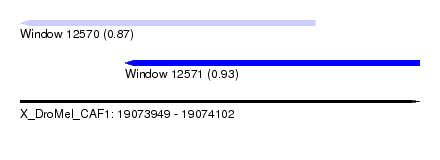

| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 19,073,949 – 19,074,102 |
| Length | 153 |
| Max. P | 0.928898 |

| Location | 19,073,949 – 19,074,062 |
|---|---|
| Length | 113 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 80.26 |
| Mean single sequence MFE | -27.13 |
| Consensus MFE | -16.55 |
| Energy contribution | -18.05 |
| Covariance contribution | 1.50 |
| Combinations/Pair | 1.15 |
| Mean z-score | -2.18 |
| Structure conservation index | 0.61 |
| SVM decision value | 0.86 |
| SVM RNA-class probability | 0.867701 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 19073949 113 - 22224390 UGUUGUUUCAAUACGAAAUACGCUAUGUGUAUUCGUGACUUAGUUAUGAGAAAUUGUCAAAUAAGUUAC-------GAGCGACGUGCAAAUUUAAUAUUUCUGUAACUUUUAAACAACCA .(((((((.(((((((((((.......(((((((((((((((...(..(....)..)....))))))))-------)).....))))).......)))))).)))...)).))))))).. ( -25.34) >DroSec_CAF1 25547 103 - 1 UGUUGUUUCAAUGCGAUAUACGAUUUGUGUAUCCGUGACUUAGUUAUGACAAAUUGUCAAAUGAGUUAC-------GAGCGACGUGCAAAUGUCAUAUUUCUGUAACUUU---------- ....(((.((..(((((((((.....)))))))(((((((((....((((.....))))..))))))))-------).))(((((....))))).......)).)))...---------- ( -29.20) >DroSim_CAF1 27467 103 - 1 UGUUGUUUCAAUACGAUAUACGAUUUGUGUAUCCGUGACUUAGUUAUGACAAAUUGUCAAAUGAGUUAC-------GAGCGACGUGCAAAUGUCAUAUUUCUGUAACUUU---------- .(((..........(((((((.....)))))))(((((((((....((((.....))))..))))))))-------))))(((((....)))))................---------- ( -26.50) >DroYak_CAF1 28033 117 - 1 UGUU---CUAAUACGAAAUAAUUGUUGCUUAUCCGUGACUUAGUUAUGACAAAUUGUCGAAUGAGUUACGAGCUACGAGCGACGUGCAAAUGUCAUGAUUCUGUAACUUCGAAACAUCGA ....---................(((((..((((((((((((....((((.....))))..))))))))).((.....))(((((....)))))..)))...))))).((((....)))) ( -27.50) >consensus UGUUGUUUCAAUACGAAAUACGAUUUGUGUAUCCGUGACUUAGUUAUGACAAAUUGUCAAAUGAGUUAC_______GAGCGACGUGCAAAUGUCAUAUUUCUGUAACUUU__________ .(((((((......(((((((.....))))))).((((((((....((((.....))))..)))))))).......)))))))..................................... (-16.55 = -18.05 + 1.50)

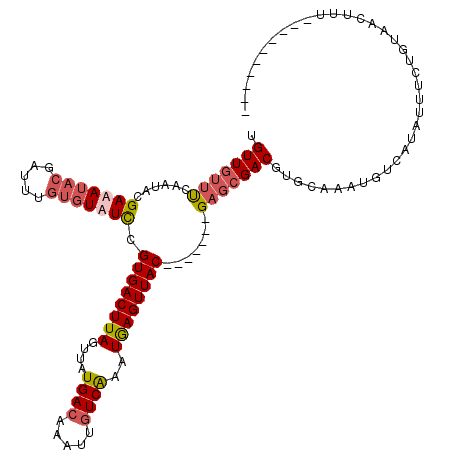
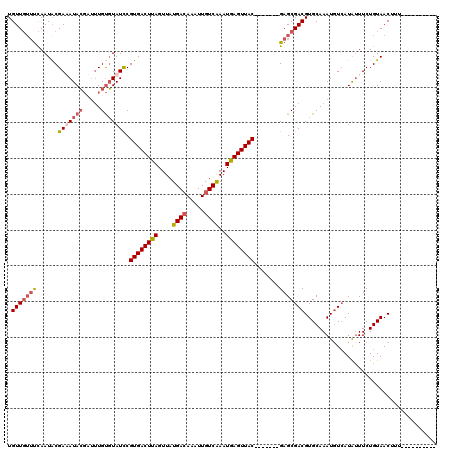
| Location | 19,073,989 – 19,074,102 |
|---|---|
| Length | 113 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 83.98 |
| Mean single sequence MFE | -28.66 |
| Consensus MFE | -18.65 |
| Energy contribution | -20.03 |
| Covariance contribution | 1.38 |
| Combinations/Pair | 1.19 |
| Mean z-score | -2.54 |
| Structure conservation index | 0.65 |
| SVM decision value | 1.19 |
| SVM RNA-class probability | 0.928898 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 19073989 113 - 22224390 CAAAUUAAGGUUUAGUGAAACAACGAGAAAAUGGGAAACAUGUUGUUUCAAUACGAAAUACGCUAUGUGUAUUCGUGACUUAGUUAUGAGAAAUUGUCAAAUAAGUUAC-------GAGC .........((....((((((((((........(....).))))))))))..))...((((((...))))))((((((((((...(..(....)..)....))))))))-------)).. ( -27.20) >DroSec_CAF1 25577 113 - 1 AAAAUUAAAGUUUACUGAAACAACGCGAGAAUGGGAAACAUGUUGUUUCAAUGCGAUAUACGAUUUGUGUAUCCGUGACUUAGUUAUGACAAAUUGUCAAAUGAGUUAC-------GAGC ...............((((((((((........(....).))))))))))..(((((((((.....)))))))(((((((((....((((.....))))..))))))))-------).)) ( -31.50) >DroSim_CAF1 27497 113 - 1 AAAAUUAAGGUUUAGUGAAACAACGCGAGAAUGGGAAACAUGUUGUUUCAAUACGAUAUACGAUUUGUGUAUCCGUGACUUAGUUAUGACAAAUUGUCAAAUGAGUUAC-------GAGC .........((....((((((((((........(....).))))))))))..))(((((((.....)))))))(((((((((....((((.....))))..))))))))-------)... ( -31.70) >DroYak_CAF1 28073 117 - 1 GGAAUUAAAGUUUAGUGAAGCAAUGCGAAACUGGCAAACAUGUU---CUAAUACGAAAUAAUUGUUGCUUAUCCGUGACUUAGUUAUGACAAAUUGUCGAAUGAGUUACGAGCUACGAGC (((((......(((((...((...))...))))).......)))---))............((((.(((....(((((((((....((((.....))))..)))))))))))).)))).. ( -24.22) >consensus AAAAUUAAAGUUUAGUGAAACAACGCGAAAAUGGGAAACAUGUUGUUUCAAUACGAAAUACGAUUUGUGUAUCCGUGACUUAGUUAUGACAAAUUGUCAAAUGAGUUAC_______GAGC .........((....((((((((((........(....).))))))))))..))(((((((.....))))))).((((((((....((((.....))))..))))))))........... (-18.65 = -20.03 + 1.38)


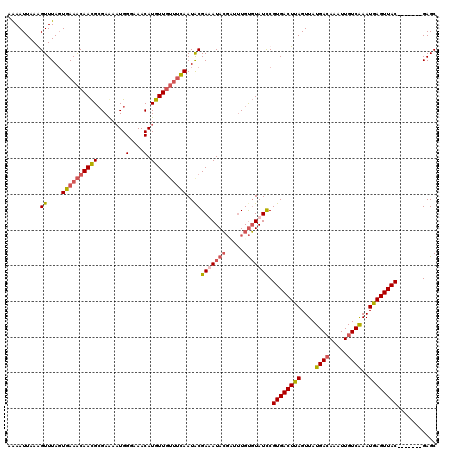
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:44:30 2006