
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 19,208,589 – 19,208,722 |
| Length | 133 |
| Max. P | 0.997069 |

| Location | 19,208,589 – 19,208,684 |
|---|---|
| Length | 95 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 77.07 |
| Mean single sequence MFE | -19.52 |
| Consensus MFE | -16.67 |
| Energy contribution | -16.91 |
| Covariance contribution | 0.25 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.06 |
| Structure conservation index | 0.85 |
| SVM decision value | 0.03 |
| SVM RNA-class probability | 0.547048 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 19208589 95 - 22224390 AAGCGCCCUUUGUGCCGAAAUCAAACUGCAAAUACAUGGCCUAAAAGCAUUGGCAAACAAGCAAACAUGGCCAUGCAUC------CUCACUCCUUUAACGC------------------- ..((((..((((.........))))..)).....(((((((.....((.(((.....)))))......)))))))....------..............))------------------- ( -18.90) >DroSec_CAF1 29950 106 - 1 AAGCGCCCUUUGUGCCGAAAUCAAACUGCAAAUACAUGGCCUAAAAGCAUUGGCAAACAAGCAAACAUGGCCAUGC---------CACACCCCCUU----CCCGCC-CGCCCUUCCUUCC ..((((..((((.........))))..)).....(((((((.....((.(((.....)))))......))))))).---------...........----...)).-............. ( -19.70) >DroSim_CAF1 5216 110 - 1 AAGCGCCCUUUGUGCCGAAAUCAAACUGCAAAUACAUGGCCUAAAAGCAUUGGCAAACAAGCAAACAUGGCCAUGC---------CACACCCCCUUACCGCCCGCC-CGCCCUUCAUUCC ..((((.....)))).(((...............(((((((.....((.(((.....)))))......))))))).---------..............((.....-.))..)))..... ( -19.90) >DroYak_CAF1 27720 108 - 1 AAGCGCCCUUUGUGCCGGAAUCAAACUGCAAAUACAUGGCCUAAAAGCAUUGGCAAACA----AACUCGGCCAUGUACUCUACUCCCCGCCCCCUUUCC--CUGCCACGCCCCU------ ..(((.......(((.(........).)))..(((((((((....((..(((.....))----).)).)))))))))......................--......)))....------ ( -19.60) >consensus AAGCGCCCUUUGUGCCGAAAUCAAACUGCAAAUACAUGGCCUAAAAGCAUUGGCAAACAAGCAAACAUGGCCAUGC_________CACACCCCCUUACC_CCCGCC_CGCCCUU______ ..((((.....))))...................(((((((........(((.....)))........)))))))............................................. (-16.67 = -16.91 + 0.25)

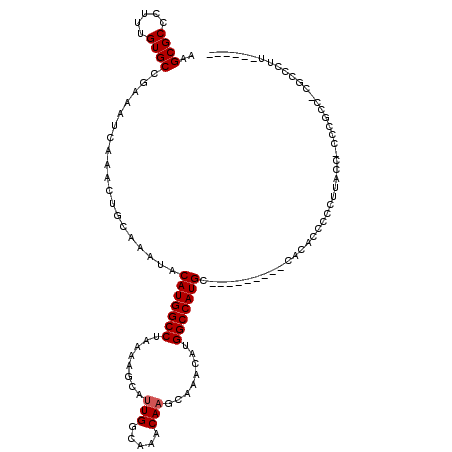

| Location | 19,208,605 – 19,208,722 |
|---|---|
| Length | 117 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 89.73 |
| Mean single sequence MFE | -36.86 |
| Consensus MFE | -26.39 |
| Energy contribution | -26.87 |
| Covariance contribution | 0.48 |
| Combinations/Pair | 1.03 |
| Mean z-score | -2.57 |
| Structure conservation index | 0.72 |
| SVM decision value | 0.86 |
| SVM RNA-class probability | 0.868354 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 19208605 117 + 22224390 -GAUGCAUGGCCAUGUUUGCUUGUUUGCCAAUGCUUUUAGGCCAUGUAUUUGCAGUUUGAUUUCGGCACAAAGGGCGCUUUGAACUGCUGUCG--UAUUUGCAUUUGCUCAAUCAAACAG -(((((((((((.((...(((((.....))).))...)))))))))))))....((((((((..(((((((((....)))))...(((.....--.....)))..)))).)))))))).. ( -35.70) >DroSec_CAF1 29980 114 + 1 ----GCAUGGCCAUGUUUGCUUGUUUGCCAAUGCUUUUAGGCCAUGUAUUUGCAGUUUGAUUUCGGCACAAAGGGCGCUUUGAACUGCUGUCC--UAUUUGCAUUGGCUCAAUCAAACAG ----.........((((((.(((...((((((((...((((.((.......((((((..(...((.(......).))..)..)))))))).))--))...)))))))).))).)))))). ( -39.51) >DroSim_CAF1 5250 114 + 1 ----GCAUGGCCAUGUUUGCUUGUUUGCCAAUGCUUUUAGGCCAUGUAUUUGCAGUUUGAUUUCGGCACAAAGGGCGCUUUGAACUGCUGUCC--UAUUUGCAUUGGCUCAAUCAAACAG ----.........((((((.(((...((((((((...((((.((.......((((((..(...((.(......).))..)..)))))))).))--))...)))))))).))).)))))). ( -39.51) >DroEre_CAF1 27218 109 + 1 -------UGGCCGAGUU----UGUUUGCCAAUGCUUUUAGGCCAUGUAUUUGCAGUUUGAUUACGGCACAAAGGGCGCUUUGAACUGCAGUCGUGUAUUUGCAUGUGCUCAAUCAAACAU -------((((((((((----((.....))).))))...)))))(((....)))((((((((..(((((....(((((........)).)))((((....))))))))).)))))))).. ( -30.80) >DroYak_CAF1 27752 114 + 1 GAGUACAUGGCCGAGUU----UGUUUGCCAAUGCUUUUAGGCCAUGUAUUUGCAGUUUGAUUCCGGCACAAAGGGCGCUUUGAACUGCAGUCG--UAUUUGCAUUUGCUCAAUCAAACGG (((((((((((((((((----((.....))).))))...))))))))))))...((((((((..(((((((((....)))))...(((((...--...)))))..)))).)))))))).. ( -38.80) >consensus ____GCAUGGCCAUGUUUGCUUGUUUGCCAAUGCUUUUAGGCCAUGUAUUUGCAGUUUGAUUUCGGCACAAAGGGCGCUUUGAACUGCUGUCG__UAUUUGCAUUUGCUCAAUCAAACAG ....((((((((.........((.....)).........)))))))).......((((((((..(((.(((((....)))))...(((............)))...))).)))))))).. (-26.39 = -26.87 + 0.48)



| Location | 19,208,605 – 19,208,722 |
|---|---|
| Length | 117 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 89.73 |
| Mean single sequence MFE | -30.26 |
| Consensus MFE | -26.33 |
| Energy contribution | -27.13 |
| Covariance contribution | 0.80 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.33 |
| Structure conservation index | 0.87 |
| SVM decision value | 2.79 |
| SVM RNA-class probability | 0.997069 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 19208605 117 - 22224390 CUGUUUGAUUGAGCAAAUGCAAAUA--CGACAGCAGUUCAAAGCGCCCUUUGUGCCGAAAUCAAACUGCAAAUACAUGGCCUAAAAGCAUUGGCAAACAAGCAAACAUGGCCAUGCAUC- ..((((((((((((...(((.....--.....)))))))...((((.....))))...))))))))........(((((((.....((.(((.....)))))......)))))))....- ( -30.40) >DroSec_CAF1 29980 114 - 1 CUGUUUGAUUGAGCCAAUGCAAAUA--GGACAGCAGUUCAAAGCGCCCUUUGUGCCGAAAUCAAACUGCAAAUACAUGGCCUAAAAGCAUUGGCAAACAAGCAAACAUGGCCAUGC---- .((((((.(((.((((((((...((--((.((((((((....((((.....)))).(....).)))))).......)).))))...))))))))...))).)))))).........---- ( -32.51) >DroSim_CAF1 5250 114 - 1 CUGUUUGAUUGAGCCAAUGCAAAUA--GGACAGCAGUUCAAAGCGCCCUUUGUGCCGAAAUCAAACUGCAAAUACAUGGCCUAAAAGCAUUGGCAAACAAGCAAACAUGGCCAUGC---- .((((((.(((.((((((((...((--((.((((((((....((((.....)))).(....).)))))).......)).))))...))))))))...))).)))))).........---- ( -32.51) >DroEre_CAF1 27218 109 - 1 AUGUUUGAUUGAGCACAUGCAAAUACACGACUGCAGUUCAAAGCGCCCUUUGUGCCGUAAUCAAACUGCAAAUACAUGGCCUAAAAGCAUUGGCAAACA----AACUCGGCCA------- ..(((((((((.(((((((((..........))))((.....))......)))))..)))))))))..........(((((....((..(((.....))----).)).)))))------- ( -24.30) >DroYak_CAF1 27752 114 - 1 CCGUUUGAUUGAGCAAAUGCAAAUA--CGACUGCAGUUCAAAGCGCCCUUUGUGCCGGAAUCAAACUGCAAAUACAUGGCCUAAAAGCAUUGGCAAACA----AACUCGGCCAUGUACUC ..((((((((((((...((((....--....))))))))...((((.....))))...))))))))......(((((((((....((..(((.....))----).)).)))))))))... ( -31.60) >consensus CUGUUUGAUUGAGCAAAUGCAAAUA__CGACAGCAGUUCAAAGCGCCCUUUGUGCCGAAAUCAAACUGCAAAUACAUGGCCUAAAAGCAUUGGCAAACAAGCAAACAUGGCCAUGC____ ..((((((((((((...(((............)))))))...((((.....))))...))))))))........(((((((........(((.....)))........)))))))..... (-26.33 = -27.13 + 0.80)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:45:35 2006