
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 19,811,631 – 19,811,907 |
| Length | 276 |
| Max. P | 0.985428 |

| Location | 19,811,631 – 19,811,751 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 89.56 |
| Mean single sequence MFE | -33.41 |
| Consensus MFE | -27.96 |
| Energy contribution | -30.12 |
| Covariance contribution | 2.16 |
| Combinations/Pair | 1.09 |
| Mean z-score | -2.17 |
| Structure conservation index | 0.84 |
| SVM decision value | 2.01 |
| SVM RNA-class probability | 0.985428 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 19811631 120 - 22224390 ACCUUGACACAAAUUAACGCAGAAUUGGCCGCCCCUUAAUCUCAUUUGCCGCCCAAUGAAAAUGCACUUCCUUCAGAAGUGUGUUGCUAGUGGCCAAUGGAGUUAAACUGCCCAAGGACA .(((((...((..(((((.(...(((((((((.........(((((........))))).((((((((((.....))))))))))....))))))))).).)))))..))..)))))... ( -37.30) >DroSec_CAF1 17505 118 - 1 ACCUUGACACAAAUUAACGCAGAAUCGGCCGCCCCUUAAUCUCAUUUUCCGCCCAAUGGAAAUGCACUUCCUUCAGAAGUGUG--ACUAGUGGCCAAUGGAGUUGAACUGCCCAAGGACA .(((((...((..(((((.(...((.((((((.............((((((.....)))))).(((((((.....))))))).--....)))))).)).).)))))..))..)))))... ( -33.50) >DroSim_CAF1 14553 118 - 1 ACCUUGACACAAAUUAACGCAGAAUUGGCCGCCCCUUAAUCUCAUUUUCCGCCCAAUGGAAAUGCACUUCCUUCAGAAGUGUG--ACUAGUGGCCAAUGGAGUUGAACUGGCCAAGGACA .(((((.(.((..(((((.(...(((((((((.............((((((.....)))))).(((((((.....))))))).--....))))))))).).)))))..))).)))))... ( -39.00) >DroEre_CAF1 9835 110 - 1 ACCUUGACACAAAUUAACACAGAAUUGGCCGCCCUUUAAUCUUAUUUGCCGCCCCAUGGAAAUGCAUUUCCUUUAGAAGUGUG--GCUAGUGG-------AG-AAAAUUGGCCAAGGACA .(((((.(...((((..(.(...(((((((((.((((.........(((..((....))....))).........)))).)))--)))))).)-------.)-..)))).).)))))... ( -25.37) >DroYak_CAF1 10072 118 - 1 ACCUUGACACAAAUUAACGCAGAAUUGGCCGCCCUUUAAUCUCAUUUUCCGCCCAAUGGAAAUGCAUUUCCUUCAGCAGUGCG--GCCAGUGGCCAAUGGAGUAAAAUUGCCCAAGGACA .(((((.................(((((((((.(...........((((((.....))))))(((..........)))).)))--))))))(((.(((........)))))))))))... ( -31.90) >consensus ACCUUGACACAAAUUAACGCAGAAUUGGCCGCCCCUUAAUCUCAUUUUCCGCCCAAUGGAAAUGCACUUCCUUCAGAAGUGUG__GCUAGUGGCCAAUGGAGUUAAACUGCCCAAGGACA .(((((...((..(((((.(...(((((((((.............((((((.....)))))).(((((((.....))))))).......))))))))).).)))))..))..)))))... (-27.96 = -30.12 + 2.16)



| Location | 19,811,751 – 19,811,867 |
|---|---|
| Length | 116 |
| Sequences | 4 |
| Columns | 117 |
| Reading direction | reverse |
| Mean pairwise identity | 80.58 |
| Mean single sequence MFE | -38.15 |
| Consensus MFE | -22.30 |
| Energy contribution | -22.24 |
| Covariance contribution | -0.06 |
| Combinations/Pair | 1.11 |
| Mean z-score | -2.07 |
| Structure conservation index | 0.58 |
| SVM decision value | 0.39 |
| SVM RNA-class probability | 0.715295 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 19811751 116 - 22224390 AGCAUGAGGAUGAAAAAAUUUCCCACAACUUCAUGCCGCGAAACGCUUUGCGGG-UGGAUGGUUGCGGGGAGGGGGGGAAGGAGGAUGGAAACGCCUAUCGCUGGUCAACGGAUGAA ..(((..(..(((.....((((((.(((((((((.(((((((....))))))))-)))..))))).))))))..((.((...(((.((....))))).)).))..))).)..))).. ( -36.60) >DroSec_CAF1 17623 117 - 1 AGCAUGAGGCAGAAAAAAUUUCCCACAACUUCAAGCCGCGAAACGCUUUGCGGGAUGGGUGGUUGGCUGGUGGGGGGGUGGGUUUAUGGAAACGCCUAUCGCUGGUCAACGGAUGAA .((.....))..........((((((....(((..(((((((....)))))))..))))))((((((..((((..(((((..((....))..))))).))))..))))))))).... ( -40.10) >DroSim_CAF1 14671 103 - 1 AGCAUGAGGCAGAAAAAAUUUCCCACAACUUCAAACCGCGAAACGCUUCGCGGGAUGGGUGGU----------GGGGGUGG----AUGGAAACGCCUAUCGCUGGUCAACGGAUGAA (((...((((.........(((((((.(((.((..(((((((....)))))))..))))).))----------)))))...----..(....)))))...))).(((....)))... ( -37.80) >DroEre_CAF1 9945 101 - 1 AGCAUAAGGCAGAAA-AAUGCCCCGCAACUUCAAGCCGCGAAUCGCUUUGGGG-UUGGGUGUU----------GGGGGG----AAAUGGAAACGCCUAUCGCUGCUCAGCGGAUGAA ..(((.((((.....-....((((.((((.((.((((.((((....)))).))-)).)).)))----------).))))----....(....))))).(((((....)))))))).. ( -38.10) >consensus AGCAUGAGGCAGAAAAAAUUUCCCACAACUUCAAGCCGCGAAACGCUUUGCGGGAUGGGUGGU__________GGGGGUGG____AUGGAAACGCCUAUCGCUGGUCAACGGAUGAA ......((((..........((((.((..((((..(((((((....)))))))..)))))).............)))).........(....))))).((((((.....))).))). (-22.30 = -22.24 + -0.06)

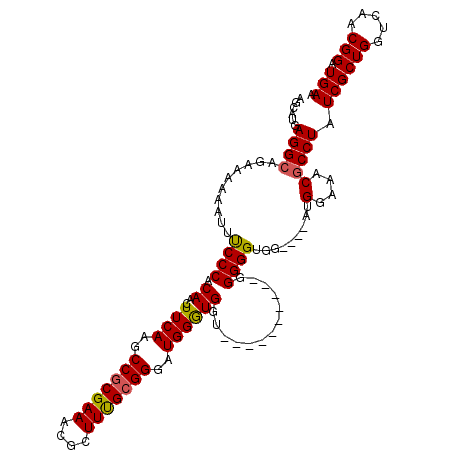
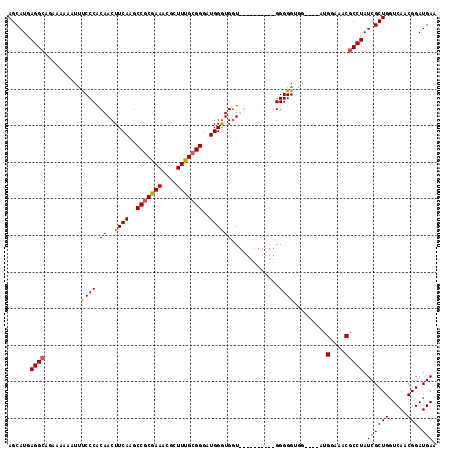
| Location | 19,811,791 – 19,811,907 |
|---|---|
| Length | 116 |
| Sequences | 4 |
| Columns | 117 |
| Reading direction | forward |
| Mean pairwise identity | 76.07 |
| Mean single sequence MFE | -28.83 |
| Consensus MFE | -19.14 |
| Energy contribution | -19.08 |
| Covariance contribution | -0.06 |
| Combinations/Pair | 1.18 |
| Mean z-score | -1.65 |
| Structure conservation index | 0.66 |
| SVM decision value | 0.36 |
| SVM RNA-class probability | 0.702734 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 19811791 116 + 22224390 CCCCCCUCCCCGCAACCAUCCA-CCCGCAAAGCGUUUCGCGGCAUGAAGUUGUGGGAAAUUUUUUCAUCCUCAUGCUAGCUUUAACCCCCAAAACGGGGUCAUAAUGGGUAAUGGGU ...............((((..(-(((((...(((...)))(((((((.(..(((..(.....)..)))).))))))).))....(((((......)))))......)))).)))).. ( -33.70) >DroSec_CAF1 17663 103 + 1 CCCCCCACCAGCCAACCACCCAUCCCGCAAAGCGUUUCGCGGCUUGAAGUUGUGGGAAAUUUUUUCUGCCUCAUGCUCGCUUUAGCCCCA--AAAGGGGUCAUAA------------ ..((((......((((....((..((((..........))))..))..))))((((...........((.....))..((....))))))--...))))......------------ ( -23.00) >DroSim_CAF1 14707 93 + 1 CCCC----------ACCACCCAUCCCGCGAAGCGUUUCGCGGUUUGAAGUUGUGGGAAAUUUUUUCUGCCUCAUGCUCGCUUUAGCCCCA--AAAGGGGUCAUAA------------ .(((----------((.((.((..(((((((....)))))))..))..)).)))))............................(((((.--...))))).....------------ ( -30.60) >DroEre_CAF1 9981 90 + 1 CCCC----------AACACCCAA-CCCCAAAGCGAUUCGCGGCUUGAAGUUGCGGGGCAUU-UUUCUGCCUUAUGCUCGCUUUAACCCCA-A--AGGGGUCAUAA------------ ....----------.........-....(((((((..((((((.....))))))(((((..-....))))).....))))))).(((((.-.--.))))).....------------ ( -28.00) >consensus CCCC__________ACCACCCAUCCCGCAAAGCGUUUCGCGGCUUGAAGUUGUGGGAAAUUUUUUCUGCCUCAUGCUCGCUUUAACCCCA__AAAGGGGUCAUAA____________ ............................(((((((((((((((.....)))))))))..........((.....)).)))))).(((((......)))))................. (-19.14 = -19.08 + -0.06)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:51:19 2006