
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 19,208,167 – 19,208,263 |
| Length | 96 |
| Max. P | 0.994268 |

| Location | 19,208,167 – 19,208,263 |
|---|---|
| Length | 96 |
| Sequences | 4 |
| Columns | 107 |
| Reading direction | forward |
| Mean pairwise identity | 74.54 |
| Mean single sequence MFE | -22.50 |
| Consensus MFE | -11.86 |
| Energy contribution | -13.80 |
| Covariance contribution | 1.94 |
| Combinations/Pair | 1.14 |
| Mean z-score | -2.74 |
| Structure conservation index | 0.53 |
| SVM decision value | 2.46 |
| SVM RNA-class probability | 0.994268 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
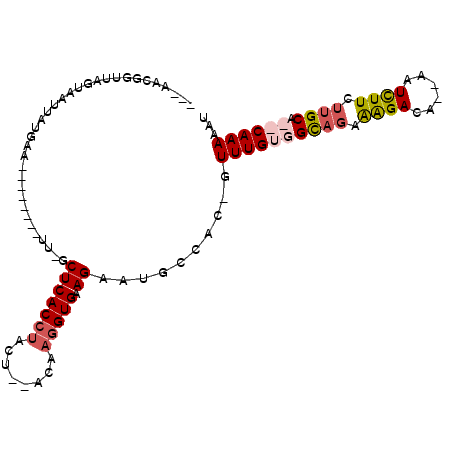
>X_DroMel_CAF1 19208167 96 + 22224390 AUUUUUGUGUGCAAGAAGAUU--UGUCUUUCUGCCACAAACUGUGGAAUGCUACACCUUGUUUAGUAGGUGAGC-AA--------UUCAUAAUUACCAAACGUUGGU ...((((((.(((..((((..--..))))..))))))))).((((((.((((.(((((........))))))))-).--------))))))...((((.....)))) ( -33.10) >DroSec_CAF1 29530 94 + 1 AUUUUUGUGUGCGAGAAGACU--UGUCCUUCUGCCACAAACUGUGGAAUUCUUCACCUUGUUUAGUGUGUGAGCCAA--------UUCAUAAUUACCAAUCGUU--- ...((((((.((.(((((...--....))))))))))))).((((((...((((((((.....)).))).)))....--------)))))).............--- ( -19.20) >DroEre_CAF1 26773 96 + 1 AUUUUUG--UGCAAGAAAAUU--UGUCUUUCUACCACAAAA-AUAACAUUCUUCACCUUGA--GGUAGGUGAGC-AAUUCAUAAUUACAUAAUUACUUCCCCUU--- (((((((--((..(((((...--....)))))..)))))))-))...(((.(((((((...--...))))))).-)))..........................--- ( -18.00) >DroYak_CAF1 27290 91 + 1 AUUUUUG--UGCAUGAAGAUUUCUGUCUCUUUGCCACAAAA-AUAGCAUUCUUCACCUUGA--AUUAGGUGAGC-AAA-------UUCAUAAUUACUAACCUUU--- (((((((--((((...((((....))))...)).)))))))-)).((.....((((((...--...))))))))-...-------...................--- ( -19.70) >consensus AUUUUUG__UGCAAGAAGAUU__UGUCUUUCUGCCACAAAA_AUAGAAUUCUUCACCUUGA__AGUAGGUGAGC_AA________UUCAUAAUUACCAACCGUU___ ...((((((.(((..(((((....)))))..)))))))))...........(((((((........))))))).................................. (-11.86 = -13.80 + 1.94)



| Location | 19,208,167 – 19,208,263 |
|---|---|
| Length | 96 |
| Sequences | 4 |
| Columns | 107 |
| Reading direction | reverse |
| Mean pairwise identity | 74.54 |
| Mean single sequence MFE | -22.27 |
| Consensus MFE | -11.00 |
| Energy contribution | -12.25 |
| Covariance contribution | 1.25 |
| Combinations/Pair | 1.19 |
| Mean z-score | -2.50 |
| Structure conservation index | 0.49 |
| SVM decision value | 1.42 |
| SVM RNA-class probability | 0.952602 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 19208167 96 - 22224390 ACCAACGUUUGGUAAUUAUGAA--------UU-GCUCACCUACUAAACAAGGUGUAGCAUUCCACAGUUUGUGGCAGAAAGACA--AAUCUUCUUGCACACAAAAAU ((((.....))))......(((--------(.-((((((((........))))).))))))).....(((((((((..((((..--..))))..))).))))))... ( -27.40) >DroSec_CAF1 29530 94 - 1 ---AACGAUUGGUAAUUAUGAA--------UUGGCUCACACACUAAACAAGGUGAAGAAUUCCACAGUUUGUGGCAGAAGGACA--AGUCUUCUCGCACACAAAAAU ---..(((((.((....)).))--------))).......((((......)))).............((((((((((((((...--..)))))).)).))))))... ( -19.90) >DroEre_CAF1 26773 96 - 1 ---AAGGGGAAGUAAUUAUGUAAUUAUGAAUU-GCUCACCUACC--UCAAGGUGAAGAAUGUUAU-UUUUGUGGUAGAAAGACA--AAUUUUCUUGCA--CAAAAAU ---..(((..((((((((((....)))).)))-)))..)))(((--....)))..........((-(((((((..((((((...--..))))))..))--))))))) ( -21.90) >DroYak_CAF1 27290 91 - 1 ---AAAGGUUAGUAAUUAUGAA-------UUU-GCUCACCUAAU--UCAAGGUGAAGAAUGCUAU-UUUUGUGGCAAAGAGACAGAAAUCUUCAUGCA--CAAAAAU ---..((((.(((((.......-------.))-))).)))).((--((........))))...((-(((((((.((..((((......))))..))))--))))))) ( -19.90) >consensus ___AACGGUUAGUAAUUAUGAA________UU_GCUCACCUACU__ACAAGGUGAAGAAUGCCAC_GUUUGUGGCAGAAAGACA__AAUCUUCUUGCA__CAAAAAU ..................................(((((((........))))).))..........((((((((((.((((......)))).)))).))))))... (-11.00 = -12.25 + 1.25)


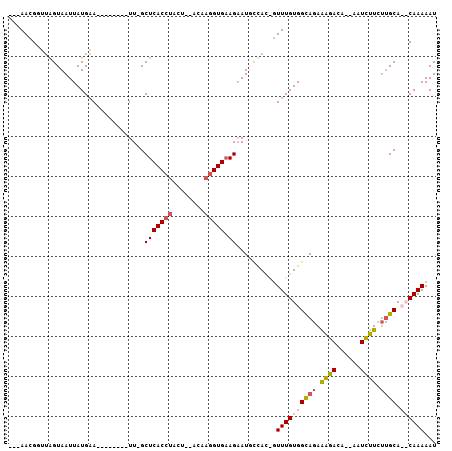
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:45:33 2006