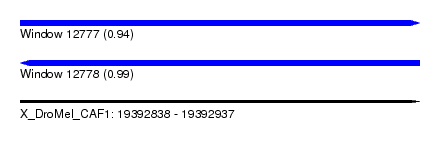
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 19,392,838 – 19,392,937 |
| Length | 99 |
| Max. P | 0.988840 |

| Location | 19,392,838 – 19,392,937 |
|---|---|
| Length | 99 |
| Sequences | 6 |
| Columns | 99 |
| Reading direction | forward |
| Mean pairwise identity | 75.32 |
| Mean single sequence MFE | -39.16 |
| Consensus MFE | -33.46 |
| Energy contribution | -32.52 |
| Covariance contribution | -0.94 |
| Combinations/Pair | 1.31 |
| Mean z-score | -4.37 |
| Structure conservation index | 0.85 |
| SVM decision value | 1.28 |
| SVM RNA-class probability | 0.939562 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 19392838 99 + 22224390 AUGCAGUGCCGCGGCAUUGGUGAGGCCAUCUCCAAUGGAUUAGUUCUCAACAUUGGAUUAGUUAUCAUCAAUGCCGGUGCACUGCACCUAACAACAUCG .(((((((((.((((((((((((.......(((((((.............))))))).......))))))))))))).))))))))............. ( -43.76) >DroVir_CAF1 41100 94 + 1 GUGCAGCCCCUCGCGAUUGAUGAUAGCAUCUCCAAUGAG-CAU---AUCACAUUGGAUUAGUUCUCAUCGAUGCCGAUGCGCUGUACCAG-CAACAACA (((((((.(.(((..((((((((.(((...(((((((..-...---....)))))))...))).))))))))..))).).)))))))...-........ ( -32.60) >DroSec_CAF1 25983 99 + 1 AUGCAGUGCCGUGGCAUUGGUGAGGACAUCUCCAAUGGCUUAGUUCUCAUCAUUGGAUUAGUUAUCAUCAAUGCCGGUGCACUGCAGCUGUCAACUUCG .((((((((..((((((((((((.(((...((((((((...........))))))))...))).))))))))))))..))))))))............. ( -43.70) >DroEre_CAF1 24534 98 + 1 AUGCAGUGCCUCGACAUUGGUGAGGGCAUCUCCAAUGGG-GAGCCAGCAUCAUUGGAUUAGUUCUCAUUAAUGCCGGAGCAUGGCACCUACGAACUGAG .(((.((((..((.(((((((((((((...(((((((..-(......)..)))))))...))))))))))))).))..)))).)))............. ( -38.90) >DroYak_CAF1 23882 98 + 1 AUGCAGUGCCACGACAUUGGUGAGGGCAUCUCCAAUGGG-GACCCGGCAUCAUUGGAUUAGUUCUCAUUAAUGCCGGAGCACUGCACCUACCAACUUCG .((((((((..((.(((((((((((((...(((((((..-(......)..)))))))...))))))))))))).))..))))))))............. ( -44.20) >DroMoj_CAF1 32398 94 + 1 GUGCAGCCCCUCGCGAUUGAUGAAAGCAUCUCCAAUGAG-UAU---UUCGCAUUGGAUUAGUUCUCAUCGAUGUCGAUGCGCUGUAUCAG-CAACAACA (((((((.(.(((..((((((((.(((...(((((((.(-...---..).)))))))...))).))))))))..))).).)))))))...-........ ( -31.80) >consensus AUGCAGUGCCUCGACAUUGGUGAGGGCAUCUCCAAUGGG_UAG___UCAUCAUUGGAUUAGUUCUCAUCAAUGCCGGUGCACUGCACCUA_CAACAUCG .((((((((..((.(((((((((.(((...(((((((.............)))))))...))).))))))))).))..))))))))............. (-33.46 = -32.52 + -0.94)



| Location | 19,392,838 – 19,392,937 |
|---|---|
| Length | 99 |
| Sequences | 6 |
| Columns | 99 |
| Reading direction | reverse |
| Mean pairwise identity | 75.32 |
| Mean single sequence MFE | -34.86 |
| Consensus MFE | -27.08 |
| Energy contribution | -29.28 |
| Covariance contribution | 2.20 |
| Combinations/Pair | 1.11 |
| Mean z-score | -2.96 |
| Structure conservation index | 0.78 |
| SVM decision value | 2.14 |
| SVM RNA-class probability | 0.988840 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 19392838 99 - 22224390 CGAUGUUGUUAGGUGCAGUGCACCGGCAUUGAUGAUAACUAAUCCAAUGUUGAGAACUAAUCCAUUGGAGAUGGCCUCACCAAUGCCGCGGCACUGCAU ............(((((((((..((((((((.(((...(((.(((((((..((.......)))))))))..)))..))).))))))))..))))))))) ( -42.30) >DroVir_CAF1 41100 94 - 1 UGUUGUUG-CUGGUACAGCGCAUCGGCAUCGAUGAGAACUAAUCCAAUGUGAU---AUG-CUCAUUGGAGAUGCUAUCAUCAAUCGCGAGGGGCUGCAC (((.(((.-((.((...((......))...(((((.......(((((((.(..---...-).))))))).......)))))....)).)).))).))). ( -26.34) >DroSec_CAF1 25983 99 - 1 CGAAGUUGACAGCUGCAGUGCACCGGCAUUGAUGAUAACUAAUCCAAUGAUGAGAACUAAGCCAUUGGAGAUGUCCUCACCAAUGCCACGGCACUGCAU .............((((((((...(((((((.(((..((...(((((((.............)))))))...))..))).)))))))...)))))))). ( -38.92) >DroEre_CAF1 24534 98 - 1 CUCAGUUCGUAGGUGCCAUGCUCCGGCAUUAAUGAGAACUAAUCCAAUGAUGCUGGCUC-CCCAUUGGAGAUGCCCUCACCAAUGUCGAGGCACUGCAU ........((((.((((.....(((((((((.((.((.....)))).))))))))).((-..((((((.((.....)).))))))..)))))))))).. ( -34.00) >DroYak_CAF1 23882 98 - 1 CGAAGUUGGUAGGUGCAGUGCUCCGGCAUUAAUGAGAACUAAUCCAAUGAUGCCGGGUC-CCCAUUGGAGAUGCCCUCACCAAUGUCGUGGCACUGCAU ............(((((((((..(((((((..((((..(...(((((((..........-..)))))))...)..))))..)))))))..))))))))) ( -39.80) >DroMoj_CAF1 32398 94 - 1 UGUUGUUG-CUGAUACAGCGCAUCGACAUCGAUGAGAACUAAUCCAAUGCGAA---AUA-CUCAUUGGAGAUGCUUUCAUCAAUCGCGAGGGGCUGCAC ........-......((((.(.(((.....(((((((.(...(((((((.(..---...-).)))))))...).))))))).....))).).))))... ( -27.80) >consensus CGAAGUUG_CAGGUGCAGUGCACCGGCAUUGAUGAGAACUAAUCCAAUGAUGA___CUA_CCCAUUGGAGAUGCCCUCACCAAUGCCGAGGCACUGCAU ............(((((((((..((((((((.(((.......(((((((.............))))))).......))).))))))))..))))))))) (-27.08 = -29.28 + 2.20)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:47:34 2006