
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 19,210,829 – 19,210,949 |
| Length | 120 |
| Max. P | 0.558782 |

| Location | 19,210,829 – 19,210,949 |
|---|---|
| Length | 120 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 80.99 |
| Mean single sequence MFE | -33.40 |
| Consensus MFE | -20.89 |
| Energy contribution | -24.82 |
| Covariance contribution | 3.94 |
| Combinations/Pair | 1.21 |
| Mean z-score | -1.68 |
| Structure conservation index | 0.63 |
| SVM decision value | -0.05 |
| SVM RNA-class probability | 0.508374 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 19210829 120 + 22224390 AUCCAAUAGUUGCAAUUGCAACCGCAUUGCAACAACGUGUGCAGCAGUGAAAACAGCAGAUCGCAAAUGAAUCUGGGUAUCUCGUGUUGGAUUAUUCGAGCCGUGUAUGUGUAUAUGUGU ........((((((((.((....)))))))))).(((((((((...((....))..(((((((....)).))))).((((..((.(((.((....)).))))).)))).))))))))).. ( -32.80) >DroSec_CAF1 32137 119 + 1 AUGCAAUAGUUGCAUUUGCAACCGCAUUGCAACAACUUGUGCAGCAGUGAAAACAGCAGAUCGAAAAUGGAUCUG-GUAGCUCGAGUUGUAUUAUUCGAGCCCUGUGUGUGUGCAUGUGU .((((((.(((((....)))))...))))))(((...((..((((((.(.......((((((.......))))))-...((((((((......)))))))))))))...))..))..))) ( -40.10) >DroEre_CAF1 29273 107 + 1 UU-CAAUAGUUGCCAUUUCAACCGCAUUGCAAUAGCAUGUGC---AGUGGAAACAGGGGAUCGAAAAUGAAUCUG-GGGGCUCGUGUUGGAUUAUUCGAGCAGGAG--------AUGUGU ..-........((((((((..((.(((((((........)))---))))....((((...((......)).))))-)).(((((.((......)).)))))..)))--------))).)) ( -26.70) >DroYak_CAF1 30032 107 + 1 AU-CAAUAGUUGCAUUUGCAACCGCAUUGCAAUAACAUGUGC---AGUGAAAACAGCAGAUCGAAAAUGGAUCUG-GCAGCUCGGGUUGGAUUAUUCGAGCAGGAG--------AUGUGU ..-.....(((((....)))))(((((..(.........(((---.((....))..((((((.......))))))-)))((((((((......))))))))..)..--------))))). ( -34.00) >consensus AU_CAAUAGUUGCAAUUGCAACCGCAUUGCAACAACAUGUGC___AGUGAAAACAGCAGAUCGAAAAUGAAUCUG_GUAGCUCGUGUUGGAUUAUUCGAGCAGGAG________AUGUGU ........((((((((.((....)))))))))).(((((((((.............((((((.......))))))....((((((((......))))))))........))))))))).. (-20.89 = -24.82 + 3.94)

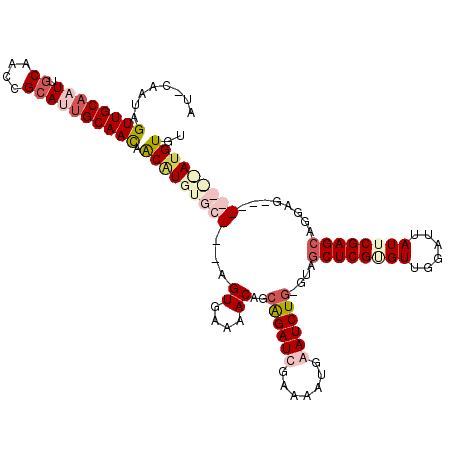

| Location | 19,210,829 – 19,210,949 |
|---|---|
| Length | 120 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 80.99 |
| Mean single sequence MFE | -26.77 |
| Consensus MFE | -16.19 |
| Energy contribution | -16.75 |
| Covariance contribution | 0.56 |
| Combinations/Pair | 1.09 |
| Mean z-score | -1.82 |
| Structure conservation index | 0.60 |
| SVM decision value | 0.05 |
| SVM RNA-class probability | 0.558782 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 19210829 120 - 22224390 ACACAUAUACACAUACACGGCUCGAAUAAUCCAACACGAGAUACCCAGAUUCAUUUGCGAUCUGCUGUUUUCACUGCUGCACACGUUGUUGCAAUGCGGUUGCAAUUGCAACUAUUGGAU ............................((((((...((((....((((((.......))))))....))))...............((((((((((....)).))))))))..)))))) ( -23.60) >DroSec_CAF1 32137 119 - 1 ACACAUGCACACACACAGGGCUCGAAUAAUACAACUCGAGCUAC-CAGAUCCAUUUUCGAUCUGCUGUUUUCACUGCUGCACAAGUUGUUGCAAUGCGGUUGCAAAUGCAACUAUUGCAU ......(((.....(((((((((((..........)))))))..-((((((.......))))))))))......)))((((..((((((((((((...))))))...))))))..)))). ( -34.70) >DroEre_CAF1 29273 107 - 1 ACACAU--------CUCCUGCUCGAAUAAUCCAACACGAGCCCC-CAGAUUCAUUUUCGAUCCCCUGUUUCCACU---GCACAUGCUAUUGCAAUGCGGUUGAAAUGGCAACUAUUG-AA ....((--------((...(((((............)))))...-.)))).....((((((...(((((((.(((---(((..(((....))).)))))).))))))).....))))-)) ( -25.40) >DroYak_CAF1 30032 107 - 1 ACACAU--------CUCCUGCUCGAAUAAUCCAACCCGAGCUGC-CAGAUCCAUUUUCGAUCUGCUGUUUUCACU---GCACAUGUUAUUGCAAUGCGGUUGCAAAUGCAACUAUUG-AU ......--------.....(((((............))))).((-((((((.......))))))..........(---(((........))))..))((((((....))))))....-.. ( -23.40) >consensus ACACAU________ACACGGCUCGAAUAAUCCAACACGAGCUAC_CAGAUCCAUUUUCGAUCUGCUGUUUUCACU___GCACAUGUUAUUGCAAUGCGGUUGCAAAUGCAACUAUUG_AU ...................(((((............)))))....((((((.......))))))..............(((...((....))..)))((((((....))))))....... (-16.19 = -16.75 + 0.56)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:45:38 2006