
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 19,826,060 – 19,826,232 |
| Length | 172 |
| Max. P | 0.994588 |

| Location | 19,826,060 – 19,826,152 |
|---|---|
| Length | 92 |
| Sequences | 4 |
| Columns | 92 |
| Reading direction | reverse |
| Mean pairwise identity | 88.41 |
| Mean single sequence MFE | -32.55 |
| Consensus MFE | -27.36 |
| Energy contribution | -27.80 |
| Covariance contribution | 0.44 |
| Combinations/Pair | 1.13 |
| Mean z-score | -1.91 |
| Structure conservation index | 0.84 |
| SVM decision value | 0.94 |
| SVM RNA-class probability | 0.886085 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
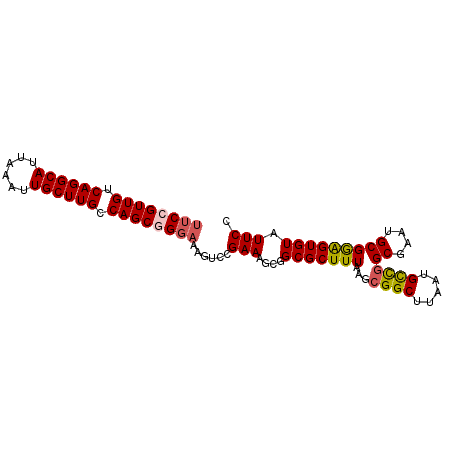
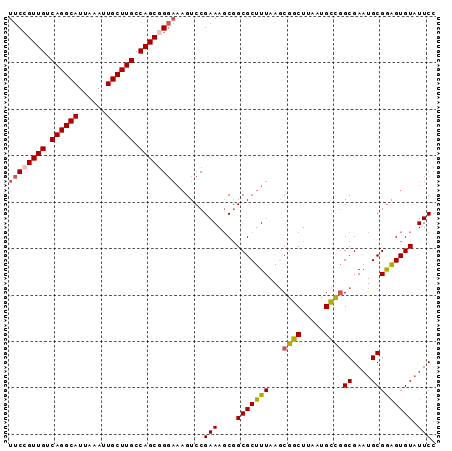
>X_DroMel_CAF1 19826060 92 - 22224390 UUCCGUUGUCAGGCAUUAAAUUGCUUGCCAGCGGGAAAGUCCGAAAUCGGCGCUUUAAGCGGCUUAAUGCUGGCGAAUGCGAAGUGUAUUCC .(((((((.((((((......)))))).)))))))((((((((....))).)))))...((((.....))))..((((((.....)))))). ( -34.70) >DroSec_CAF1 31850 92 - 1 UUCGGUUGUCAGGCAUUAUAUUGCUUGCCAGCGGGAAAGUCCGAAAGCGGCGCUUUAAGCGGCUUAAUGCCGGCGAAUGCGGAGUGUAUUCC ....((((.((((((......)))))).))))((((.(.((((....((.(((.....))(((.....)))).))....)))).)...)))) ( -32.80) >DroSim_CAF1 30010 92 - 1 UUCCGUUGUCAGGCAUUAAAUUGCUUGCCAGCGGGAAAGUCCGAAAGCAGCGCUUUAAGCGGCUUAAUGCCGGCGACUGCGGAGUGUAUUCC ((((((((.((((((......)))))).))))))))..........(((((((......((((.....))))))).))))((((....)))) ( -36.90) >DroEre_CAF1 22110 87 - 1 UGCCGUUGUCAGGCAUUAAAUUGCUUGCCAGCCG----GUCCGAAAGCGGCGCUUUAAACGGCUCUAUGUUUGCGAAUGCGGGGUGU-UUCA .(((((((.((((((......)))))).).((((----...(....)))))......))))))......(((((....)))))....-.... ( -25.80) >consensus UUCCGUUGUCAGGCAUUAAAUUGCUUGCCAGCGGGAAAGUCCGAAAGCGGCGCUUUAAGCGGCUUAAUGCCGGCGAAUGCGGAGUGUAUUCC ((((((((.((((((......)))))).))))))))......(((....(((((((...((((.....))))((....))))))))).))). (-27.36 = -27.80 + 0.44)

| Location | 19,826,119 – 19,826,232 |
|---|---|
| Length | 113 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 91.88 |
| Mean single sequence MFE | -27.97 |
| Consensus MFE | -22.93 |
| Energy contribution | -23.61 |
| Covariance contribution | 0.68 |
| Combinations/Pair | 1.07 |
| Mean z-score | -2.55 |
| Structure conservation index | 0.82 |
| SVM decision value | 2.49 |
| SVM RNA-class probability | 0.994588 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
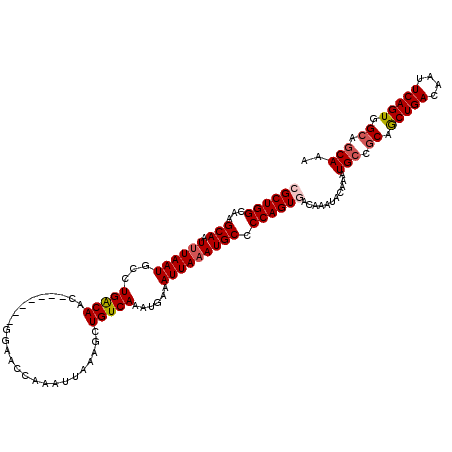
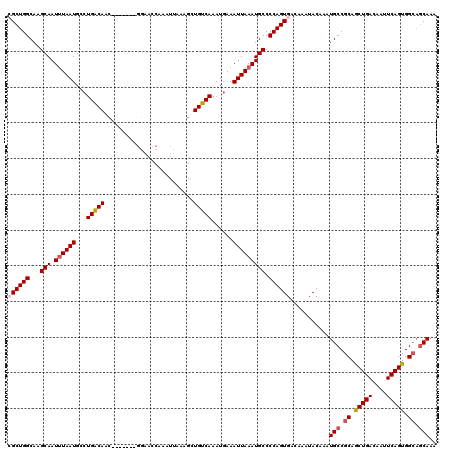
>X_DroMel_CAF1 19826119 113 + 22224390 CGCUGGCAAGCAAUUUAAUGCCUGACAAC-------GGAACCAAAUUAAAGAUGUCAAAUGAAAUUAAAUGCCCCAGUGACAAAUACAAAUGCCGCAGCUGACAAUUCAGUGGCAGCAAA ((((((...(((.((((((...(((((.(-------..............).)))))......))))))))).))))))...........(((.((.(((((....))))).)).))).. ( -28.24) >DroSec_CAF1 31909 110 + 1 CGCUGGCAAGCAAUAUAAUGCCUGACAAC-------CGAACCAAAUUAAAGCUGUCAAAUGAAAUUAAAUGCCCCAGUGACAAAUACAAAUGCCGCAGCUGACAAUUCAGUGG---CAAA ((((((...(((...((((...(((((..-------................)))))......))))..))).))))))...........((((...(((((....)))))))---)).. ( -22.27) >DroSim_CAF1 30069 113 + 1 CGCUGGCAAGCAAUUUAAUGCCUGACAAC-------GGAACCAAAUUAAAGCUGUCAAAUGAAAUUAAAUGCCCCAGUGACAAAUACAAAUGCCGCAGCUGACAAUUCAGUGGCAGCAAA ((((((...(((.((((((...(((((.(-------..............).)))))......))))))))).))))))...........(((.((.(((((....))))).)).))).. ( -28.24) >DroEre_CAF1 22164 113 + 1 GGCUGGCAAGCAAUUUAAUGCCUGACAAC-------GGCACCAAAUUAAAGCUGUCAAAUUAAAUUAAAUGCCCCAGUGACAAAUACAAAUGCCGCAACUGACAACUCAGUGGCAGCAAA .(((((...(((.((((((........((-------(((...........)))))........))))))))).)))))............(((.((.(((((....))))).)).))).. ( -27.89) >DroYak_CAF1 23064 120 + 1 GGCUGGCAGGCAAUUUAAUGGCUGGCAAGCAAGCCUGCAACCAAAUUAAAGCUGUCAAAUGAAAUUAAAUGCCCCAGUGACAAAUACAAAUGCCGCAGCUGACAAUUCAGUGGCAGCAAA .(((((..((((.((((((.((.(((......))).))................((....)).)))))))))))))))............(((.((.(((((....))))).)).))).. ( -33.20) >consensus CGCUGGCAAGCAAUUUAAUGCCUGACAAC_______GGAACCAAAUUAAAGCUGUCAAAUGAAAUUAAAUGCCCCAGUGACAAAUACAAAUGCCGCAGCUGACAAUUCAGUGGCAGCAAA ((((((...(((.((((((...(((((.........................)))))......))))))))).))))))...........(((.((.(((((....))))).)).))).. (-22.93 = -23.61 + 0.68)

| Location | 19,826,119 – 19,826,232 |
|---|---|
| Length | 113 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 91.88 |
| Mean single sequence MFE | -32.73 |
| Consensus MFE | -25.48 |
| Energy contribution | -25.64 |
| Covariance contribution | 0.16 |
| Combinations/Pair | 1.10 |
| Mean z-score | -2.16 |
| Structure conservation index | 0.78 |
| SVM decision value | 1.17 |
| SVM RNA-class probability | 0.925235 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 19826119 113 - 22224390 UUUGCUGCCACUGAAUUGUCAGCUGCGGCAUUUGUAUUUGUCACUGGGGCAUUUAAUUUCAUUUGACAUCUUUAAUUUGGUUCC-------GUUGUCAGGCAUUAAAUUGCUUGCCAGCG ..((((((..((((....))))..))))))........(((((.((..(.......)..))..)))))...............(-------((((.((((((......)))))).))))) ( -31.50) >DroSec_CAF1 31909 110 - 1 UUUG---CCACUGAAUUGUCAGCUGCGGCAUUUGUAUUUGUCACUGGGGCAUUUAAUUUCAUUUGACAGCUUUAAUUUGGUUCG-------GUUGUCAGGCAUUAUAUUGCUUGCCAGCG ....---..((..(((((..((((((((......((..((((.....))))..)).......))).))))).)))))..))...-------((((.((((((......)))))).)))). ( -29.02) >DroSim_CAF1 30069 113 - 1 UUUGCUGCCACUGAAUUGUCAGCUGCGGCAUUUGUAUUUGUCACUGGGGCAUUUAAUUUCAUUUGACAGCUUUAAUUUGGUUCC-------GUUGUCAGGCAUUAAAUUGCUUGCCAGCG ..((((((..((((....))))..)))))).......((((((.((..(.......)..))..))))))..............(-------((((.((((((......)))))).))))) ( -32.00) >DroEre_CAF1 22164 113 - 1 UUUGCUGCCACUGAGUUGUCAGUUGCGGCAUUUGUAUUUGUCACUGGGGCAUUUAAUUUAAUUUGACAGCUUUAAUUUGGUGCC-------GUUGUCAGGCAUUAAAUUGCUUGCCAGCC ..((((((.(((((....))))).)))))).................((((..(((((((((((((((((..............-------))))))))..)))))))))..)))).... ( -35.44) >DroYak_CAF1 23064 120 - 1 UUUGCUGCCACUGAAUUGUCAGCUGCGGCAUUUGUAUUUGUCACUGGGGCAUUUAAUUUCAUUUGACAGCUUUAAUUUGGUUGCAGGCUUGCUUGCCAGCCAUUAAAUUGCCUGCCAGCC ..((((((..((((....))))..)))))).............((((((((((((((......((.(((((.......)))))))((((.(....).)))))))))).))))..)))).. ( -35.70) >consensus UUUGCUGCCACUGAAUUGUCAGCUGCGGCAUUUGUAUUUGUCACUGGGGCAUUUAAUUUCAUUUGACAGCUUUAAUUUGGUUCC_______GUUGUCAGGCAUUAAAUUGCUUGCCAGCG .........((..(((((..((((((((......((..((((.....))))..)).......))).))))).)))))..))..........((((.((((((......)))))).)))). (-25.48 = -25.64 + 0.16)

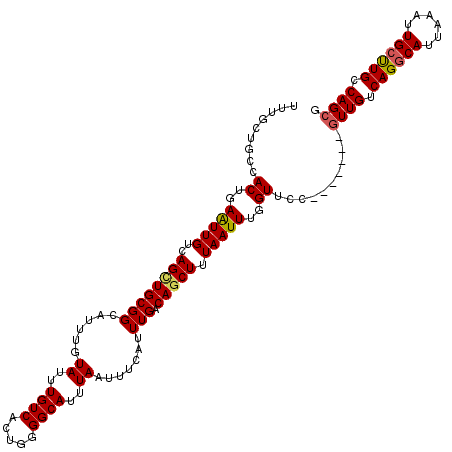
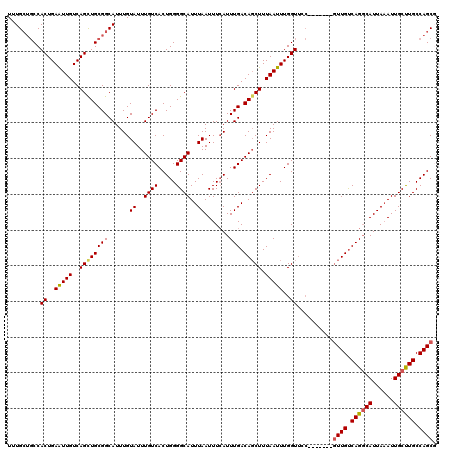
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:51:36 2006