
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 19,193,634 – 19,193,734 |
| Length | 100 |
| Max. P | 0.927591 |

| Location | 19,193,634 – 19,193,734 |
|---|---|
| Length | 100 |
| Sequences | 3 |
| Columns | 103 |
| Reading direction | forward |
| Mean pairwise identity | 80.26 |
| Mean single sequence MFE | -26.87 |
| Consensus MFE | -16.01 |
| Energy contribution | -16.13 |
| Covariance contribution | 0.12 |
| Combinations/Pair | 1.15 |
| Mean z-score | -2.01 |
| Structure conservation index | 0.60 |
| SVM decision value | 0.15 |
| SVM RNA-class probability | 0.605750 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
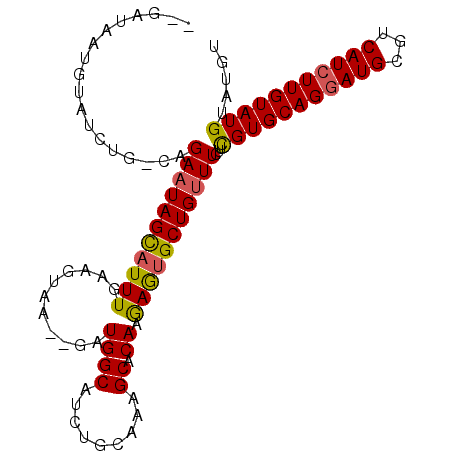
>X_DroMel_CAF1 19193634 100 + 22224390 AAGAUAAUGUAUCUG-CAGAAUAGCAUUUGAAGUAA--GAUGGCAUCUGCAAAGCACAAGAGCGCUGUUUCUCCGUGCAGGAUGCGUCAUCUUGUAUGUAUGU (((((.(((((((((-((((....(((((......)--))))...)))))...((((.((((......))))..)))).)))))))).))))).......... ( -29.20) >DroSec_CAF1 15253 100 + 1 --GAUAAUGUAUCUG-CAGAAUAGCAUUUGAAGUGAGAGAUGGCAUCUGCAAAGCACAAGAGUGCUGAUUCUUCGUGCAGGAUGCGUCAUCUUGUAUGUAUGU --(((.(((((((((-((((....(((((........)))))...)))))...((((((((((....)))))).)))).)))))))).)))............ ( -29.70) >DroEre_CAF1 12510 87 + 1 ---A-----UAUCUGCAGGAAUAGUAUUUGAAGUAA--GAUGGCAUCGGAAAAGCACAAAAAUGCUGUUU-UUUGUGCAGCAUGCGUCAUCUUGUAUG----- ---.-----....(((((((...((((((((.((..--....)).)))))...(((((((((......))-)))))))....)))....)))))))..----- ( -21.70) >consensus __GAUAAUGUAUCUG_CAGAAUAGCAUUUGAAGUAA__GAUGGCAUCUGCAAAGCACAAGAGUGCUGUUUCUUCGUGCAGGAUGCGUCAUCUUGUAUGUAUGU ..................(((((((((((...........((((.........)).)).)))))))))))...(((((((((((...)))))))))))..... (-16.01 = -16.13 + 0.12)


| Location | 19,193,634 – 19,193,734 |
|---|---|
| Length | 100 |
| Sequences | 3 |
| Columns | 103 |
| Reading direction | reverse |
| Mean pairwise identity | 80.26 |
| Mean single sequence MFE | -21.37 |
| Consensus MFE | -13.34 |
| Energy contribution | -13.57 |
| Covariance contribution | 0.23 |
| Combinations/Pair | 1.11 |
| Mean z-score | -2.44 |
| Structure conservation index | 0.62 |
| SVM decision value | 1.18 |
| SVM RNA-class probability | 0.927591 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
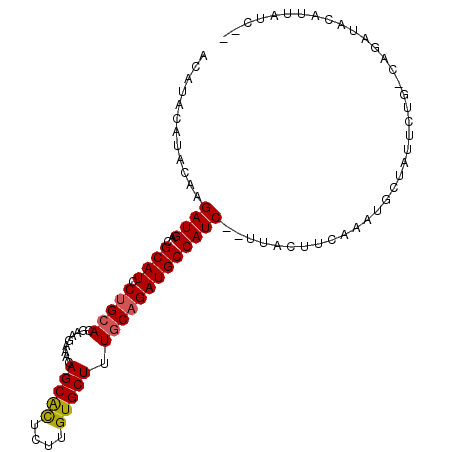
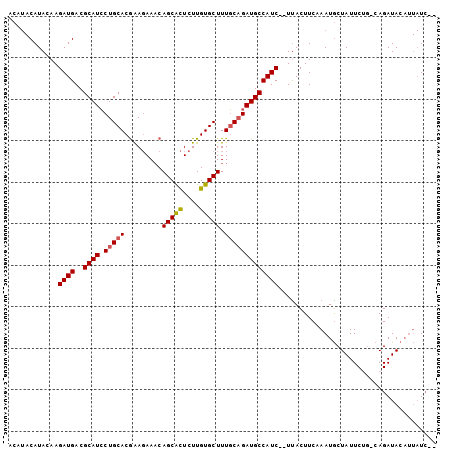
>X_DroMel_CAF1 19193634 100 - 22224390 ACAUACAUACAAGAUGACGCAUCCUGCACGGAGAAACAGCGCUCUUGUGCUUUGCAGAUGCCAUC--UUACUUCAAAUGCUAUUCUG-CAGAUACAUUAUCUU ..........((((((..(((((.((((.(......)(((((....))))).)))))))))))))--))........(((......)-))............. ( -24.20) >DroSec_CAF1 15253 100 - 1 ACAUACAUACAAGAUGACGCAUCCUGCACGAAGAAUCAGCACUCUUGUGCUUUGCAGAUGCCAUCUCUCACUUCAAAUGCUAUUCUG-CAGAUACAUUAUC-- ...........(((((..(((((.((((.((....))(((((....))))).))))))))))))))...........(((......)-))...........-- ( -23.20) >DroEre_CAF1 12510 87 - 1 -----CAUACAAGAUGACGCAUGCUGCACAAA-AAACAGCAUUUUUGUGCUUUUCCGAUGCCAUC--UUACUUCAAAUACUAUUCCUGCAGAUA-----U--- -----.....((((((..((((...(((((((-((......))))))))).......))))))))--)).........................-----.--- ( -16.70) >consensus ACAUACAUACAAGAUGACGCAUCCUGCACGAAGAAACAGCACUCUUGUGCUUUGCAGAUGCCAUC__UUACUUCAAAUGCUAUUCUG_CAGAUACAUUAUC__ ............((((..((((.(((((.........(((((....))))).)))))))))))))...................................... (-13.34 = -13.57 + 0.23)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:45:16 2006