGAAGCCGGAAUUUCUGG---- ...........((((...(((((((....((((................)))).)))))))...((((((......)))--))).......))))..((((((......)))))).---- ( -27.79) >DroYak_CAF1 11427 120 - 1 GGCAAUAAAAAUGUUAUAUGCCGGCAAAAGUCGGACCAGAGAUAACAAACGAUGUCCGGCAGGAAUUUUAGAGAAAUGCCACGUCUUAAAGCACCAACAGAAGUCAGAAUCUCUGUAUCG .((........((((((.(((((((....)))))..))...))))))...(((((..((((...............))))))))).....))....(((((..(....)..))))).... ( -25.86) >consensus GGCAAUAAAAAUGUUAUAUGCCGGCAAAAGUCGGACCAGAGAUAACGAACGAUGUCCGGCAGGAAUUUUAGAAAAAUGCAAAGUGUUUCAGGACCAACAGAAGCCGGAAUCUCUGUAUCG ((((((((.....)))).))))(((....)))....(((((((...........((((((..........(((((((.....))))))).(......)....)))))))))))))..... (-20.90 = -21.78 + 0.88)

| Location | 19,881,846 – 19,881,964 |
|---|---|
| Length | 118 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 89.22 |
| Mean single sequence MFE | -26.07 |
| Consensus MFE | -19.59 |
| Energy contribution | -19.90 |
| Covariance contribution | 0.31 |
| Combinations/Pair | 1.04 |
| Mean z-score | -1.53 |
| Structure conservation index | 0.75 |
| SVM decision value | 0.07 |
| SVM RNA-class probability | 0.569450 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
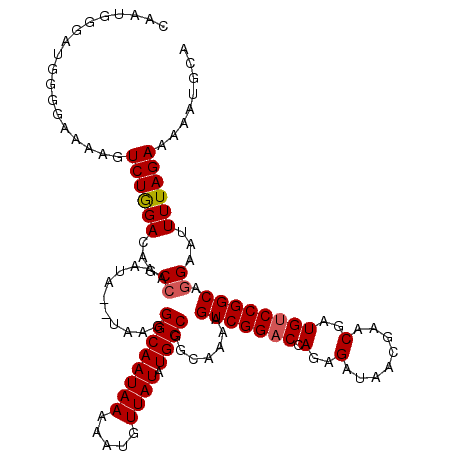
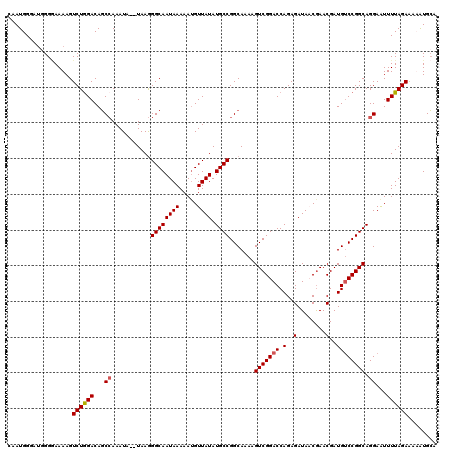
>X_DroMel_CAF1 19881846 118 - 22224390 CUAUGGUAUGGGGAAAAGUCUGGACAGCCAAAUA--UAAGGGCAAUAAAAAUGUUAUAUGCCGGCAAAAGUCGGACCAGAGAUACCGAACGAUGUCCGGCAGGAGUUUUAGAAAUAUGCA .....(((((...((((.(((.....(((.....--....)))...............((((((((....((((..........))))....)).))))))))).)))).....))))). ( -28.80) >DroSec_CAF1 35603 116 - 1 CAAUGG----GGGAAAAGUCUAGACAGCCAAAUAUAUAAGGGCAAUAAAAAUGUUAUAUGCCGGCAAAAGUCGGACCAGAGAUACCGAACGAUGUCCGGCAGGAAUUUUAGAAAUAUGCA ......----........((((((...((...........((((.......))))...((((((((....((((..........))))....)).))))))))...))))))........ ( -24.20) >DroEre_CAF1 12935 117 - 1 AAAUGGGAUGGGAAAAAGUCUGGACAUCGAAAUA--UAAAGGCAAUAAAAAGGUUAUAUGCCGGCAAAAGUCGGACCAGAGAUAACGAACGAUGGCCGGCAGGAAUUUUAGAAAAGUGA- ......((((..............))))......--.............((((((...(((((((....((((................)))).)))))))..))))))..........- ( -23.83) >DroYak_CAF1 11467 118 - 1 CAAUGGGAUGGGUAAAAGUCUGGACAGCCAAAUA--UAAGGGCAAUAAAAAUGUUAUAUGCCGGCAAAAGUCGGACCAGAGAUAACAAACGAUGUCCGGCAGGAAUUUUAGAGAAAUGCC .....((((........))))((((((((.....--....)))........((((((.(((((((....)))))..))...)))))).....)))))((((...............)))) ( -27.46) >consensus CAAUGGGAUGGGGAAAAGUCUGGACAGCCAAAUA__UAAGGGCAAUAAAAAUGUUAUAUGCCGGCAAAAGUCGGACCAGAGAUAACGAACGAUGUCCGGCAGGAAUUUUAGAAAAAUGCA ..................((((((...((...........((((((((.....)))).)))).......(((((((.(..(........)..)))))))).))...))))))........ (-19.59 = -19.90 + 0.31)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:52:27 2006