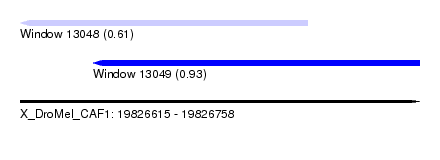

| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 19,826,615 – 19,826,758 |
| Length | 143 |
| Max. P | 0.925166 |

| Location | 19,826,615 – 19,826,718 |
|---|---|
| Length | 103 |
| Sequences | 5 |
| Columns | 106 |
| Reading direction | reverse |
| Mean pairwise identity | 86.91 |
| Mean single sequence MFE | -27.12 |
| Consensus MFE | -22.16 |
| Energy contribution | -22.68 |
| Covariance contribution | 0.52 |
| Combinations/Pair | 1.12 |
| Mean z-score | -1.51 |
| Structure conservation index | 0.82 |
| SVM decision value | 0.15 |
| SVM RNA-class probability | 0.607461 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
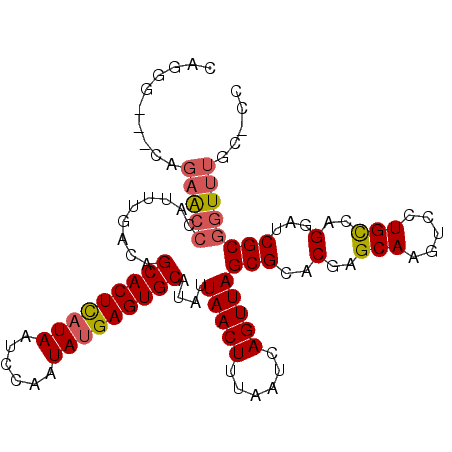
>X_DroMel_CAF1 19826615 103 - 22224390 CAGGG---AAGAACCCAUUUGACAGCACUUAUAAUCCAAUAUGAGUGCAUAUUAACUUUAAUCAGUUAGCGCACGAGCAAGUCCUGCCAGGAUCGCGGUUUGCCCC ..(((---..(((((.........(((((((((......)))))))))....(((((......)))))((((....))..((((.....)))).)))))))..))) ( -29.90) >DroSec_CAF1 32363 100 - 1 CAGGG---CAGAACCCAUUUGACAGCACUCAUAAUCCAAUAUGAGUGCAUAUUAACUUUAAUCAGUUAGCGCACGAGCAAGUCCUGUCAGGAUCGCGGUUUGC--- ....(---((((.(((.((((((((((((((((......)))))))))....(((((......)))))((......))......)))))))...).)))))))--- ( -27.80) >DroSim_CAF1 30526 100 - 1 CAGGG---CAGAACCCAUUUGACAGCACUCAUAAUCCAAUAUGAGUGCAUAUUAACUUUAAUCAGUUAGCGCACGAGCAAGUCCUGCCAGGAUCGCGGUUUGC--- ....(---((((.((.........(((((((((......)))))))))....(((((......)))))((((....))..((((.....)))).)))))))))--- ( -27.50) >DroEre_CAF1 22644 91 - 1 ------------GA-CACU--CCAGCACUCUUAAUGCAAUAUGAGUGCAUAUUAACUUUAAUCAGUUAGCGCACGAGCAAGUCCUGCCAGGAUCGCGGUUUCUUCC ------------((-.(((--...((((((............))))))....(((((......)))))((((....))..((((.....)))).))))).)).... ( -19.30) >DroYak_CAF1 23559 106 - 1 CGGGGGAACAGAACCCAUUUGACAGCACUCAUAAUCCAAUAUGAGUGCAUAUUAACUUUAAUCAGUUAGCGCACGAGCAAGUCCUGCCAGGAACGCGGUUUUUGCC (((((........)))........(((((((((......)))))))))...((((((......))))))....)).(((((..((((.......))))..))))). ( -31.10) >consensus CAGGG___CAGAACCCAUUUGACAGCACUCAUAAUCCAAUAUGAGUGCAUAUUAACUUUAAUCAGUUAGCGCACGAGCAAGUCCUGCCAGGAUCGCGGUUUGC_CC ..........(((((.........(((((((((......)))))))))....(((((......)))))(((..(..(((.....)))..)...))))))))..... (-22.16 = -22.68 + 0.52)


| Location | 19,826,641 – 19,826,758 |
|---|---|
| Length | 117 |
| Sequences | 3 |
| Columns | 117 |
| Reading direction | reverse |
| Mean pairwise identity | 97.72 |
| Mean single sequence MFE | -30.60 |
| Consensus MFE | -30.39 |
| Energy contribution | -30.50 |
| Covariance contribution | 0.11 |
| Combinations/Pair | 1.03 |
| Mean z-score | -2.06 |
| Structure conservation index | 0.99 |
| SVM decision value | 1.17 |
| SVM RNA-class probability | 0.925166 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
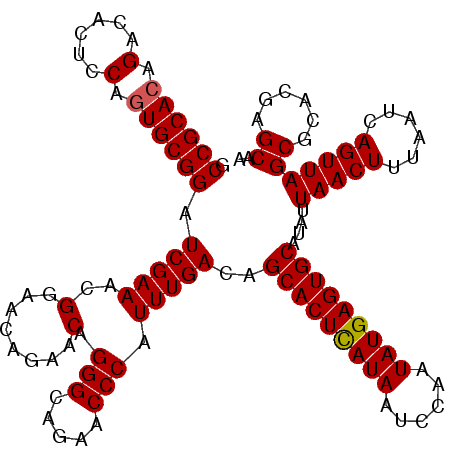
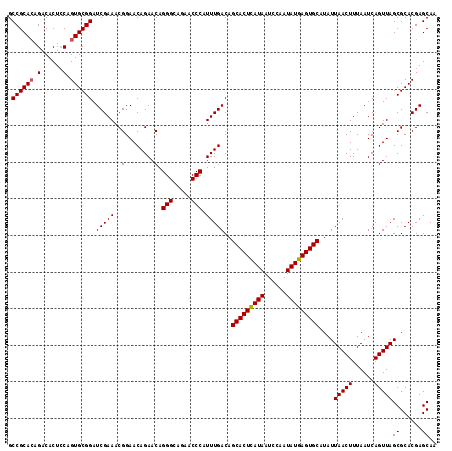
>X_DroMel_CAF1 19826641 117 - 22224390 GCCGCAAAGACACUACAGUGCGGAUCGAAACGGAACAGAACAGGGAAGAACCCAUUUGACAGCACUUAUAAUCCAAUAUGAGUGCAUAUUAACUUUAAUCAGUUAGCGCACGAGCAA .(((((..(......)..))))).(((((..(........).(((.....))).)))))..(((((((((......)))))))))....(((((......)))))((......)).. ( -27.60) >DroSec_CAF1 32386 117 - 1 GCCGCACAGACACUCCAGUGCGGAUCGAAACGGAACAGAACAGGGCAGAACCCAUUUGACAGCACUCAUAAUCCAAUAUGAGUGCAUAUUAACUUUAAUCAGUUAGCGCACGAGCAA .((((((..........)))))).(((((..(........).(((.....))).)))))..(((((((((......)))))))))....(((((......)))))((......)).. ( -32.10) >DroSim_CAF1 30549 117 - 1 GCCGCACAGACACUCCAGUGCGGAUCGAAACGGAACAGAACAGGGCAGAACCCAUUUGACAGCACUCAUAAUCCAAUAUGAGUGCAUAUUAACUUUAAUCAGUUAGCGCACGAGCAA .((((((..........)))))).(((((..(........).(((.....))).)))))..(((((((((......)))))))))....(((((......)))))((......)).. ( -32.10) >consensus GCCGCACAGACACUCCAGUGCGGAUCGAAACGGAACAGAACAGGGCAGAACCCAUUUGACAGCACUCAUAAUCCAAUAUGAGUGCAUAUUAACUUUAAUCAGUUAGCGCACGAGCAA .((((((.(......).)))))).(((((..(........).(((.....))).)))))..(((((((((......)))))))))....(((((......)))))((......)).. (-30.39 = -30.50 + 0.11)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:51:37 2006