
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 2,099,801 – 2,099,912 |
| Length | 111 |
| Max. P | 0.945908 |

| Location | 2,099,801 – 2,099,912 |
|---|---|
| Length | 111 |
| Sequences | 4 |
| Columns | 119 |
| Reading direction | forward |
| Mean pairwise identity | 83.38 |
| Mean single sequence MFE | -34.34 |
| Consensus MFE | -27.06 |
| Energy contribution | -29.12 |
| Covariance contribution | 2.06 |
| Combinations/Pair | 1.15 |
| Mean z-score | -2.20 |
| Structure conservation index | 0.79 |
| SVM decision value | 1.36 |
| SVM RNA-class probability | 0.945908 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 2099801 111 + 22224390 GAACAGGUGUCCCAUCCGAAGC-UCUAUUCAGGGAGGCACACCGAUUCCGGGUACAGUU-----UCGGCUUAAUCCGGAAUCACUUUGCACCCCUGAUUUGAUCCGUAA--CCCCCACC .....((((.......((....-((...((((((..(((....((((((((((..(((.-----...)))..))))))))))....)))..))))))...))..))...--....)))) ( -36.34) >DroSec_CAF1 487 116 + 1 GAACAGGUGUCCCAUCCGAAGU-UCUAUUCAGGGGCGCACACCGAUUCCGGGUACAGUACCGUACCGGCUUAAUCCGGAAUCACUUUGCACCCCUGAUUUGAUCCGUAA--CCCCCACC .....((((.......((....-((...(((((((.(((....((((((.(((((......)))))((......))))))))....))).)))))))...))..))...--....)))) ( -39.54) >DroEre_CAF1 957 112 + 1 GAACAGGUGUCCCAUCCAGAGCAUCUCUUCAGGAAUGCACACCGAUUCCGGGUACAUUU-----UCGGUUCGAUCUGGAAUCGCUUUGCAGUCCUGAUUUGAUCCAUAA--CCCCCACC .....((((........((((...))))((((((.((((...(((((((((((..((..-----...))...)))))))))))...)))).))))))............--....)))) ( -33.40) >DroYak_CAF1 181 114 + 1 GAACAGGUGUCCCAUCCGCAGCAUCUCUUCAGGAAUGCACAGCGAUUCCGGACACAGUC-----UCGGUUCGAUCUGAAAUCGCUUUGCAGCCCCGAUUUGAUUCACAAUCCCCCCACC (((.((((((..(....)..)))))).))).((..((((.(((((((((((((...)))-----.)))..........))))))).))))..)).((......)).............. ( -28.10) >consensus GAACAGGUGUCCCAUCCGAAGC_UCUAUUCAGGAAUGCACACCGAUUCCGGGUACAGUU_____UCGGCUCAAUCCGGAAUCACUUUGCACCCCUGAUUUGAUCCAUAA__CCCCCACC .....((((...........(.(((...((((((.((((.(.(((((((((((..((((.......))))..))))))))))).).)))).))))))...))).)..........)))) (-27.06 = -29.12 + 2.06)
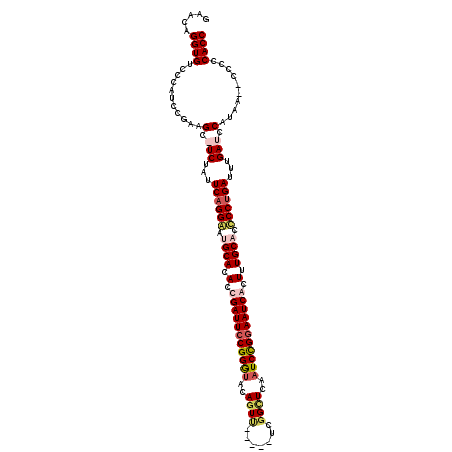


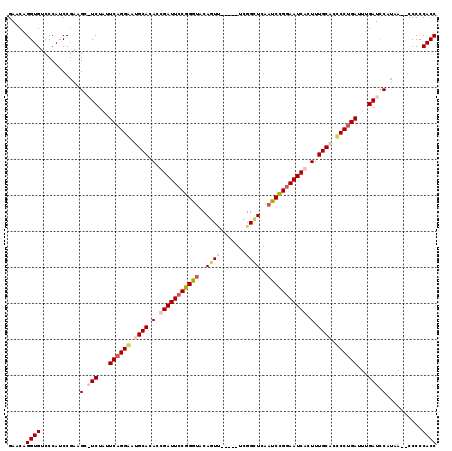
| Location | 2,099,801 – 2,099,912 |
|---|---|
| Length | 111 |
| Sequences | 4 |
| Columns | 119 |
| Reading direction | reverse |
| Mean pairwise identity | 83.38 |
| Mean single sequence MFE | -40.22 |
| Consensus MFE | -34.10 |
| Energy contribution | -33.79 |
| Covariance contribution | -0.31 |
| Combinations/Pair | 1.32 |
| Mean z-score | -1.49 |
| Structure conservation index | 0.85 |
| SVM decision value | 0.16 |
| SVM RNA-class probability | 0.614289 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 2099801 111 - 22224390 GGUGGGGG--UUACGGAUCAAAUCAGGGGUGCAAAGUGAUUCCGGAUUAAGCCGA-----AACUGUACCCGGAAUCGGUGUGCCUCCCUGAAUAGA-GCUUCGGAUGGGACACCUGUUC ((((....--...((((((...(((((((.(((.(.(((((((((....((....-----..))....))))))))).).))).)))))))...))-..)))).......))))..... ( -40.14) >DroSec_CAF1 487 116 - 1 GGUGGGGG--UUACGGAUCAAAUCAGGGGUGCAAAGUGAUUCCGGAUUAAGCCGGUACGGUACUGUACCCGGAAUCGGUGUGCGCCCCUGAAUAGA-ACUUCGGAUGGGACACCUGUUC ((((....--...((((((...(((((((((((.(.(((((((((......))((((((....)))))).))))))).).)))))))))))...))-..)))).......))))..... ( -50.94) >DroEre_CAF1 957 112 - 1 GGUGGGGG--UUAUGGAUCAAAUCAGGACUGCAAAGCGAUUCCAGAUCGAACCGA-----AAAUGUACCCGGAAUCGGUGUGCAUUCCUGAAGAGAUGCUCUGGAUGGGACACCUGUUC ((((...(--((..(.(((...((((((.((((.(.(((((((...(((...)))-----....(....)))))))).).)))).))))))...))).)....)))....))))..... ( -33.30) >DroYak_CAF1 181 114 - 1 GGUGGGGGGAUUGUGAAUCAAAUCGGGGCUGCAAAGCGAUUUCAGAUCGAACCGA-----GACUGUGUCCGGAAUCGCUGUGCAUUCCUGAAGAGAUGCUGCGGAUGGGACACCUGUUC ((((......(((((.(((...(((((..((((.(((((((((.....)).(((.-----(((...))))))))))))).))))..)))))...)))..)))))......))))..... ( -36.50) >consensus GGUGGGGG__UUACGGAUCAAAUCAGGGCUGCAAAGCGAUUCCAGAUCAAACCGA_____AACUGUACCCGGAAUCGGUGUGCAUCCCUGAAGAGA_GCUUCGGAUGGGACACCUGUUC ((((.........((((((...(((((((((((.(.(((((((((.......................))))))))).).)))))))))))...))...)))).......))))..... (-34.10 = -33.79 + -0.31)

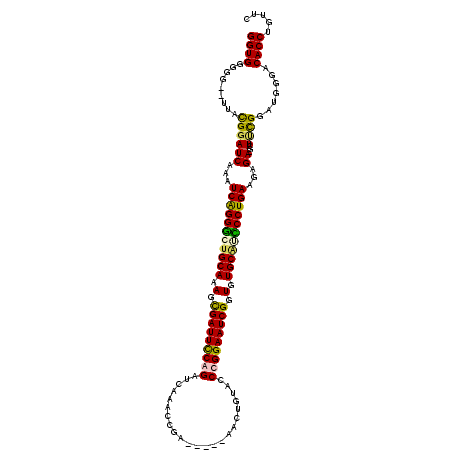
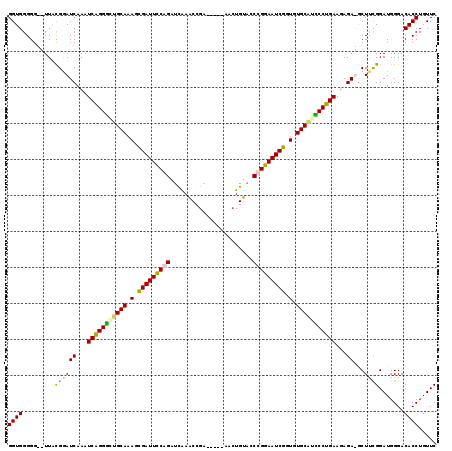
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:53:15 2006