
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 19,600,909 – 19,601,021 |
| Length | 112 |
| Max. P | 0.564934 |

| Location | 19,600,909 – 19,601,021 |
|---|---|
| Length | 112 |
| Sequences | 4 |
| Columns | 117 |
| Reading direction | forward |
| Mean pairwise identity | 76.96 |
| Mean single sequence MFE | -21.23 |
| Consensus MFE | -10.41 |
| Energy contribution | -14.29 |
| Covariance contribution | 3.88 |
| Combinations/Pair | 1.04 |
| Mean z-score | -2.13 |
| Structure conservation index | 0.49 |
| SVM decision value | -0.05 |
| SVM RNA-class probability | 0.510144 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 19600909 112 + 22224390 CAAAUACCAAUGCACGCACA--CACAACAUACACGAAAUAUCAUUGUAUACAAUGGACAAUGAUUUCGAUAUUUUUA--AACCAUUAUCUGUGUGCUUUUUUUGUUUUUUGGUG-GA ....((((((.(((.(((((--((..........(((..((((((((.........)))))))))))((((......--......)))))))))))......)))...))))))-.. ( -24.50) >DroSec_CAF1 7246 113 + 1 CAAAUACCAAUGCACGCACACACACAACAUAUACCCAAAAUCAUUGUUUGCAAUCGACAAUGAUUUCGAUAUUUUUA--AACCAUUAUCUGUGUGCUUUUCCUAUU-UUUGGUG-GA ....((((((.(((((((...................(((((((((((.......))))))))))).((((......--......)))))))))))..........-.))))))-.. ( -24.10) >DroSim_CAF1 7173 113 + 1 CAAAUACCAAUGCACGCACACACACAACAUAUACGAAAAAUCAUUGUUUGCAGUCGACAAUGAUUUCGAUAUUUUUA--AACCAUUAUCUGUGUGCUUUUCCUAUU-UUUGGUG-GA ....((((((.(((((((...................(((((((((((.......))))))))))).((((......--......)))))))))))..........-.))))))-.. ( -24.10) >DroAna_CAF1 8983 105 + 1 CAAAUACACAUGCACGCACACUCAG-ACAUACACAAUACACAAUUAAUUA-AUUG------GAUUAAAACAUUUUCAUUAAAUAUUC-UUGUGUGAUAUUCG--AU-UUUGGUGCCA ...........((((.((.(.((..-(..(((((((....(((((....)-))))------((...........))...........-)))))))...)..)--).-).)))))).. ( -12.20) >consensus CAAAUACCAAUGCACGCACACACACAACAUACACGAAAAAUCAUUGUUUACAAUCGACAAUGAUUUCGAUAUUUUUA__AACCAUUAUCUGUGUGCUUUUCCUAUU_UUUGGUG_GA ....((((((.(((((((...................(((((((((((.......))))))))))).((((..............)))))))))))............))))))... (-10.41 = -14.29 + 3.88)

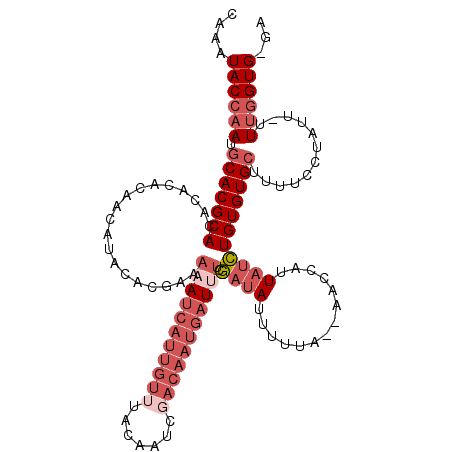
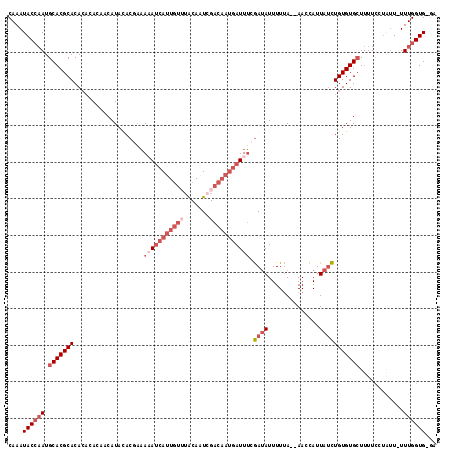
| Location | 19,600,909 – 19,601,021 |
|---|---|
| Length | 112 |
| Sequences | 4 |
| Columns | 117 |
| Reading direction | reverse |
| Mean pairwise identity | 76.96 |
| Mean single sequence MFE | -26.93 |
| Consensus MFE | -12.88 |
| Energy contribution | -15.38 |
| Covariance contribution | 2.50 |
| Combinations/Pair | 1.17 |
| Mean z-score | -2.24 |
| Structure conservation index | 0.48 |
| SVM decision value | 0.06 |
| SVM RNA-class probability | 0.564934 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 19600909 112 - 22224390 UC-CACCAAAAAACAAAAAAAGCACACAGAUAAUGGUU--UAAAAAUAUCGAAAUCAUUGUCCAUUGUAUACAAUGAUAUUUCGUGUAUGUUGUG--UGUGCGUGCAUUGGUAUUUG ..-.(((((...((.......(((((((((((((((((--(..........))))))))))).....((((((.(((....)))))))))...))--)))))))...)))))..... ( -27.91) >DroSec_CAF1 7246 113 - 1 UC-CACCAAA-AAUAGGAAAAGCACACAGAUAAUGGUU--UAAAAAUAUCGAAAUCAUUGUCGAUUGCAAACAAUGAUUUUGGGUAUAUGUUGUGUGUGUGCGUGCAUUGGUAUUUG ..-.(((((.-.....(....(((((((.((((((...--((......(((((((((((((.........))))))))))))).))..)))))).)))))))...).)))))..... ( -30.30) >DroSim_CAF1 7173 113 - 1 UC-CACCAAA-AAUAGGAAAAGCACACAGAUAAUGGUU--UAAAAAUAUCGAAAUCAUUGUCGACUGCAAACAAUGAUUUUUCGUAUAUGUUGUGUGUGUGCGUGCAUUGGUAUUUG ..-.(((((.-.....(....(((((((.((((((((.--.......)))(((((((((((.........)))))))))))........))))).)))))))...).)))))..... ( -27.70) >DroAna_CAF1 8983 105 - 1 UGGCACCAAA-AU--CGAAUAUCACACAA-GAAUAUUUAAUGAAAAUGUUUUAAUC------CAAU-UAAUUAAUUGUGUAUUGUGUAUGU-CUGAGUGUGCGUGCAUGUGUAUUUG ..((((((..-.(--((.((((.((((((-((((((((.....))))))))).(..------((((-(....)))))..)..)))))))))-.)))...)).))))........... ( -21.80) >consensus UC_CACCAAA_AAUAGGAAAAGCACACAGAUAAUGGUU__UAAAAAUAUCGAAAUCAUUGUCCAUUGCAAACAAUGAUUUUUCGUAUAUGUUGUGUGUGUGCGUGCAUUGGUAUUUG .....................(((((((.((((((((..........)))(((((((((((.........)))))))))))........))))).)))))))((((....))))... (-12.88 = -15.38 + 2.50)


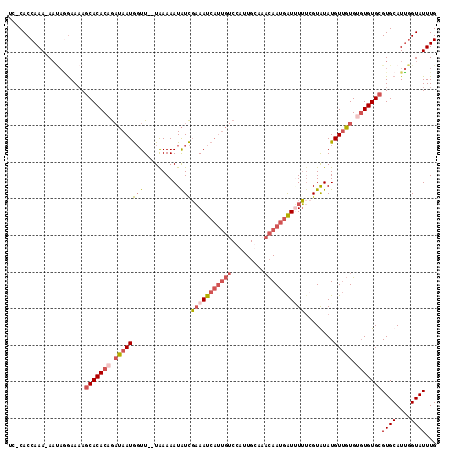
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:49:35 2006